Abstract
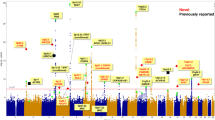
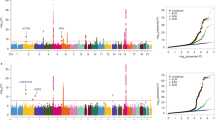
Gene expression variation is an important mechanism underlying susceptibility to complex disease. In comparison with tobacco-related lung carcinogenesis, lung cancer in nonsmokers may involve important and etiologically distinct causal pathways. In this study, we conducted a genome-wide association study on spontaneous lung tumor incidence in inbred mice and identified a major susceptibility locus on mouse chromosome 2 (rs27328255, P=6.68 × 10−7). We then evaluated the correlations of polymorphisms with the transcription of positional candidate genes in normal lungs. Single-nucleotide polymorphism rs27328255 was consistently and strongly associated (P=7.42 × 10−9) in cis with transcript levels of Xrn2. We further showed that Xrn2 promotes proliferation and inhibits squamous differentiation in human lung epithelial cells and polymorphisms in human homolog XRN2 are associated with human lung cancer (rs2025811, P=1.90 × 10−3, OR=1.20). We conclude that genetic variants regulating Xrn2 expression in cis are determinants of spontaneous lung tumor susceptibility in mice and have genetic equivalents in lung cancer susceptibility in human beings. Identifying Xrn2 as a major candidate for spontaneous lung cancer has important implications for the diagnosis and treatment of lung cancer as well as delineation of the mechanisms underlying the genesis of lung cancer in nonsmokers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Amos CI, Wu X, Broderick P, Gorlov IP, Gu J, Eisen T et al. (2008). Genome-wide association scan of tag SNPs identifies a susceptibility locus for lung cancer at 15q25.1. Nat Genet 40: 616–622.
Borczuk AC, Gorenstein L, Walter KL, Assaad AA, Wang L, Powell CA . (2003). Non-small-cell lung cancer molecular signatures recapitulate lung developmental pathways. Am J Pathol 163: 1949–1960.
Borczuk AC, Powell CA . (2007). Expression profiling and lung cancer development. Proc Am Thorac Soc 4: 127–132.
Clark AG, Hubisz MJ, Bustamante CD, Williamson SH, Nielsen R . (2005). Ascertainment bias in studies of human genome-wide polymorphism. Genome Res 15: 1496–1502.
Cookson W, Liang L, Abecasis G, Moffatt M, Lathrop M . (2009). Mapping complex disease traits with global gene expression. Nat Rev Genet 10: 184–194.
Couzin J, Kaiser J . (2007). Genome-wide association. Closing the net on common disease genes. Science 316: 820–822.
Doll R, Peto R . (1981). The causes of cancer: quantitative estimates of avoidable risks of cancer in the United States today. J Natl Cancer Inst 66: 1191–1308.
Foley JF, Anderson MW, Stoner GD, Gaul BW, Hardisty JF, Maronpot RR . (1991). Proliferative lesions of the mouse lung: progression studies in strain A mice. Exp Lung Res 17: 157–168.
Frazer KA, Eskin E, Kang HM, Bogue MA, Hinds DA, Beilharz EJ et al. (2007). A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature 448: 1050–1053.
Hung RJ, McKay JD, Gaborieau V, Boffetta P, Hashibe M, Zaridze D et al. (2008). A susceptibility locus for lung cancer maps to nicotinic acetylcholine receptor subunit genes on 15q25. Nature 452: 633–637.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Thun MJ . (2009). Cancer statistics, 2009. CA Cancer J Clin 59: 225–249 caac.20006.
Kang HM, Zaitlen NA, Wade CM, Kirby A, Heckerman D, Daly MJ et al. (2008). Efficient control of population structure in model organism association mapping. Genetics 178: 1709–1723.
Ke Y, Reddel RR, Gerwin BI, Miyashita M, McMenamin M, Lechner JF et al. (1988). Human bronchial epithelial cells with integrated SV40 virus T antigen genes retain the ability to undergo squamous differentiation. Differentiation 38: 60–66.
Kendall J, Liu Q, Bakleh A, Krasnitz A, Nguyen KC, Lakshmi B et al. (2007). Oncogenic cooperation and coamplification of developmental transcription factor genes in lung cancer. Proc Natl Acad Sci USA 104: 16663–16668.
Kimura S, Hara Y, Pineau T, Fernandez-Salguero P, Fox CH, Ward JM et al. (1996). The T/ebp null mouse: thyroid-specific enhancer-binding protein is essential for the organogenesis of the thyroid, lung, ventral forebrain, and pituitary. Genes Dev 10: 60–69.
Kuntz E, Hoeller U, Greatrix B, Lankin C, Seifert N, Acharya S et al. (2006). Beta-carotene and apocarotenals promote retinoid signaling in BEAS-2B human bronchioepithelial cells. Arch Biochem Biophys 455: 48–60.
Kwei KA, Kim YH, Girard L, Kao J, Pacyna-Gengelbach M, Salari K et al. (2008). Genomic profiling identifies TITF1 as a lineage-specific oncogene amplified in lung cancer. Oncogene 27: 3635–3640.
Li J, Zheng H, Ji C, Fei X, Zheng M, Gao Y et al. (2005). A novel splice variant of human XRN2 gene is mainly expressed in blood leukocyte. DNA Seq 16: 143–146.
Liu P, Vikis HG, Wang D, Lu Y, Wang Y, Schwartz AG et al. (2008). Familial aggregation of common sequence variants on 15q24-25.1 in lung cancer. J Natl Cancer Inst 100: 1326–1330.
Liu P, Wang Y, Vikis H, Maciag A, Wang D, Lu Y et al. (2006). Candidate lung tumor susceptibility genes identified through whole-genome association analyses in inbred mice. Nat Genet 38: 888–895.
Matakidou A, Eisen T, Houlston RS . (2005). Systematic review of the relationship between family history and lung cancer risk. Br J Cancer 93: 825–833.
McKay JD, Hung RJ, Gaborieau V, Boffetta P, Chabrier A, Byrnes G et al. (2008). Lung cancer susceptibility locus at 5p15.33. Nat Genet 40: 1404–1406.
Meuwissen R, Berns A . (2005). Mouse models for human lung cancer. Genes Dev 19: 643–664.
Moffatt MF, Kabesch M, Liang L, Dixon AL, Strachan D, Heath S et al. (2007). Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448: 470–473.
Morley M, Molony CM, Weber TM, Devlin JL, Ewens KG, Spielman RS et al. (2004). Genetic analysis of genome-wide variation in human gene expression. Nature 430: 743–747.
Park MH, Cho SA, Yoo KH, Yang MH, Ahn JY, Lee HS et al. (2007). Gene expression profile related to prognosis of acute myeloid leukemia. Oncol Rep 18: 1395–1402.
Powell CA, Bueno R, Borczuk AC, Caracta CF, Richards WG, Sugarbaker DJ et al. (2003). Patterns of allelic loss differ in lung adenocarcinomas of smokers and nonsmokers. Lung Cancer 39: 23–29.
Rafnar T, Sulem P, Stacey SN, Geller F, Gudmundsson J, Sigurdsson A et al. (2009). Sequence variants at the TERT-CLPTM1 L locus associate with many cancer types. Nat Genet 41: 221–227.
Said JW, Nash G, Sassoon AF, Shintaku IP, Banks-Schlegel S . (1983). Involucrin in lung tumors. A specific marker for squamous differentiation. Lab Invest 49: 563–568.
Schadt EE, Monks SA, Drake TA, Lusis AJ, Che N, Colinayo V et al. (2003). Genetics of gene expression surveyed in maize, mouse and man. Nature 422: 297–302.
Shigematsu H, Gazdar AF . (2006). Somatic mutations of epidermal growth factor receptor signaling pathway in lung cancers. Int J Cancer 118: 257–262.
Smith R, Owen LA, Trem DJ, Wong JS, Whangbo JS, Golub TR et al. (2006). Expression profiling of EWS/FLI identifies NKX2.2 as a critical target gene in Ewing's sarcoma. Cancer Cell 9: 405–416.
Tanaka H, Yanagisawa K, Shinjo K, Taguchi A, Maeno K, Tomida S et al. (2007). Lineage-specific dependency of lung adenocarcinomas on the lung development regulator TTF-1. Cancer Res 67: 6007–6011.
Tang H, Ross A, Capel B . (2008). Expression and functional analysis of Gm114, a putative mammalian ortholog of Drosophila bam. Dev Biol 318: 73–81.
Thorgeirsson TE, Geller F, Sulem P, Rafnar T, Wiste A, Magnusson KP et al. (2008). A variant associated with nicotine dependence, lung cancer and peripheral arterial disease. Nature 452: 638–642.
Thun MJ, Henley SJ, Burns D, Jemal A, Shanks TG, Calle EE . (2006). Lung cancer death rates in lifelong nonsmokers. J Natl Cancer Inst 98: 691–699.
Tian J, Mahmood R, Hnasko R, Locker J . (2006). Loss of Nkx2.8 deregulates progenitor cells in the large airways and leads to dysplasia. Cancer Res 66: 10399–10407.
Wacholder S, Chatterjee N, Caporaso N . (2008). Intermediacy and gene-environment interaction: the example of CHRNA5-A3 region, smoking, nicotine dependence, and lung cancer. J Natl Cancer Inst 100: 1488–1491.
Walts AE, Said JW, Siegel MB, Banks-Schlegel S . (1985). Involucrin, a marker of squamous and urothelial differentiation. An immunohistochemical study on its distribution in normal and neoplastic tissues. J Pathol 145: 329–340.
Wang Y, Broderick P, Webb E, Wu X, Vijayakrishnan J, Matakidou A et al. (2008). Common 5p15.33 and 6p21.33 variants influence lung cancer risk. Nat Genet 40: 1407–1409.
Weir BA, Woo MS, Getz G, Perner S, Ding L, Beroukhim R et al. (2007). Characterizing the cancer genome in lung adenocarcinoma. Nature 450: 893–898.
Wong MP, Fung LF, Wang E, Chow WS, Chiu SW, Lam WK et al. (2003). Chromosomal aberrations of primary lung adenocarcinomas in nonsmokers. Cancer 97: 1263–1270.
You M, Bergman G . (1998). Preclinical and clinical models of lung cancer chemoprevention. Hematol Oncol Clin North Am 12: 1037–1053.
Zeka A, Mannetje A, Zaridze D, Szeszenia-Dabrowska N, Rudnai P, Lissowska J et al. (2006). Lung cancer and occupation in nonsmokers: a multicenter case-control study in Europe. Epidemiology 17: 615–623.
Acknowledgements
We thank the Broad Institute of Harvard/Massachusetts General Hospital and Massachusetts Institute of Technology, the Wellcome Trust Center for Human Genetics at Oxford University, and Perlegen Sciences for releasing inbred laboratory mouse SNP data. We thank the Mouse Phenome Project for collecting mouse SNP data. We thank the International Agency for Research on Cancer (Lyon, France) for releasing lung cancer GWAS data. We also thank investigators for generating spontaneous lung tumor data in inbred mice. The mouse SNP and phenotype data were crucial for the key findings described in this paper. We thank the staff of The Vanderbilt Microarray Shared Resource for microarray processing. MY was supported by grants from the US National Institutes of Health (CA099187, CA099147, ES012063, and ES013340). We thank Dr Jay W Tichelaar for his critical comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Lu, Y., Liu, P., James, M. et al. Genetic variants cis-regulating Xrn2 expression contribute to the risk of spontaneous lung tumor. Oncogene 29, 1041–1049 (2010). https://doi.org/10.1038/onc.2009.396
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.396
Keywords
This article is cited by
-
XRN2 interactome reveals its synthetic lethal relationship with PARP1 inhibition
Scientific Reports (2020)
-
XRN2 promotes EMT and metastasis through regulating maturation of miR-10a
Oncogene (2017)