Abstract
Background
Increased maternal interleukin (IL)-17A and activated microglia are pivotal factors contributing to the pathological phenotypes of maternal immune activation (MIA), developing neurodevelopmental disorders in offspring. This study aimed to determine whether IL-17A affects the microglial microRNA (miRNA) profiles.
Methods
The miRNA expression profiles of primary cultured microglia stimulated with recombinant IL-17A were examined comprehensively using miRNA sequencing and validated through qRT-PCR. The expressions of miRNAs target genes identified using bioinformatics, were investigated in microglia transfected with mimic miRNA. The target gene’s expression was also examined in the fetal brains of the MIA mouse model induced by maternal lipopolysaccharide (LPS) administration.
Results
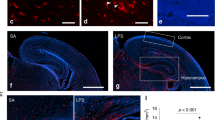
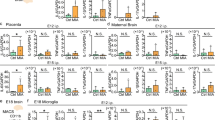
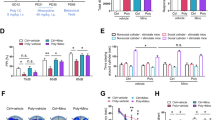
Primary cultured microglia expressed the IL-17A receptor and increased proinflammatory cytokines and nitric oxide synthase 2 upon treatment with IL-17A. Among the three miRNAs with |log2FC | >1, only mmu-miR-206-3p expression was significantly up-regulated by IL-17A. Transfection with the mmu-miR-206-3p mimic resulted in a significant decrease in the expression of Hdac4 and Igf1, target genes of mmu-miR-206-3p. Hdac4 expression also significantly decreased in the LPS-induced MIA model.
Conclusions
IL-17A affected microglial miRNA profiles with upregulated mmu-miR-206-3p. These findings suggest that targeting the IL-17A/mmu-miR-206-3p pathway may be a new strategy for predicting MIA-related neurodevelopmental deficits and providing preventive interventions.
Impact
-
Despite the growing evidence of interleukin (IL)-17A and microglia in the pathology of maternal immune activation (MIA), the downstream of IL-17A in microglia is not fully known.
-
IL-17A altered microRNA profiles and upregulated the mmu-miR-206-3p expression in microglia. The mmu-miR-206-3p reduced autism spectrum disorder (ASD) related gene expressions, Hdac4 and Igf1.
-
The Hdac4 expression was also reduced in the brain of MIA offspring.
-
The hsa-miR-206 sequence is consistent with that of mmu-miR-206-3p.
-
This study may provide clues to pathological mechanisms leading to predictions and interventions for ASD children born to mothers with IL-17A-related disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated and analysed during the current study are available in the DDBJ Sequence Read Archive (DRA015999). All other relevant data are available within the article file from the authors upon reasonable request.
References
Estes, M. L. & McAllister, A. K. Maternal immune activation: Implications for neuropsychiatric disorders. Science 353, 772–777 (2016).
Knuesel, I. et al. Maternal immune activation and abnormal brain development across Cns disorders. Nat. Rev. Neurol. 10, 643–660 (2014).
Taylor, M. J. et al. Etiology of autism spectrum disorders and autistic traits over time. JAMA Psychiatry 77, 936–943 (2020).
Wong, H. & Hoeffer, C. Maternal Il-17a in autism. Exp. Neurol. 299, 228–240 (2018).
Choi, G. B. et al. The maternal interleukin-17a pathway in mice promotes autism-like phenotypes in offspring. Science 351, 933–939 (2016).
Imai, K. et al. Neuroprotective potential of molecular hydrogen against perinatal brain injury via suppression of activated microglia. Free Radic. Biol. Med. 91, 154–163 (2016).
Imai, K. et al. Administration of molecular hydrogen during pregnancy improves behavioral abnormalities of offspring in a maternal immune activation model. Sci. Rep. 8, 9221 (2018).
Otero, A. M. & Antonson, A. M. At the crux of maternal immune activation: Viruses, microglia, microbes, and Il-17a. Immunol. Rev. 311, 205–223 (2022).
Das Sarma, J. et al. Functional interleukin-17 receptor a is expressed in central nervous system glia and upregulated in experimental autoimmune encephalomyelitis. J. Neuroinflammation 6, 14 (2009).
Kawanokuchi, J. et al. Production and functions of Il-17 in Microglia. J. Neuroimmunol. 194, 54–61 (2008).
Yu, D. et al. Microglial Gpr56 is the molecular target of maternal immune activation-induced parvalbumin-positive interneuron deficits. Sci. Adv. 8, eabm2545 (2022).
O’Connell, R. M., Rao, D. S. & Baltimore, D. Microrna regulation of inflammatory responses. Annu Rev. Immunol. 30, 295–312 (2012).
Komada, M. & Nishimura, Y. Epigenetics and neuroinflammation associated with neurodevelopmental disorders: A microglial perspective. Front Cell Dev. Biol. 10, 852752 (2022).
Almehmadi, K. A., Tsilioni, I. & Theoharides, T. C. Increased expression of Mir-155p5 in amygdala of children with autism spectrum disorder. Autism Res 13, 18–23 (2020).
Sunwoo, J. S. et al. Maternal immune activation alters brain microrna expression in mouse offspring. Ann. Clin. Transl. Neurol. 5, 1264–1276 (2018).
Suzumura, A., Sawada, M. & Takayanagi, T. Production of interleukin-12 and expression of its receptors by murine microglia. Brain Res 787, 139–142 (1998).
Yoshida, K. et al. Unique mirna profiling of squamous cell carcinoma arising from ovarian mature teratoma: Comprehensive mirna sequence analysis of its molecular background. Carcinogenesis 40, 1435–1444 (2019).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. Mirbase: Tools for microrna genomics. Nucleic Acids Res 36, D154–D158 (2008).
Kang, J. et al. Identification of key micrornas in diabetes mellitus erectile dysfunction rats with stem cell therapy by bioinformatic analysis of deep sequencing data. World J. Mens. Health 40, 663–677 (2022).
Zhang, L., Lu, D., Liu, M., Zhang, M. & Peng, Q. Identification and interaction analysis of key mirnas in medullary thyroid carcinoma by bioinformatics analysis. Mol. Med Rep. 20, 2316–2324 (2019).
Mizuno, M. et al. The role of E2f8 in the human placenta. Mol. Med. Rep. 19, 293–301 (2019).
ElAtta, A. A., Ali, Y., Bassyouni, I. & Talaat, R. Correlation of myomir-206 and proinflammatory cytokines (Il-16 and Il-17) in patients with rheumatoid arthritis. Reumatologia 57, 72–77 (2019).
Guan, W. et al. Hippocampal Mir-206-3p participates in the pathogenesis of depression via regulating the expression of Bdnf. Pharm. Res. 174, 105932 (2021).
Li, Q., Zhang, J., Gao, Z., Zhang, Y. & Gu, J. Gut microbiota-induced microrna-206-3p increases anxiety-like behaviors by inhibiting expression of Cited2 and Stk39. Micro. Pathog. 176, 106008 (2023).
Miao, Z. et al. Anxiety-related behaviours associated with microrna-206-3p and Bdnf expression in pregnant female mice following psychological social stress. Mol. Neurobiol. 55, 1097–1111 (2018).
Xie, B. et al. Increased serum Mir-206 level predicts conversion from amnestic mild cognitive impairment to Alzheimer’s disease: A 5-year follow-up study. J. Alzheimers Dis. 55, 509–520 (2017).
Moon, J. et al. Early diagnosis of Alzheimer’s disease from elevated olfactory mucosal Mir-206 level. Sci. Rep. 6, 20364 (2016).
Lee, S. T. et al. Mir-206 regulates brain-derived neurotrophic factor in Alzheimer disease model. Ann. Neurol. 72, 269–277 (2012).
Quagliato, L. A., de Matos, U. & Nardi, A. E. Maternal immune activation generates anxiety in offspring: A translational meta-analysis. Transl. Psychiatry 11, 245 (2021).
Sullivan, B. P. et al. Skeletal muscle Igf-1 is lower at rest and after resistance exercise in humans with obesity. Eur. J. Appl. Physiol. 120, 2835–2846 (2020).
Wu, H. Y., Wang, X. H., Liu, K. & Zhang, J. L. Lncrna Malat1 regulates trophoblast cells migration and invasion Via Mir-206/Igf-1 axis. Cell Cycle 19, 39–52 (2020).
Yu, Q., Zhao, B., He, Q., Zhang, Y. & Peng, X. B. Microrna-206 is required for osteoarthritis development through its effect on apoptosis and autophagy of articular chondrocytes via modulating the phosphoinositide 3-kinase/protein kinase B-Mtor pathway by targeting insulin-like growth factor-1. J. Cell Biochem. 120, 5287–5303 (2019).
Simon, L. et al. Decreased myoblast differentiation in chronic binge alcohol-administered simian immunodeficiency virus-infected male macaques: Role of decreased Mir-206. Am. J. Physiol. Regul. Integr. Comp. Physiol. 313, R240–r250 (2017).
Wen, J., He, T., Qi, F. & Chen, H. Mir-206-3p alleviates chronic constriction injury-induced neuropathic pain through targeting Hdac4. Exp. Anim. 68, 213–220 (2019).
Di Pietro, L. et al. Potential therapeutic targets for Als: Mir206, Mir208b and Mir499 are modulated during disease progression in the skeletal muscle of patients. Sci. Rep. 7, 9538 (2017).
Pan, Y. et al. Activation of Ampk inhibits Tgf-Β1-induced airway smooth muscle cells proliferation and its potential mechanisms. Sci. Rep. 8, 3624 (2018).
Guida, N. et al. The Mir206-Jund circuit mediates the neurotoxic effect of methylmercury in cortical neurons. Toxicol. Sci. 163, 569–578 (2018).
Toma, C. et al. Common and rare variants of microrna genes in autism spectrum disorders. World J. Biol. Psychiatry 16, 376–386 (2015).
Du, Y. et al. Genome-wide, integrative analysis implicates exosome-derived microrna dysregulation in Schizophrenia. Schizophr. Bull. 45, 1257–1266 (2019).
Hansen, T. et al. Brain expressed micrornas implicated in Schizophrenia etiology. PLoS One 2, e873 (2007).
Hauberg, M. E. et al. Schizophrenia risk variants affecting microrna function and site-specific regulation of Nt5c2 by Mir-206. Eur. Neuropsychopharmacol. 26, 1522–1526 (2016).
Tian, T. et al. Mirna profiling in the hippocampus of attention-deficit/hyperactivity disorder rats. J. Cell Biochem 120, 3621–3629 (2019).
Kolevzon, A. et al. Clinical trial of insulin-like growth factor-1 in phelan-mcdermid syndrome. Mol. Autism 13, 17 (2022).
Linker, S. B., Mendes, A. P. D. & Marchetto, M. C. Igf-1 treatment causes unique transcriptional response in neurons from individuals with idiopathic autism. Mol. Autism 11, 55 (2020).
Arjunan, A., Sah, D. K., Woo, M. & Song, J. Identification of the molecular mechanism of insulin-like growth factor-1 (Igf-1): A promising therapeutic target for neurodegenerative diseases associated with metabolic syndrome. Cell Biosci. 13, 16 (2023).
Costales, J. & Kolevzon, A. The therapeutic potential of insulin-like growth factor-1 in central nervous system disorders. Neurosci. Biobehav. Rev. 63, 207–222 (2016).
Wlodarczyk, A. et al. A novel microglial subset plays a key role in myelinogenesis in developing brain. EMBO J. 36, 3292–3308 (2017).
Szczesny, E. et al. The impact of prenatal stress on insulin-like growth factor-1 and pro-inflammatory cytokine expression in the brains of adult male rats: The possible role of suppressors of cytokine signaling proteins. J. Neuroimmunol. 276, 37–46 (2014).
Grozinger, C. M., Hassig, C. A. & Schreiber, S. L. Three proteins define a class of human histone deacetylases related to yeast Hda1p. Proc. Natl. Acad. Sci. USA 96, 4868–4873 (1999).
Wu, Y. et al. Aberrant expression of histone deacetylases 4 in cognitive disorders: Molecular mechanisms and a potential target. Front Mol. Neurosci. 9, 114 (2016).
Horvath, R. J., Nutile-McMenemy, N., Alkaitis, M. S. & Deleo, J. A. Differential migration, Lps-induced cytokine, chemokine, and no expression in immortalized Bv-2 and hapi cell lines and primary microglial cultures. J. Neurochem. 107, 557–569 (2008).
Henn, A. et al. The suitability of Bv2 cells as alternative model system for primary microglia cultures or for animal experiments examining brain inflammation. Altex 26, 83–94 (2009).
Chang, C. H., Kuek, E. J. W., Su, C. L. & Gean, P. W. Microrna-206 regulates stress-provoked aggressive behaviors in post-weaning social isolation mice. Mol. Ther. Nucleic Acids 20, 812–822 (2020).
Xing, H., Guo, S., Zhang, Y., Zheng, Z. & Wang, H. Upregulation of microrna-206 enhances lipopolysaccharide-induced inflammation and release of amyloid-Β by targeting insulin-like growth factor 1 in microglia. Mol. Med. Rep. 14, 1357–1364 (2016).
Acknowledgements
We express our gratitude to the members of the Department of Obstetrics and Gynecology at the Nagoya University Graduate School of Medicine. We received technical support from the Division of Medical Research Engineering of Nagoya University Graduate School of Medicine. Funding: This work was financially supported by the Strategic Professional Development Program for Young Researchers (MEXT) and the Japan Society for the Promotion of Science Grant-in-Aid for Scientific Research (Grant Numbers 17K11230 and 22K09638).
Author information
Authors and Affiliations
Contributions
Conception and design; Y.I., T.U., K.I. and T.K., Acquisition of data; Y.I., R.M., K.F., S.T. and T.K., Analysis and interpretation of data; Y.I., T.U., K.I., R.M., K.Y., A.Y. and T.K., Drafting the article or revising it critically for important intellectual content; Y.I., T.U., K.I., A.Y., H.K., and T.K.. Final approval of the version to be published; all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Iitani, Y., Miki, R., Imai, K. et al. Interleukin-17A stimulation induces alterations in Microglial microRNA expression profiles. Pediatr Res 95, 167–173 (2024). https://doi.org/10.1038/s41390-023-02825-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-023-02825-6