Abstract
Background
Small bowel adenocarcinoma is a rare disease. The genomic profiling tumours according to clinical characteristics and its impact on the prognosis remains unclear.
Methods
A pooled analysis of clinical data, genomic profiling and MisMatch Repair (MMR) status from three databases was performed.
Results
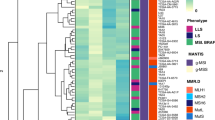
A total of 188 tumour samples were analysed. A predisposing disease was reported in 22.3%, mainly Lynch syndrome and Crohn’s disease. The tumours were localized in 80.2% and metastatic in 18.8%. The most frequent mutations were KRAS (42.0%) among them 7/79 are G12C, TP53 (40.4%), APC (19.1%), PIK3CA (18.6%), SMAD4 (12.8%) and ERBB2 (9.6%). Mutation distribution differed according to predisposing disease for TP53, ERBB2, IDH1, FGFR3, FGFR1 and KDR. KRAS and SMAD4 mutations were more frequent in metastatic tumour, whereas ERBB2 mutations were absent in metastatic tumour. For localized tumour, APC mutation was independently associated with a poor overall survival (OS) (p = 0.0254). 31.8% of localized tumours and 11.3% of metastatic tumours were dMMR (29.8% of the entire cohort). A dMMR status was associated with a better OS (HR = 0.61 [0.39–0.96], p = 0.0316).
Conclusions
There is a different genomic profile according to the stage and predisposing disease. dMMR and APC mutation in localized tumour predict a better prognosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Aparicio T, Pachev A, Laurent-Puig P, Svrcek M. Epidemiology, Risk Factors and Diagnosis of Small Bowel Adenocarcinoma. Cancers. 2022;14:2268.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Bouvier AM, Robaszkiewicz M, Jooste V, Cariou M, Drouillard A, Bouvier V, et al. Trends in incidence of small bowel cancer according to histology: a population-based study. J Gastroenterol. 2020;55:181–8.
Legué LM, Bernards N, Gerritse SL, van Oudheusden TR, de Hingh IHJT, Creemers GJM, et al. Trends in incidence, treatment and survival of small bowel adenocarcinomas between 1999 and 2013: a population-based study in The Netherlands. Acta Oncol Stock Swed. 2016;55:1183–9.
Aparicio T, Henriques J, Manfredi S, Tougeron D, Bouché O, Pezet D, et al. Small bowel adenocarcinoma: Results from a nationwide prospective ARCAD-NADEGE cohort study of 347 patients. Int J Cancer. 2020;147:967–77.
Haan JC, Buffart TE, Eijk PP, van de Wiel MA, van Wieringen WN, Howdle PD, et al. Small bowel adenocarcinoma copy number profiles are more closely related to colorectal than to gastric cancers. Ann Oncol J Eur Soc Med Oncol. 2012;23:367–74.
Schrock AB, Devoe CE, McWilliams R, Sun J, Aparicio T, Stephens PJ, et al. Genomic Profiling of Small-Bowel Adenocarcinoma. JAMA Oncol. 2017;3:1546–53.
Laforest A, Aparicio T, Zaanan A, Silva FP, Didelot A, Desbeaux A, et al. ERBB2 gene as a potential therapeutic target in small bowel adenocarcinoma. Eur J Cancer Oxf Engl 1990. 2014;50:1740–6.
Hänninen UA, Katainen R, Tanskanen T, Plaketti RM, Laine R, Hamberg J, et al. Exome-wide somatic mutation characterization of small bowel adenocarcinoma. PLoS Genet. 2018;14:e1007200.
Aparicio T, Svrcek M, Henriques J, Afchain P, Lièvre A, Tougeron D, et al. Panel gene profiling of small bowel adenocarcinoma: Results from the NADEGE prospective cohort. Int J Cancer. 2021;148:1731–42.
Adam L, San Lucas FA, Fowler R, Yu Y, Wu W, Liu Y, et al. DNA Sequencing of Small Bowel Adenocarcinomas Identifies Targetable Recurrent Mutations in the ERBB2 Signaling Pathway. Clin Cancer Res J Am Assoc Cancer Res. 2019;25:641–51.
Casadei-Gardini A, Lonardi S, Smiroldo V, Canale M, Passardi A, Silvestris N, et al. Extensive molecular reclassification: new perspectives in small bowel adenocarcinoma? Med Oncol Northwood Lond Engl. 2021;38:17.
Marabelle A, Le DT, Ascierto PA, Di Giacomo AM, De Jesus-Acosta A, Delord JP, et al. Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair-Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. J Clin Oncol J Am Soc Clin Oncol. 2020;38:1–10.
Aparicio T, Svrcek M, Zaanan A, Beohou E, Laforest A, Afchain P, et al. Small bowel adenocarcinoma phenotyping, a clinicobiological prognostic study. Br J Cancer. 2013;109:3057–66.
Vanoli A, Grillo F, Guerini C, Neri G, Arpa G, Klersy C, et al. Prognostic Role of Mismatch Repair Status, Histotype and High-Risk Pathologic Features in Stage II Small Bowel Adenocarcinomas. Ann Surg Oncol. 2021;28:1167–77.
Jun SY, Kim M, Jin Gu M, Kyung Bae Y, Chang HK, Sun Jung E, et al. Clinicopathologic and prognostic associations of KRAS and BRAF mutations in small intestinal adenocarcinoma. Mod Pathol J U S Can Acad Pathol Inc. 2016;29:402–15.
Alvi MA, McArt DG, Kelly P, Fuchs MA, Alderdice M, McCabe CM, et al. Comprehensive molecular pathology analysis of small bowel adenocarcinoma reveals novel targets with potential for clinical utility. Oncotarget. 2015;6:20863–74.
Pandya K, Overman MJ, Gulhati P. Molecular Landscape of Small Bowel Adenocarcinoma. Cancers. 2022;14:1287.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med J Am Coll Med Genet. 2015;17:405–24.
Hammoudi N, Lehmann-Che J, Lambert J, Amoyel M, Maggiori L, Salfati D, et al. Prognosis and molecular characteristics of IBD-associated colorectal cancer: Experience from a French tertiary-care center. Dig Liver Dis J Ital Soc Gastroenterol Ital Assoc Study Liver. 2023;55:1280–7. S1590-8658(23)00225-6
Yaeger R, Weiss J, Pelster MS, Spira AI, Barve M, Ou SHI, et al. Adagrasib with or without Cetuximab in Colorectal Cancer with Mutated KRAS G12C. N. Engl J Med. 2023;388:44–54.
Strickler JH, Satake H, George TJ, Yaeger R, Hollebecque A, Garrido-Laguna I, et al. Sotorasib in KRAS p.G12C-Mutated Advanced Pancreatic Cancer. N. Engl J Med. 2023;388:33–43.
Kopetz S, Grothey A, Yaeger R, Van Cutsem E, Desai J, Yoshino T, et al. Encorafenib, Binimetinib, and Cetuximab in BRAF V600E-Mutated Colorectal Cancer. N. Engl J Med. 2019;381:1632–43.
Yaeger R, Shah MA, Miller VA, Kelsen JR, Wang K, Heins ZJ, et al. Genomic Alterations Observed in Colitis-Associated Cancers Are Distinct From Those Found in Sporadic Colorectal Cancers and Vary by Type of Inflammatory Bowel Disease. Gastroenterology. 2016;151:278–87.
Jorissen RN, Christie M, Mouradov D, Sakthianandeswaren A, Li S, Love C, et al. Wild-type APC predicts poor prognosis in microsatellite-stable proximal colon cancer. Br J Cancer. 2015;113:979–88.
Mondaca S, Walch H, Nandakumar S, Chatila WK, Schultz N, Yaeger R. Specific Mutations in APC, but Not Alterations in DNA Damage Response, Associate With Outcomes of Patients With Metastatic Colorectal Cancer. Gastroenterology. 2020;159:1975–8.e4.
Liao X, Li G, McBride R, Houldsworth J, Harpaz N, Polydorides AD. Clinicopathological and Molecular Characterisation of Crohn’s Disease-associated Small Bowel Adenocarcinomas. J Crohns Colitis. 2020;14:287–94.
Donahu TF, Bagrodia A, Audenet F, Donoghue MTA, Cha EK, Sfakianos JP, et al. Genomic Characterization of Upper-Tract Urothelial Carcinoma in Patients With Lynch Syndrome. JCO Precis Oncol [Internet]. 2018 [cité 20 avr 2020];2018. Disponible sur: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6404976/
Kondelin J, Salokas K, Saarinen L, Ovaska K, Rauanheimo H, Plaketti RM, et al. Comprehensive evaluation of coding region point mutations in microsatellite-unstable colorectal cancer. EMBO Mol Med. 2018;10:e8552.
Boyer C, Sefrioui D, Cohen R, Chautard R, Perrier M, Lebrun H, et al. Prognosis and chemosensitivity of non-colorectal alimentary tract cancers with microsatellite instability. Dig Liver Dis J Ital Soc Gastroenterol Ital Assoc Study Liver. 2023;55:123–30.
Coutzac C, Bibeau F, Ben Abdelghani M, Aparicio T, Cohen R, Coquan E, et al. Immunotherapy in MSI/dMMR tumors in the perioperative setting: The IMHOTEP trial. Dig Liver Dis J Ital Soc Gastroenterol Ital Assoc Study Liver. 2022;54:1335–41.
André T, Tougeron D, Piessen G, de la Fouchardière C, Louvet C, Adenis A, et al. Neoadjuvant Nivolumab Plus Ipilimumab and Adjuvant Nivolumab in Localized Deficient Mismatch Repair/Microsatellite Instability-High Gastric or Esophagogastric Junction Adenocarcinoma: The GERCOR NEONIPIGA Phase II Study. J Clin Oncol J Am Soc Clin Oncol. 2023;41:255–65.
Chalabi M, Fanchi LF, Dijkstra KK, Van den Berg JG, Aalbers AG, Sikorska K, et al. Neoadjuvant immunotherapy leads to pathological responses in MMR-proficient and MMR-deficient early-stage colon cancers. Nat Med. 2020;26:566–76.
Bennett CM, Coleman HG, Veal PG, Cantwell MM, Lau CCL, Murray LJ. Lifestyle factors and small intestine adenocarcinoma risk: A systematic review and meta-analysis. Cancer Epidemiol. 2015;39:265–73.
Ewans J, Aparicio T, Le Malicot K, Nakamura K, Homma Y, Mc Williams R. GLOBAL BALLAD: An International Rare Cancers Initiative trial to evaluate the potential benefit of adjuvant chemotherapy for small bowel adenocarcinoma (IRCI 002). In Chicago: J Clin Oncol; 2016. p. TPS4154.
Acknowledgements
BIONADEGE and AGEO study were granted by INCa and sponsor by Assistance Publique Hôpitaux de Paris (Délégation à la Recherche Clinique). ARCAD-NADEGE cohort was granted by A.R.C.A.D. and sponsored by GERCOR.
Funding
BIONADEGE was supported by a grant from INCa (Programme Hospitalier de Recherche Translationelle Cancer, PRTK14 N°091) and a grant n° NA 2009 from the A.R.CA.D. foundation. AGEO study was supported by a grant from INCa (Programme Hospitalier de Recherche Clinique, PHRC AOM 09204).
Author information
Authors and Affiliations
Contributions
Study design: TA, MS, JH, DV and PLP. Data acquisition: TA, MS, AZ, SM, ACG, DT, JMG, DP, ET, GP, PA, CL, MP, TL, VB, FDF, SLD, SC, PLP. Statistical analysis: JH and DV. Manuscript preparation: TA, MS, JH, DV and PLP. Manuscript review: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study was performed in accordance with the Declaration of Helsinki and was authorised by the ethics committee “Ile de France II” No. ID-RCB: 2008-A01058-47 for the French part and by the local Area Vasta Emilia Nord Ethics committee for the Italian part.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aparicio, T., Henriques, J., Svrcek, M. et al. Genomic profiling of small bowel adenocarcinoma: a pooled analysis from 3 databases. Br J Cancer (2024). https://doi.org/10.1038/s41416-024-02687-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41416-024-02687-7