Abstract
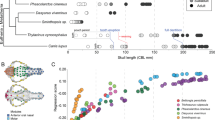
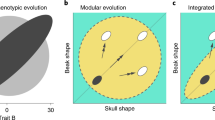
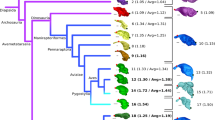
A central question in evolutionary developmental biology is how highly conserved developmental systems can generate the remarkable phenotypic diversity observed among distantly related species. In part, this paradox reflects our limited knowledge about the potential for species to both respond to selection and generate novel variation. Consequently, the developmental links between small-scale microevolutionary variations within populations to larger macroevolutionary patterns among species remain unbridged. Domesticated species, such as the pigeon, are unique resources for addressing this question, because a history of strong artificial selection has significantly increased morphological diversity, offering a direct comparison of the developmental potential of a single species to broader evolutionary patterns. Here, we demonstrate that patterns of variation and covariation within and between the face and braincase in domesticated breeds of the pigeon are predictive of avian cranial evolution. These results indicate that selection on variation generated by a conserved developmental system is sufficient to explain the evolution of crania as different in shape as the albatross or eagle, parakeet or hummingbird. These ‘rules’ of craniofacial variation are a common pattern in the evolution of a broad diversity of vertebrate species and may ultimately reflect structural limitations of a shared embryonic bauplan on functional variation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Darwin, C. The Variation of Animals and Plants under Domestication Vol. 1 (John Murray, 1868).
Darwin, C. On the Origin of Species by Means of Natural Selection (John Murray, 1859).
Stringham S. A. et al. Divergence, convergence, and the ancestry of feral populations in the domestic rock pigeon. Curr. Biol. 22, 302–308 (2012).
Shapiro, M. D. Genomic diversity and evolution of the head crest in the rock pigeon. Science 339, 1063–1067 (2013).
Domyan, E. T. & Shapiro, M. D. Pigeonetics takes flight: evolution, development, and genetics of intraspecific variation. Dev. Biol. http://dx.doi.org/10.1016/j.ydbio.2016.11.008 (in the press).
Wright, S. Genic and organismic selection. Evolution 34, 825–843 (1980).
Cheverud, J. M. Phenotypic, genetic, and environmental integration in the cranium. Evolution 36, 499–516 (1982).
Hallgrímsson, B. et al. Deciphering the palimpsest: studying the relationship between morphological integration and phenotypic covariation. Evol. Biol. 36, 355–376 (2009).
Cheverud, J. M. Developmental integration and the evolution of pleiotropy. Am. Zool. 36, 44–50 (1996).
Wagner G. P. & Altenberg, L. Complex adaptations and the evolution of evolvability. Evolution 50, 967–976 (1996).
Maynard Smith J. et al. Developmental constraints and evolution. Q. Rev. Biol. 60, 265–287 (1985).
Wagner, G. P., Pavlicev, M. & Cheverud, J. The road to modularity. Nat. Rev. Genet. 8, 921–931 (2007).
Johnston, R. F. Evolution in the rock dove: skeletal morphology. Auk 109, 530–542 (1992).
Baptista, L. F., Gomez Martinez, J. E. & Horblit, H. M. Darwin’s pigeons and the evolution of the columbiforms: recapitulation of ancient genes. Acta Zool. Mex. 25, 719–741 (2009).
Abzhanov, A. et al. Bmp4 and morphological variation of beaks in Darwin’s finches. Science 305, 1462–1465 (2004).
Wu, P. et al. Molecular shaping of the beak. Science 305, 1465–1466 (2004).
Abzhanov, A. et al. The calmodulin pathway and evolution of elongated beak morphology in Darwin’s finches. Nature 442, 563–567 (2006).
Wu, P. et al. Morphoregulation of avian beaks: comparative mapping of growth zone activities and morphological evolution. Dev. Dynam. 235, 1400–1412 (2006).
Fritz, J. A. et al. Shared developmental programme strongly constrains beak shape diversity in songbirds. Nat. Commun. 5, 3700 (2015).
Linde-Medina, M. & Newman, S. A. Limb, tooth, beak: three modes of development and evolutionary innovation of form. J. Bioscience 39, 211–223 (2014).
Mallarino, R. et al. Two developmental modules establish 3D beak-shape variation in Darwin’s finches. Proc. Natl Acad. Sci. USA 108, 4057–4062 (2011).
Bright, J. A. et al. The shapes of bird beaks are highly controlled by nondietary factors. Proc. Natl Acad. Sci. USA 113, 5352–5357 (2016).
Drake, A. G. & Klingenberg, C. P. Large-scale diversification of skull shape in domestic dogs: disparity and modularity. Am. Nat. 175, 289–301 (2010).
Martínez-Abadías, N. et al. Pervasive genetic integration directs the evolution of human skull shape. Evolution 66, 1010–1023 (2010).
Young, N. M. et al. Facial surface morphology predicts variation in internal skeletal shape. Am. J. Orthod. Dent. Orthop. 149, 501–508 (2016).
Porto, A. et al. The evolution of modularity in the mammalian skull I: Morphological integration patterns and magnitudes. Evol. Biol. 36, 118–135 (2009).
Young, N. M. et al. Embryonic bauplans and the developmental origins of facial diversity and constraint. Development 141, 1059–1063 (2014).
Marcucio, R. S., Young, N. M., Hu, D. & Hallgrímsson, B. Mechanisms that underlie co-variation of the brain and face. Genesis 49, 177–189 (2011).
Warton, D. I. et al. smatr 3—an R package for estimation and inference about allometric lines. Methods Ecol. Evol. 3, 257–259 (2012).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2016); https://www.R-project.org
Marroig, G. & Cheverud, J. M. A comparison of phenotypic variation and covariation patterns and the role of phylogeny, ecology, and ontogeny during cranial evolution of new world monkeys. Evolution 55, 2576–2600 (2001).
Wagner, G. P. On the eigenvalue distribution of genetic and phenotypic dispersion matrices: Evidence for a non-random origin of quantitative genetic variation. J. Math. Biol. 21, 77–95 (1984).
Pavlicev, M., Cheverud, J. M. & Wagner, G. P. Measuring morphological integration using eigenvalue variance. Evol. Biol. 36, 157–170 (2009).
Young, N. M., Wagner, G. P. & Hallgrímsson, B. Development and the evolvability of human limbs. Proc. Natl Acad. Sci. USA 107, 3400–3405 (2010).
Zelditch, M. L., Swiderski, D. L. & Sheets, H. D. Geometric Morphometrics for Biologists: A Primer (Academic, 2012).
Nealen, P. M. & Ricklefs, R. Early diversification of the avian brain:body relationship. J. Zool. 253, 391–404 (2001).
Klingenberg, C. P. Morphometric integration and modularity in configurations of landmarks: tools for evaluating a priori hypotheses. Evol. Dev. 11, 405–421 (2009).
Adams, D. C. Evaluating modularity in morphometric data: challenges with the RV coefficient and a new test measure. Methods Ecol. Evol. 7, 565–572 (2016).
Adams, D. C. & Otarola-Castillo, E. geomorph: an R package for the collection and analysis of geometric morphometric shape data. Methods Ecol. Evol. 4, 393–399 (2013).
Bookstein, F. L. Integration, disintegration, and self-similarity: characterizing the scales of shape variation in landmark data. Evol. Biol. 42, 395–426 (2015).
Jetz, W. et al. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Maddison, W. P. & Maddison, D. R. Mesquite: A Modular System for Evolutionary Analysis Version 3.10 (accessed 7 January 2016); http://mesquiteproject.org
Klingenberg, C. P. & Gidaszewski, N. A. Testing and quantifying phylogenetic signals and homoplasy in morphometric data. Syst. Biol. 59, 245–261 (2010).
Blomberg, S. P., Garland, T. & Ives, A. R. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57, 717–745 (2003).
Adams, D. C. A generalized Κ statistic for estimating phylogenetic signal from shape and other high-dimensional multivariate data. Syst. Biol. 63, 685–697 (2014).
Sakamoto, M. & Ruta, M. Convergence and divergence in the evolution of cat skulls: Temporal and spatial patterns of morphological diversity. PLoS ONE 7, e39752 (2012).
Klingenberg, C. P. MorphoJ: an integrated software package for geometric morphometrics. Mol. Ecol. Resour. 11, 353–357 (2011).
Acknowledgements
We thank J. DeCarlo for kindly donating specimens of domestic pigeon breeds, A. Goode for preparing their skeletons and R. Johnston for generously sharing his original pigeon data. Non-pigeon avian CT data were provided courtesy of the University of Texas High Resolution X-ray CT Facility (UTCT) (National Science Foundation grant number IIS-0208675). Research reported in this publication was supported by the National Institute of Dental and Craniofacial Research of the National Institutes of Health under Award Numbers F32DE018596 (to N.M.Y.), R01DE019638 (to R.S.M. and B.H.) and R01DE021708 (to R.S.M. and B.H.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
N.M.Y. designed the research. N.M.Y. and J.W.F. collected pigeon specimens. N.M.Y. and M.L.-M. performed the analyses. N.M.Y., M.L.-M., B.H., and R.S.M. contributed to the interpretation of the results. N.M.Y. drafted the paper. All authors contributed to the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Tables 1–7, Supplementary Figures 1–10. (PDF 2647 kb)
Rights and permissions
About this article
Cite this article
Young, N., Linde-Medina, M., Fondon, J. et al. Craniofacial diversification in the domestic pigeon and the evolution of the avian skull. Nat Ecol Evol 1, 0095 (2017). https://doi.org/10.1038/s41559-017-0095
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-017-0095
This article is cited by
-
Comparative valuation of the chlorpyrifos, acetamiprid, and lambda-cyhalothrin toxicity and their hematological and histopathological consequences in pigeons
Environmental Science and Pollution Research (2023)
-
MusMorph, a database of standardized mouse morphology data for morphometric meta-analyses
Scientific Data (2022)
-
Cranial shape diversification in horses: variation and covariation patterns under the impact of artificial selection
BMC Ecology and Evolution (2021)
-
Patterns of skeletal integration in birds reveal that adaptation of element shapes enables coordinated evolution between anatomical modules
Nature Ecology & Evolution (2021)
-
Digital dissection of the head of the rock dove (Columba livia) using contrast-enhanced computed tomography
Zoological Letters (2019)