Abstract
Tumours evade immune control by creating hostile microenvironments that perturb T cell metabolism and effector function1,2,3,4. However, it remains unclear how intra-tumoral T cells integrate and interpret metabolic stress signals. Here we report that ovarian cancer—an aggressive malignancy that is refractory to standard treatments and current immunotherapies5,6,7,8—induces endoplasmic reticulum stress and activates the IRE1α–XBP1 arm of the unfolded protein response9,10 in T cells to control their mitochondrial respiration and anti-tumour function. In T cells isolated from specimens collected from patients with ovarian cancer, upregulation of XBP1 was associated with decreased infiltration of T cells into tumours and with reduced IFNG mRNA expression. Malignant ascites fluid obtained from patients with ovarian cancer inhibited glucose uptake and caused N-linked protein glycosylation defects in T cells, which triggered IRE1α–XBP1 activation that suppressed mitochondrial activity and IFNγ production. Mechanistically, induction of XBP1 regulated the abundance of glutamine carriers and thus limited the influx of glutamine that is necessary to sustain mitochondrial respiration in T cells under glucose-deprived conditions. Restoring N-linked protein glycosylation, abrogating IRE1α–XBP1 activation or enforcing expression of glutamine transporters enhanced mitochondrial respiration in human T cells exposed to ovarian cancer ascites. XBP1-deficient T cells in the metastatic ovarian cancer milieu exhibited global transcriptional reprogramming and improved effector capacity. Accordingly, mice that bear ovarian cancer and lack XBP1 selectively in T cells demonstrate superior anti-tumour immunity, delayed malignant progression and increased overall survival. Controlling endoplasmic reticulum stress or targeting IRE1α–XBP1 signalling may help to restore the metabolic fitness and anti-tumour capacity of T cells in cancer hosts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source Data are provided for Figs. 1–4 and Extended Data Figs. 1–8. The NCBI GEO (Gene Expression Omnibus) accession number for RNA sequencing data reported in this paper is GSE118430. The datasets generated during the current study are available from the corresponding authors upon reasonable request.
References
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015).
Ho, P. C. et al. Phosphoenolpyruvate is a metabolic checkpoint of anti-tumor T cell responses. Cell 162, 1217–1228 (2015).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 374–388 (2016).
Anderson, K. G., Stromnes, I. M. & Greenberg, P. D. Obstacles posed by the tumor microenvironment to T cell activity: a case for synergistic therapies. Cancer Cell 31, 311–325 (2017).
Chae, C. S., Teran-Cabanillas, E. & Cubillos-Ruiz, J. R. Dendritic cell rehab: new strategies to unleash therapeutic immunity in ovarian cancer. Cancer Immunol. Immunother. 66, 969–977 (2017).
Matulonis, U. A. et al. Ovarian cancer. Nat. Rev. Dis. Primers 2, 16061 (2016).
Hamanishi, J. et al. Safety and antitumor activity of anti-PD-1 antibody, nivolumab, in patients with platinum-resistant ovarian cancer. J. Clin. Oncol. 33, 4015–4022 (2015).
Kershaw, M. H. et al. A phase I study on adoptive immunotherapy using gene-modified T cells for ovarian cancer. Clin. Cancer Res. 12, 6106–6115 (2006).
Yoshida, H., Matsui, T., Yamamoto, A., Okada, T. & Mori, K. XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107, 881–891 (2001).
Lee, A. H., Iwakoshi, N. N. & Glimcher, L. H. XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol. Cell. Biol. 23, 7448–7459 (2003).
Cubillos-Ruiz, J. R., Bettigole, S. E. & Glimcher, L. H. Tumorigenic and immunosuppressive effects of endoplasmic reticulum stress in cancer. Cell 168, 692–706 (2017).
Cubillos-Ruiz, J. R. et al. ER stress sensor XBP1 controls anti-tumor immunity by disrupting dendritic cell homeostasis. Cell 161, 1527–1538 (2015).
Yan, D., Wang, H. W., Bowman, R. L. & Joyce, J. A. STAT3 and STAT6 signaling pathways synergize to promote cathepsin secretion from macrophages via IRE1α activation. Cell Rep. 16, 2914–2927 (2016).
Condamine, T. et al. Lectin-type oxidized LDL receptor-1 distinguishes population of human polymorphonuclear myeloid-derived suppressor cells in cancer patients. Sci. Immunol. 1, aaf8943 (2016).
Kipps, E., Tan, D. S. & Kaye, S. B. Meeting the challenge of ascites in ovarian cancer: new avenues for therapy and research. Nat. Rev. Cancer 13, 273–282 (2013).
Bamias, A. et al. Significant differences of lymphocytes isolated from ascites of patients with ovarian cancer compared to blood and tumor lymphocytes. Association of CD3+CD56+ cells with platinum resistance. Gynecol. Oncol. 106, 75–81 (2007).
Curiel, T. J. et al. Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nat. Med. 10, 942–949 (2004).
Lukesova, S. et al. Comparative study of various subpopulations of cytotoxic cells in blood and ascites from patients with ovarian carcinoma. Contemp. Oncol. (Pozn) 19, 290–299 (2015).
Knutson, K. L. et al. Regulatory T cells, inherited variation, and clinical outcome in epithelial ovarian cancer. Cancer Immunol. Immunother. 64, 1495–1504 (2015).
Kim, K. S. et al. Hypoxia enhances lysophosphatidic acid responsiveness in ovarian cancer cells and lysophosphatidic acid induces ovarian tumor metastasis in vivo. Cancer Res. 66, 7983–7990 (2006).
Denzel, M. S. & Antebi, A. Hexosamine pathway and (ER) protein quality control. Curr. Opin. Cell Biol. 33, 14–18 (2015).
Yang, C. et al. Glutamine oxidation maintains the TCA cycle and cell survival during impaired mitochondrial pyruvate transport. Mol. Cell 56, 414–424 (2014).
Wellen, K. E. et al. The hexosamine biosynthetic pathway couples growth factor-induced glutamine uptake to glucose metabolism. Genes Dev. 24, 2784–2799 (2010).
Wang, H. et al. Endoplasmic reticulum stress up-regulates Nedd4-2 to induce autophagy. FASEB J. 30, 2549–2556 (2016).
Jeon, Y. J. et al. Regulation of glutamine carrier proteins by RNF5 determines breast cancer response to ER stress-inducing chemotherapies. Cancer Cell 27, 354–369 (2015).
Hatanaka, T., Hatanaka, Y. & Setou, M. Regulation of amino acid transporter ATA2 by ubiquitin ligase Nedd4-2. J. Biol. Chem. 281, 35922–35930 (2006).
Scarlett, U. K. et al. Ovarian cancer progression is controlled by phenotypic changes in dendritic cells. J. Exp. Med. 209, 495–506 (2012).
Conejo-Garcia, J. R. et al. Tumor-infiltrating dendritic cell precursors recruited by a β-defensin contribute to vasculogenesis under the influence of Vegf-A. Nat. Med. 10, 950–958 (2004).
So, J. S. et al. Silencing of lipid metabolism genes through IRE1α-mediated mRNA decay lowers plasma lipids in mice. Cell Metab. 16, 487–499 (2012).
Roby, K. F. et al. Development of a syngeneic mouse model for events related to ovarian cancer. Carcinogenesis 21, 585–591 (2000).
Lee, A. H., Scapa, E. F., Cohen, D. E. & Glimcher, L. H. Regulation of hepatic lipogenesis by the transcription factor XBP1. Science 320, 1492–1496 (2008).
Iwawaki, T., Akai, R., Yamanaka, S. & Kohno, K. Function of IRE1 alpha in the placenta is essential for placental development and embryonic viability. Proc. Natl Acad. Sci. USA 106, 16657–16662 (2009).
Lee, A. H., Iwakoshi, N. N., Anderson, K. C. & Glimcher, L. H. Proteasome inhibitors disrupt the unfolded protein response in myeloma cells. Proc. Natl Acad. Sci. USA 100, 9946–9951 (2003).
Yuan, M., Breitkopf, S. B., Yang, X. & Asara, J. M. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Maglott, D., Ostell, J., Pruitt, K. D. & Tatusova, T. Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res. 39, D52–D57 (2011).
Acknowledgements
Our research was supported by the Irvington Institute Fellowship Program of the Cancer Research Institute (J.R.C.-R.), the Ann Schreiber Mentored Investigator Award of the Ovarian Cancer Research Fund Alliance (J.R.C.-R.), the Ovarian Cancer Academy Early-Career Investigator Award W81XWH-16-1-0438 of the Department of Defense (J.R.C.-R.), the Stand Up to Cancer Innovative Research Grant SU2C-AACR-IRG-03-16 (J.R.C.-R.), the Jacquie Liggett Fellowship Award of Hearing the Ovarian Cancer Whisper (J.R.C.-R.), Weill Cornell Medicine Funds (J.R.C.-R. and L.H.G.), NIH grant R01CA112663 (L.H.G.) and NIH SIG grant 1S10 OD017992-01 (S.Z.) for the Orbitrap Fusion mass spectrometer. We thank T. Iwawaki at Kanazawa Medical University for sharing the Ern1-floxed mouse strain; J. McCormick for expert assistance with cell sorting; L. Cohen-Gould and J. Jimenez for electron microscopy analysis; G. Zhang, Z. Cheng and T. Su for metabolic tracing experiments; all members of the Weill Cornell Epigenomics Facility for assistance with RNA sequencing, T. Walther for help collecting patient samples; J. M. Pérez-Sáez and J. Trillo-Tinoco for assistance with some experimental analyses and helpful suggestions; and L. Cantley and M. Goncalves for sharing valuable instruments and resources. We also thank all members of the Cubillos-Ruiz, Morales and Glimcher laboratories for helpful suggestions and critical reading of this manuscript.
Reviewer information
Nature thanks T. Curiel and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
M.S. designed and conducted most of the in vitro and in vivo experiments, analysed data and wrote the manuscript. T.A.S., C.-S.C., S.C., C.T., M.R., R.A.C., K.K.P., H.R.S., M.J.P.C. and J.P.C performed in vitro and in vivo experiments and analysed data. M.R.R performed in vivo experiments using p53/KRAS hosts and analysed data. C.K. and G.M. contributed to the design of certain Seahorse-related experiments and shared resources. S.E.B. performed mixed BM chimaeras, in vitro experiments and edited the manuscript. A.V.K. carried out computational analyses of RNA sequencing data. I.M. and S.Z. performed mass spectrometry experiments and analysed proteomics data. K.H. and D.Z. performed surgeries and provided patient specimens. P.C.R., G.A.R. and J.R.C.-G. contributed to the design of certain experiments, provided ideas and models, shared resources, analysed data and reviewed the manuscript. L.H.G. designed the research, provided models and resources, analysed data and reviewed the manuscript. J.R.C.-R. conceived the idea, designed and conducted the research, analysed data, wrote the manuscript and directed the project. J.R.C.-R. is lead senior author for this paper.
Corresponding authors
Ethics declarations
Competing interests
J.R.C.-R. and L.H.G. are co-founders of and scientific advisors for Quentis Therapeutics. S.E.B. is co-founder and employee of Quentis Therapeutics. L.H.G. also serves on the board of directors of and holds equity in GlaxoSmithKline Pharmaceuticals.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Reduced glucose uptake and defective N-linked protein glycosylation promotes IRE1α–XBP1 activation in human CD4+ T cells exposed to ascites.
a, Human CD4+ T cells were activated by CD3 and CD28 stimulation for 16 h in the absence or presence of supernatants from ovarian cancer ascites at the indicated concentrations. XBP1 expression was determined by qPCR (10%, n = 58; 50%, n = 25; 100%, n = 30). Data were normalized to endogenous expression of ACTB in each sample. Results are presented as per cent increase in expression compared with untreated controls. b, Histograms for FACS-based XBP1s staining in CD4+ T cells treated as indicated. Tm, tunicamycin; 4μ8C, inhibitor of the IRE1α RNase domain. Data were validated by three independent experiments. c, d, CD4+ T cells were treated with vitamin E (VitE, 50 μM) or vehicle (ethanol, 0.1%) for 1 h and then stimulated with anti-CD3 and anti-CD28 beads for 16 h in the presence of ascites. Intracellular reactive oxygen species staining by DCFDA (c) and XBP1 expression (d) (n = 10). Data are expressed as per cent response change compared with untreated controls. e, Glucose concentration in regular culture medium (cRPMI) and in seven independent samples of ovarian cancer ascites. Each dot represents the mean of two measurements. f, XBP1 expression in the samples described in a are displayed, based on the final glucose concentration in the culture after addition of ascites. Three independent ascites samples were used: A10 (triangles), A15 (circles) and A17 (squares) at three different concentrations (10, 50 and 100%). g, GLUT1 surface expression on the indicated cell types present in ovarian cancer ascites from six independent patients was determined by GLUT1.RBD staining (n = 6). Quantitative analysis (left) and representative histograms (right). h, Glucose uptake and XBP1s protein expression in activated CD4+ T cells exposed to three different ascites samples at 10 and 100% for 16 h (n = 14). Results are presented as relative to untreated controls. i, CD4+ T cells were activated with anti-CD3 and anti-CD28 beads in the presence or absence of ascites, for 16 h. Cells were lysed and the enriched glycoprotein fractions were analysed for N-linked glycosylation events by LC–MS/MS. Total ion chromatograms for N-glycosylation at amino acid 315 in DPP2 are shown. Numbers in blue indicate abundance of each glycan through quantification of the corresponding peak area. Data are representative of two independent experiments with similar results. j, CD4+ T cells were treated with 10 mM GlcNAc for 1 h and stimulated by CD3 and CD28 for 16 h in the presence of 10% ascites. Quantitative analyses for XBP1s protein by FACS (left, n = 6) and XBP1 gene expression by qPCR (right, n = 15) are presented as per cent response change compared with untreated controls. k, Relative basal and maximal OCR for CD4+ T cells exposed to 10% ascites analysed in Fig. 2g (n = 16 total from five independent experiments). Data are expressed as per cent response change compared with untreated controls. l, m, CD4+ T cells were activated by CD3 and CD28 stimulation for 16 h in the absence or presence of tunicamycin (Tm) at the indicated concentrations (n = 3). l, XBP1 expression was determined by qPCR. m, OCR profile of Tm-treated CD4+ T cells are shown relative to the untreated control. n, o, Relative quantification of basal (left) and maximal (right) OCR (n), and ECAR measurements (o) in all independent samples analysed in Fig. 2i (n = 9 total from three independent experiments). p, q, Relative quantification of basal (left) and maximal (right) OCR (p), and ECAR measurements (q) for the specimens described in Fig. 2j. (n = 5 total from two independent experiments). Data are presented as relative expression compared with matching controls that were not exposed to ascites. Data are shown as mean ± s.e.m. (a, c, d, f, g, k–m). n values represent biologically independent samples (a, c–h, j–q). One-way ANOVA with Bonferroni’s multiple comparisons test (a, l); two-tailed Student’s t-test (c, k); one-way ANOVA with Tukey’s multiple comparisons test (d, g); two-tailed paired Student’s t-test (j, n–q); nonparametric Spearman’s rank correlation test, Spearman coefficient (r) with P value (two-tailed); 95% confidence interval −0.8811 to −0.1626 (h). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. NS, not significant; MFI, mean fluorescence intensity; gMFI, geometric mean fluorescence intensity.
Extended Data Fig. 2 Characterization of mice devoid of XBP1 in T cells.
a, Deletion efficiency was analysed by qPCR using a primer set that specifically detects the exon 2 region of Xbp1. Data were normalized to endogenous expression of Actb and presented as relative expression compared with wild-type littermates (n = 8). b, Absolute cell numbers in the thymus, spleen and lymph nodes. c, FACS-based phenotyping of double-negative (CD4−CD8−), double-positive (CD4+CD8+) or single-positive (CD4+ or CD8+) thymocytes. d–g, Frequency of TCRβ+ cells (d, f) and CD4+ or CD8+ cells (gated on TCRβ+ cells) (e, g) in lymph nodes or spleen. h, i, Expression of CD44 and CD62L on both CD4+ (h) and CD8+ (i) TCRβ+ subsets in the spleen. j, Frequency of splenic TCRβ+CD4+FoxP3+ T cells. k, l, Frequency of non-T-cell populations among total live cells in spleen (k) and lymph nodes (l). b–g, n = 5; h–j, n = 3; k, l, Xbp1f/f (n = 4), Xbp1f/fCd4cre (n = 5). m, Reconstitution efficiency of CD4+ and CD8+ T cells in bone marrow and spleen from mixed bone marrow chimaeras (n = 3 per chimaera type). Chimaeras were generated with a mixture of wild-type bone marrow (CD45.1+) plus either Xbp1f/f or Xbp1f/fVav1cre bone marrow (CD45.2+). n, Flow cytometry assessing cell proliferation of CD4+ T cells stained with the division-tracking dye (Cell Trace Violet). Cells were left unstimulated or stimulated for 72 and 96 h with plate-bound anti-CD3 (5 μg ml−1) and soluble anti-CD28 (1 μg ml−1). Histograms (left) and proliferation index (right) are shown (n = 4). o, Cell-cycle analysis of CD4+ T cells activated for 72 h by staining with propidium iodide. Representative plots from two experiments. p, Transmission electron microscopy of in vitro activated wild-type versus XBP1-deficient CD4+ T cells. Naive CD4+ T cells isolated from three biologically independent mice were activated with plate-bound anti-CD3 and soluble anti-CD28 antibodies for 48 h. White arrowheads indicate the ER; M, mitochondria; magnification 12,000× (left); 50,000× (right). Average mitochondrial area of independent cells was estimated using ImageJ software. Xbp1f/f (n = 19); Xbp1f/fCd4cre (n = 29). q, Histogram (left) and quantification (right) for mitochondrial staining (Mitotracker) in in vitro activated CD4+ T cells (n = 2 from two independent experiments). r, Activated wild-type versus XBP1-deficient CD4+ T cells were incubated in glucose-containing, glucose-depleted or 2-deoxyglucose (2-DG, 10 mM)-treated medium for 6 h and PGC1α expression was analysed by immunoblot. β-Actin was used as loading control. Representative plots from two independent experiments. Data are shown as mean ± s.e.m. (a–n, p). n values represent biologically independent samples (a–n, p, q). Two-tailed Student’s t-tests (b–l, n, p); *P < 0.05; NS, not significant.
Extended Data Fig. 3 XBP1 inhibits glutamine influx in response to glucose deprivation.
a, b, Naive splenic CD4+ T cells isolated from wild-type mice were activated by CD3 and CD28 stimulation for 48 h and then incubated for 6 h in the indicated medium. a, Expression of gene markers related to ER stress (n = 4 from two independent experiments). Data are shown as per cent response change compared with control in the presence of medium containing glucose and glutamine. b, Maximal OCR was measured in CD4+ T cells in the presence or absence of glucose, and treated with corresponding medium (untreated, n = 5) or inhibitors blocking pyruvate (UK5099, n = 5), glutamine (BPTES, n = 4) or fatty acid (etomoxir, n = 4) oxidation. Data are presented as per cent response change compared with untreated control in the presence of glucose. c–i, Naive splenic CD4+ T cells isolated from wild-type (solid bars) or XBP1-deficient (hatched bars) mice were activated by CD3 and CD28 stimulation for 48 h, followed by culture in the presence or absence of glucose for 4.5 h, and then pulsed with [U-13C]glutamine for an additional 1.5 h in the same culture condition. Relative abundance of 13C-labelled metabolites and TCA intermediates including glutamine (c), glutamate (d), α-ketoglutarate (e), succinate (f), malate (g), citrate (h) and aspartate (i) was determined by LC–MS/MS. Data were normalized to cell number in all cases and are representative of two independent experiments with n = 2 biologically distinct samples per group. Data are shown as mean ± s.e.m. n values represent biologically independent samples (a, b). One-way ANOVA with Bonferroni’s multiple comparisons test (a); one-way ANOVA with Tukey’s multiple comparisons test (b); *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 4 XBP1 controls the abundance of glutamine transporters in glucose-deprived T cells.
a, b, Pre-activated wild-type or XBP1-deficient CD4+ T cells were incubated in the indicated medium for 6 h and then stained on poly-l-lysine coated discs using antibodies specific for ASCT2 (green) or SNAT2 (red). Nuclei are depicted in blue (DAPI staining). a, Representative confocal images of the indicated T cells from three experiments. b, The mean fluorescence intensity (MFI) of each glutamine transporter on about 50 individual cells from three independent slides (n = 150) was computationally quantified using the ImageJ software by two independent investigators in a blinded manner. Individual dots depict the average MFI of each independent analysis (n = 6). c, Naive splenic CD4+ T cells isolated from wild-type or XBP1-deficient mice were activated by CD3 and CD28 stimulation for 48 h and then incubated in medium that lacks glucose for 6 h. mRNA expression of genes encoding glutamine transporters was determined by qPCR (n = 6 from three experiments). Data were normalized to endogenous expression of Actb in each case. d, e, Pre-activated mouse CD4+ T cells were incubated in the indicated medium for 6 h in the presence or absence of proteasome inhibitor MG132 (10 μM). d, Protein levels of the glutamine transporter SNAT1 were determined by immunoblot analysis, in which β-actin was used as loading control. Representative image from five independent experiments. e, Densitometric quantification of SNAT1 (n = 5). Results are presented as relative expression compared with untreated control T cells incubated in glucose-containing medium. Data are shown as mean ± s.e.m. (b, c). n values represent biologically independent samples (c, e). Two-tailed Student’s t-tests (b); two-tailed paired Student’s t-tests (e); *P < 0.05, ***P < 0.001; NS, not significant.
Extended Data Fig. 5 Restoring glutamine influx enhances mitochondrial function in CD4+ T cells exposed to ascites.
a, b, Immunoblot (a) and densitometric quantification (b) of levels of ASCT2 and SNAT1 protein in human CD4+ T cells exposed to ovarian cancer ascites at the indicated concentrations for 16 h. β-Actin was used as loading control. Data are shown as the relative expression compared with untreated (0%) controls. n = 4 for 10% ascites; n = 4 for 50% ascites; n = 2 for 100% ascites. Data were generated from two independent experiments. c, d, Human CD4+ T cells were activated by CD3 and CD28 stimulation for 16 h in the absence or presence of 25% supernatants of ovarian cancer ascites, and DMKG (5 mM) was added to the cell culture during the last 4 h of incubation. OCR profile (c) and quantification of maximal OCR (d). Data are presented as relative expression compared with untreated controls incubated in the absence of ascites (n = 17 total from two independent experiments). e, f, Human CD4+ T cells activated by CD3 and CD28 stimulation and IL-2 (50 U ml−1) for 36 h were transduced with GFP-expressing retroviruses containing no insert (control) or the gene that encodes human SNAT1. GFP+ cells were sorted 3 days after transduction and expanded for an additional 48 h in the presence of CD3 and CD28 stimulation, and IL-2 (50 U ml−1). After 20 h of resting, cells were restimulated with CD3 and CD28 antibodies in the absence or presence of supernatants of ovarian cancer ascites for 16 h. e, Sorting plots showing GFP expression by CD4+ T cells that were left untreated or transduced with either control or SNAT1-expressing viruses. Representative plots from two experiments. f, Relative SLC38A1 expression levels in sorted cells after transduction (n = 6 total from two independent experiments). n values represent biologically independent samples (b, d, f). Data are shown as mean ± s.e.m. (b, c, f). One-way ANOVA with Bonferroni’s multiple comparisons test (b); two-tailed paired Student’s t-test (d); two-tailed Student’s t-test (f). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 6 IRE1α–XBP1 activation and ER stress responses in ovarian cancer-associated T cells isolated from mouse models of disease.
a, b, Expression of marker genes (Xbp1, Xbp1s, Hspa5 and Ddit3) for ER stress was determined by qPCR. Circles, CD4+ T cells; squares, CD8+ T cells. a, CD45+TCRβ+CD4+ and CD45+TCRβ+CD8+ cells were sorted from tumours (n = 8), spleens (n = 4) and lymph nodes (n = 4) of mice bearing advanced ovarian tumours driven by p53 and KRAS. b, CD45+TCRβ+CD4+ and CD45+TCRβ+CD8 cells were isolated from malignant ascites (n = 6), spleens (n = 8) and lymph nodes (n = 8) of mice bearing aggressive ID8-Defb29/Vegfa ovarian cancer (OvCa), and from spleens (n = 6) of naive mice as control. c, d, Expression of target genes (Bloc1s1 and Gm2a) of canonical regulated IRE1α-dependent decay (RIDD)—in CD4+ and CD8+ T cells isolated from different tissues of mice that bore ovarian cancer—was analysed by qPCR. c, T cells were sorted from tumours (CD4+ T cells, n = 5; CD8+ T cells, n = 4), spleens (n = 2) and lymph nodes (n = 2) of mice bearing advanced ovarian tumours driven by p53 and KRAS. d, T cells were isolated from malignant ascites (CD4+ T cells, n = 8; CD8+ T cells, n = 6), spleens (n = 13) and lymph nodes (n = 8) of mice that bore aggressive ID8-Defb29/Vegfa ovarian cancer. Data were normalized to Actb. Data are shown as mean ± s.e.m. (a–d). n values represent biologically independent samples (a–d). One-way ANOVA with Tukey’s multiple comparisons test (a, b); *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 7 IRE1α–XBP1 signalling alters function of T cells associated with ovarian cancer, and promotes malignant progression.
a, Transcriptional profiling of splenic CD44highCD62LlowCD4+ T cells sorted from naive wild-type versus XBP1-deficient mice. Expression of the differentially regulated genes identified in Fig. 4a is shown (n = 3 per group). b, c, e, f, Analysis of wild-type versus XBP1-deficient CD4+ T cells isolated from mice that bore metastatic ovarian cancer (n = 4 per group). b, Expression of previously reported target genes of regulated IRE1α-dependent mRNA decay (RIDD). c, Relative gene expression of master transcription factors controlling helper T cell differentiation. d, Intracellular staining for FoxP3 (left) and proportion of FoxP3+CD4+ T cells from wild-type (n = 5) and XBP1-deficient (n = 6) mice that bore metastatic ovarian cancer for 21 days. e, Predicted upstream regulators associated with the transcriptional changes observed. f, Enriched cellular functions in XBP1-deficient CD4+ T cells at tumour sites. Z-scores greater than 2 indicate functions predicted to be significantly increased in XBP1-deficient CD4+ T cells. g, Intracellular staining for IFNγ in CD45+CD3+CD4+ T cells from wild-type and conditional XBP1-deficient mice that bore metastatic ovarian cancer for 29 days (late stage). Representative plots from three independent experiments. h, Intracellular staining for perforin in CD45+CD3+CD8+ T cells from wild-type and conditional XBP1-deficient mice that bore metastatic ovarian cancer for 23 days. i, Intracellular staining for IFNγ in CD45+CD3+CD8+ T cells from wild-type and conditional XBP1-deficient mice that bore metastatic ovarian cancer for 29 days (late stage). Representative plots from two independent experiments. j, In vivo glucose uptake by CD44highCD4+ tumour-infiltrating lymphocytes in Xbp1f/f (n = 6) or Xbp1f/fCd4cre (n = 7) female mice that bore metastatic ovarian cancer. k, Representative TMRE staining for CD45+TCRβ+CD44+CD4+ T cells associated with ovarian cancer, from tumour-bearing Xbp1f/f (n = 3) or Xbp1f/fCd4cre (n = 4) mice. l, Peritoneal wash samples were collected from mice at 24 days after tumour challenge and the proportion of CD4+ T cells associated with ovarian cancer that expressed PD-1, CTLA4 and TIM3 in wild-type (n = 9) or XBP1-deficient (n = 8) mice was determined. m, Female mice (n = 4 per group) were implanted with ID8-Defb29/Vegfa ovarian cancer cells in the flank, and tumour growth was monitored over time (left). Tumours were resected at day 34 and final size was confirmed ex vivo (right). n, Ascites accumulation (left) in Ern1f/f (n = 5) or Ern1f/fCd4cre (n = 6) mice that bore ID8-Defb29/Vegfa ovarian cancer and overall survival (right, n = 6 per group). Ern1 is the gene that encodes IRE1α. Data are shown as mean ± s.e.m. (c, d, j–l). Boxes represent median ± interquartile range and whiskers indicate minimum and maximum (m, n). n values represent biologically independent mice (a–d, j–n). Two-tailed Student’s t-tests (d, j–l, m, n); log-rank test for survival (n). *P < 0.05, NS, not significant; gMFI, geometric mean fluorescence intensity.
Extended Data Fig. 8 IRE1α–XBP1 regulates IFNγ production in human CD4+ T cells exposed to ascites.
a, IFNG expression in CD4+ T cells receiving CD3 and CD28 activation for 16 h under increasing concentrations of supernatants of ovarian cancer ascites; 10% (n = 12); 50% (n = 3); 100% (n = 3). b, CD4+ T cells were activated for 12 h, incubated for additional 36 h in the presence of 25% ascites, and IFNγ in culture supernatants was determined by ELISA. Data were normalized to final viable cell counts in each case. n = 11 independent responder CD4+ T cells in all cases with the exception of A14 (n = 9), A15 (n = 10) and A17 (n = 3). c–h, Pre-activated CD4+ T cells were treated with 4μ8C for 2 h, and 25% ascites was subsequently added to the culture for additional 12 h. c, IFNγ in culture supernatants was quantified by ELISA (n = 14). Data are presented as relative expression compared with matching controls that were not exposed to ascites. d, FACS histogram (left) and quantitative analysis (right) for intracellular IFNγ in CD4+ T cells (n = 16). e, Maximal OCR presented as per cent response change compared with untreated controls (n = 7). f, The frequency of viable cells among total cells was determined by staining with dead cell exclusion dye (n = 12). The frequency of T-bet+ (g) and RORγt+ (h) populations among CD4+ T cells was determined by intracellular staining and presented as relative expression compared with controls that were not treated with ascites (n = 12). i, IFNG expression by SNAT1-overexpressing human CD4+ T cells exposed to 10% ovarian cancer ascites (n = 6 from two independent experiments) or incubated in glucose-free medium (n = 3 from two independent experiments). Data were normalized to endogenous expression of GAPDH in each sample. Data are presented as relative expression compared with control-virus-transduced T cells that were not exposed to ascites or glucose-deprived. Data are shown as mean ± s.e.m. (a–c, f–i). n values represent biologically independent samples (a–i). One-way ANOVA with Bonferroni’s multiple comparisons test (a, b); one-way ANOVA with Tukey’s multiple comparisons test (c, f–h); two-tailed paired Student’s t-tests (d, e); two-tailed Student’s t-tests (i); *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; gMFI, geometric mean fluorescence intensity.
Extended Data Fig. 9 Proposed model.
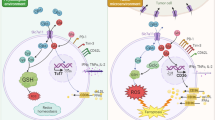
a, Under normal (glucose-rich) conditions, T cells can effectively glycosylate proteins in the ER while fuelling mitochondrial respiration through glycolysis. These processes endow T cells with competent effector function and anti-cancer capacity. b, In the tumour microenvironment, glucose availability could be limited and T cells may also express low levels of glucose transporters such as GLUT1. Glucose restriction not only dampens glycolysis, but also impairs optimal N-linked protein glycosylation in T cells, leading to ER stress and IRE1α–XBP1 activation. XBP1 controls the abundance of glutamine transporters in T cells that are experiencing ER stress, and consequently limits the influx of glutamine necessary to sustain mitochondrial respiration under glucose deprivation. Therefore, T cells become dysfunctional and incapable of controlling malignant progression. Disabling IRE1α–XBP1 signalling may be useful to enhance T cell mitochondrial function and anti-cancer capacity in a harsh tumour microenvironment.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and Supplementary Tables 1–3.
Source data
Rights and permissions
About this article
Cite this article
Song, M., Sandoval, T.A., Chae, CS. et al. IRE1α–XBP1 controls T cell function in ovarian cancer by regulating mitochondrial activity. Nature 562, 423–428 (2018). https://doi.org/10.1038/s41586-018-0597-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-018-0597-x
Keywords
This article is cited by
-
Construction of an ER stress-related prognostic signature for predicting prognosis and screening the effective anti-tumor drug in osteosarcoma
Journal of Translational Medicine (2024)
-
Emerging mechanisms of the unfolded protein response in therapeutic resistance: from chemotherapy to Immunotherapy
Cell Communication and Signaling (2024)
-
Hypoxia-activated XBP1s recruits HDAC2-EZH2 to engage epigenetic suppression of ΔNp63α expression and promote breast cancer metastasis independent of HIF1α
Cell Death & Differentiation (2024)
-
Amino acid metabolism in immune cells: essential regulators of the effector functions, and promising opportunities to enhance cancer immunotherapy
Journal of Hematology & Oncology (2023)
-
Tumor microenvironment in ovarian cancer peritoneal metastasis
Cancer Cell International (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.