Abstract
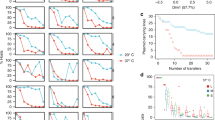
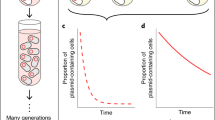
Plasmids are thought to play a key role in bacterial evolution by acting as vehicles for horizontal gene transfer, but the role of plasmids as catalysts of gene evolution remains unexplored. We challenged populations of Escherichia coli carrying the blaTEM-1 β-lactamase gene on either the chromosome or a multicopy plasmid (19 copies per cell) with increasing concentrations of ceftazidime. The plasmid accelerated resistance evolution by increasing the rate of appearance of novel TEM-1 mutations, thereby conferring resistance to ceftazidime, and then by amplifying the effect of TEM-1 mutations due to the increased gene dosage. Crucially, this dual effect was necessary and sufficient for the evolution of clinically relevant levels of resistance. Subsequent evolution occurred by mutations in a regulatory RNA that increased the plasmid copy number, resulting in marginal gains in ceftazidime resistance. These results uncover a role for multicopy plasmids as catalysts for the evolution of antibiotic resistance in bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Wiedenbeck, J. & Cohan, F. M. Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol. Rev. 35, 957–976 (2011).
Baltrus, D. A. Exploring the costs of horizontal gene transfer. Trends Ecol. Evol. 28, 489–495 (2013).
Vogwill, T. & MacLean, R. C. The genetic basis of the fitness costs of antimicrobial resistance: a meta-analysis approach. Evol. Appl. 8, 284–295 (2014).
Stewart, F. M. & Levin, B. R. The population biology of bacterial plasmids: a priori conditions for the existence of conjugationally transmitted factors. Genetics 87, 209–228 (1977).
Levin, B. R. & Stewart, F. M. The population biology of bacterial plasmids: a priori conditions for the existence of mobilizable nonconjugative factors. Genetics 94, 425–443 (1980).
Bergstrom, C. T., Lipsitch, M. & Levin, B. R. Natural selection, infectious transfer and the existence conditions for bacterial plasmids. Genetics 155, 1505–1519 (2000).
Harrison, E. & Brockhurst, M. A. Plasmid-mediated horizontal gene transfer is a coevolutionary process. Trends Microbiol. 20, 262–267 (2012).
San Millan, A. et al. Positive selection and compensatory adaptation interact to stabilize non-transmissible plasmids. Nat. Commun. 5, 5208 (2014).
Harrison, E., Guymer, D., Spiers, A. J., Paterson, S. & Brockhurst, M. A. Parallel compensatory evolution stabilizes plasmids across the parasitism–mutualism continuum. Curr. Biol. 25, 2034–2039 (2015).
Peña-Miller, R., Rodríguez-González, R., MacLean, R. C. & San Millan, A. Evaluating the effect of horizontal transmission on the stability of plasmids under different selection regimes. Mob. Genet. Elements 22, 1–5 (2015).
Modi, R. I., Castilla, L. H., Puskas-Rozsa, S., Helling, R. B. & Adams, J. Genetic changes accompanying increased fitness in evolving populations of Escherichia coli . Genetics 130, 241–249 (1992).
Stoesser, N. et al. Evolutionary history of the global emergence of the Escherichia coli epidemic clone ST131. MBio 7, e02162 (2016).
Latorre, A., Gil, R., Silva, F. J. & Moya, A. Chromosomal stasis versus plasmid plasticity in aphid endosymbiont Buchnera aphidicola . Heredity 95, 339–347 (2005).
Zhang, Z., Qian, W. & Zhang, J. Positive selection for elevated gene expression noise in yeast. Mol. Syst. Biol. 5, 299 (2009).
Martinez, J. L. & Baquero, F. Mutation frequencies and antibiotic resistance. Antimicrob. Agents Chemother. 44, 1771–1777 (2000).
Couce, A., Rodriguez-Rojas, A. & Blazquez, J. Bypass of genetic constraints during mutator evolution to antibiotic resistance. Proc. Biol. Sci. 282, 20142698 (2015).
Sano, E., Maisnier-Patin, S., Aboubechara, J. P., Quinones-Soto, S. & Roth, J. R. Plasmid copy number underlies adaptive mutability in bacteria. Genetics 198, 919–933 (2014).
Hendrickson, H., Slechta, E. S., Bergthorsson, U., Andersson, D. I. & Roth, J. R. Amplification-mutagenesis: evidence that "directed" adaptive mutation and general hypermutability result from growth with a selected gene amplification. Proc. Natl Acad. Sci. USA 99, 2164–2169 (2002).
Sarno, R., McGillivary, G., Sherratt, D. J., Actis, L. A. & Tolmasky, M. E. Complete nucleotide sequence of Klebsiella pneumoniae multiresistance plasmid pJHCMW1. Antimicrob. Agents Chemother. 46, 3422–3427 (2002).
Søndergaard, A. et al. Molecular organization of small plasmids bearing blaTEM-1 and conferring resistance to beta-lactams in Haemophilus influenzae . Antimicrob. Agents Chemother. 56, 4958–4960 (2012).
Vignoli, R. et al. New TEM-derived extended-spectrum beta-lactamase and its genomic context in plasmids from Salmonella enterica serovar derby isolates from Uruguay. Antimicrob. Agents Chemother. 50, 781–784 (2006).
Bershtein, S., Segal, M., Bekerman, R., Tokuriki, N. & Tawfik, D. S. Robustness–epistasis link shapes the fitness landscape of a randomly drifting protein. Nature 444, 929–932 (2006).
Stiffler, M. A., Hekstra, D. R. & Ranganathan, R. Evolvability as a function of purifying selection in TEM-1 beta-lactamase. Cell 160, 882–892 (2015).
Schenk, M. F., Szendro, I. G., Krug, J. & de Visser, J. A. Quantifying the adaptive potential of an antibiotic resistance enzyme. PLoS Genet. 8, e1002783 (2012).
San Millan, A. et al. Small-plasmid-mediated antibiotic resistance is enhanced by increases in plasmid copy number and bacterial fitness. Antimicrob. Agents Chemother. 59, 3335–3341 (2015).
San Millan, A., Heilbron, K. & MacLean, R. C. Positive epistasis between co-infecting plasmids promotes plasmid survival in bacterial populations. ISME J. 8, 601–612 (2014).
Reguera, J. A., Baquero, F., Perez-Diaz, J. C. & Martinez, J. L. Synergistic effect of dosage and bacterial inoculum in TEM-1 mediated antibiotic resistance. Eur. J. Clin. Microbiol. Infect. Dis. 7, 778–779 (1988).
Martinez, J. L. et al. Resistance to beta-lactam/clavulanate. Lancet 2, 1473 (1987).
Artemova, T., Gerardin, Y., Dudley, C., Vega, N. M. & Gore, J. Isolated cell behavior drives the evolution of antibiotic resistance. Mol. Syst. Biol. 11, 822 (2015).
Gullberg, E., Albrecht, L. M., Karlsson, C., Sandegren, L. & Andersson, D. I. Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. MBio 5, e01918–14 (2014).
Salverda, M. L., De Visser, J. A. & Barlow, M. Natural evolution of TEM-1 β-lactamase: experimental reconstruction and clinical relevance. FEMS Microbiol. Rev. 34, 1015–1036 (2010).
Tavio, M. M., Aquili, V. D., Vila, J. & Poveda, J. B. Resistance to ceftazidime in Escherichia coli associated with AcrR, MarR and PBP3 mutations and overexpression of sdiA. J. Med. Microbiol. 63, 56–65 (2014).
Hirakawa, H., Nishino, K., Yamada, J., Hirata, T. & Yamaguchi, A. Beta-lactam resistance modulated by the overexpression of response regulators of two-component signal transduction systems in Escherichia coli . J. Antimicrob. Chemother. 52, 576–582 (2003).
Cai, S. J. & Inouye, M. EnvZ-OmpR interaction and osmoregulation in Escherichia coli . J. Biol. Chem. 277, 24155–24161 (2002).
Jacquier, H. et al. In vivo selection of a complex mutant TEM (CMT) from an inhibitor-resistant TEM (IRT) during ceftazidime therapy. J. Antimicrob. Chemother. 68, 2792–2796 (2013).
Negri, M. C., Lipsitch, M., Blazquez, J., Levin, B. R. & Baquero, F. Concentration-dependent selection of small phenotypic differences in TEM beta-lactamase-mediated antibiotic resistance. Antimicrob. Agents Chemother. 44, 2485–2491 (2000).
Nasvall, J., Sun, L., Roth, J. R. & Andersson, D. I. Real-time evolution of new genes by innovation, amplification, and divergence. Science 338, 384–387 (2012).
Sandegren, L. & Andersson, D. I. Bacterial gene amplification: implications for the evolution of antibiotic resistance. Nat. Rev. Microbiol. 7, 578–588 (2009).
Tomizawa, J., Itoh, T., Selzer, G. & Som, T. Inhibition of ColE1 RNA primer formation by a plasmid-specified small RNA. Proc. Natl Acad. Sci. USA 78, 1421–1425 (1981).
Tomizawa, J. Control of ColE1 plasmid replication: the process of binding of RNA I to the primer transcript. Cell 38, 861–870 (1984).
Lacatena, R. M. & Cesareni, G. Interaction between RNA1 and the primer precursor in the regulation of Co1E1 replication. J. Mol. Biol. 170, 635–650 (1983).
Lacatena, R. M. & Cesareni, G. Base pairing of RNA I with its complementary sequence in the primer precursor inhibits ColE1 replication. Nature 294, 623–626 (1981).
Breakpoint Tables for Interpretation of MICs and Zone Diameters v. 6.0. (The European Committee on Antimicrobial Susceptibility Testing, 2016); http://www.eucast.org
Wang, X., Minasov, G. & Shoichet, B. K. Evolution of an antibiotic resistance enzyme constrained by stability and activity trade-offs. J. Mol. Biol. 320, 85–95 (2002).
Ohno, S. Evolution by Gene Duplication (Springer, 1970).
Toll-Riera, M., San Millan, A., Wagner, A. & MacLean, R. C. The genomic basis of evolutionary innovation in Pseudomonas aeruginosa. PLoS Genet. 12, e1006005 (2016).
Andersson, D. I. & Hughes, D. Gene amplification and adaptive evolution in bacteria. Annu. Rev. Genet. 43, 167–195 (2009).
Lindsey, H. A., Gallie, J., Taylor, S. & Kerr, B. Evolutionary rescue from extinction is contingent on a lower rate of environmental change. Nature 494, 463–467 (2013).
Baquero, F. & Negri, M. C. Selective compartments for resistant microorganisms in antibiotic gradients. Bioessays 19, 731–736 (1997).
Performance standards for antimicrobial susceptibility testing. 19th ed. (Clinical and Laboratory Standards Institute, 2009).
Escudero, J. A. et al. Unmasking the ancestral activity of integron integrases reveals a smooth evolutionary transition during functional innovation. Nat. Commun. 7, 10937 (2016).
Demarre, G., Frumerie, C., Gopaul, D. N. & Mazel, D. Identification of key structural determinants of the IntI1 integron integrase that influence attC x attI1 recombination efficiency. Nucleic Acids Res. 35, 6475–6489 (2007).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Chaveroche, M. K., Ghigo, J. M. & d’Enfert, C. A rapid method for efficient gene replacement in the filamentous fungus Aspergillus nidulans. Nucleic Acids Res. 28, E97 (2000).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Le Roux, F., Binesse, J., Saulnier, D. & Mazel, D. Construction of a Vibrio splendidus mutant lacking the metalloprotease gene vsm by use of a novel counterselectable suicide vector. Appl. Environ. Microbiol. 73, 777–784 (2007).
Lee, C., Kim, J., Shin, S. G. & Hwang, S. Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli . J. Biotechnol. 123, 273–280 (2006).
Deatherage, D. E. & Barrick, J. E. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–1882 (2014).
Barrick, J. E. et al. Identifying structural variation in haploid microbial genomes from short-read resequencing data using breseq. BMC Genomics 15, 1039 (2014).
Deatherage, D. E., Traverse, C. C., Wolf, L. N. & Barrick, J. E. Detecting rare structural variation in evolving microbial populations from new sequence junctions using breseq. Front. Genet. 5, 468 (2014).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Acknowledgements
A.S.M. thanks E. Frago for his help with the statistical analyses. This work was supported by funding from the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013)/ERC grant (StG-2011-281591). A.S.M. is supported by a Miguel Servet fellowship from the Instituto de Salud Carlos III (MS15/00012) co-financed by the European Social Fund and The European Development Regional Fund ‘A way to achieve Europe’ (ERDF). J.A.E. is supported by a Marie Curie Intra-European Fellowship for Career Development (FP-7-PEOPLE-2011-IEF, ICADIGE). Work in the Mazel lab was supported by the Institut Pasteur, the Centre National de la Recherche Scientifique (CNRS-UMR3525) and the European Union Seventh Framework Programme (FP7-HEALTH- 2011-single-stage), the ‘Evolution and Transfer of Antibiotic Resistance’ (EvoTAR, FP7- HEALTH-282004).
We thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics funded by Wellcome Trust grant reference 090532/Z/09/Z and Medical Research Council Hub grant G0900747 91070 for generation of the high-throughput sequencing data.
Author information
Authors and Affiliations
Contributions
A.S.M. and R.C.M. were responsible for the conceptualization of this study; A.S.M., J.A.E., and D.R.G. designed the methodology; formal analysis was by D.R.G.; A.S.M. and J.A.E. carried out the investigations; A.S.M. and R.C.M. prepared the original draft of the manuscript and also undertook the reviewing and editing; R.C.M. and D.M. were responsible for funding acquisition and for supervision; D.M. oversaw the resources; project administration was performed by A.S.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1,2, Supplementary Tables 1–3 and Supplementary References. (PDF 214 kb)
Supplementary Data 1
Description of the mutations affecting the chromosome and pBGT in the different populations and clones of MG, MG::blaTEM-1 and MG/pBGT analysed in this work. The position and nature of the mutations are indicated, as well as the genes affected and their activity. The frequencies of the mutations in the different populations and clones are also indicated. (XLSX 70 kb)
Rights and permissions
About this article
Cite this article
San Millan, A., Escudero, J., Gifford, D. et al. Multicopy plasmids potentiate the evolution of antibiotic resistance in bacteria. Nat Ecol Evol 1, 0010 (2017). https://doi.org/10.1038/s41559-016-0010
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41559-016-0010
This article is cited by
-
Plasmid-mediated phenotypic noise leads to transient antibiotic resistance in bacteria
Nature Communications (2024)
-
Phage-plasmids promote recombination and emergence of phages and plasmids
Nature Communications (2024)
-
Profile and resistance levels of 136 integron resistance genes
npj Antimicrobials and Resistance (2023)
-
Regulatory fine-tuning of mcr-1 increases bacterial fitness and stabilises antibiotic resistance in agricultural settings
The ISME Journal (2023)
-
Cooperative antibiotic resistance facilitates horizontal gene transfer
The ISME Journal (2023)