Abstract
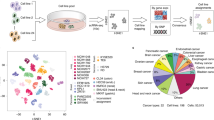
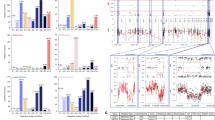
The oncogenic phenotype is complex, resulting from the accumulation of multiple somatic mutations that lead to the deregulation of growth regulatory and cell fate controlling activities and pathways. The ability to dissect this complexity, so as to reveal discrete aspects of the biology underlying the oncogenic phenotype, is critical to understanding the various mechanisms of disease as well as to reveal opportunities for novel therapeutic strategies. Previous work has characterized the process of anchorage-independent growth of cancer cells in vitro as a key aspect of the tumor phenotype, particularly with respect to metastatic potential. Nevertheless, it remains a major challenge to translate these cell biology findings into the context of human tumors. We previously used DNA microarray assays to develop expression signatures, which have the capacity to identify subtle distinctions in biological states and can be used to connect in vitro and in vivo states. Here we describe the development of a signature of anchorage-independent growth, show that the signature exhibits characteristics of deregulated mitochondrial function and then demonstrate that the signature identifies human tumors with the potential for metastasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bild A, Yao G, Chang JT, Wang Q, Potti A, Chasse D et al. (2006). Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439: 353–357.
Calvo S, Jain M, Xie X, Sheth SA, Chang B, Goldberger OA . (2006). Systematic identification of human mitochondrial disease genes through integrative genomics. Nat Genet 38: 576–582.
Campbell PM, Der CJ . (2004). Oncogenic Ras and its role in tumor cell invasion and metastasis. Semin Cancer Biol 14: 105–114.
Chang JT, Nevins JR . (2006). GATHER: a systems approach to interpreting genomic signatures. Bioinformatics 22: 2926–2933.
Chen EI, Hewel J, Krueger JS, Tiraby C, Weber MR, Kralli A et al. (2007). Adaptation of energy metabolism in breast cancer brain metastases. Cancer Res 67: 1472–1486.
Cifone MA, Fidler IJ . (1980). Correlation of patterns of anchorage-independent growth with in vivo behavior of cells from a murine fibrosarcoma. Proc Natl Acad Sci USA 77: 1039–1043.
Dang CV, O'Donnell KA, Zeller KI, Nguyen T, Osthus RC, Li F . (2006). The c-Myc target gene network. Semin Cancer Biol 16: 253–264.
Eccles SA, Welch DR . (2007). Metastasis: recent discoveries and novel treatment strategies. Lancet 369: 1742–1757.
Edelman E, Porrello A, Guinney J, Balakumaran B, Bild A, Febbo PG et al. (2006). Analysis of sample set enrichment scores: assaying the enrichment of sets of genes for individual samples in genome-wide expression profiles. Bioinformatics 22: e108–116.
Funes JM, Quintero M, Henderson S, Martinez D, Qureshi U, Westwood C et al. (2007). Transformation of human mesenchymal stem cells increases their dependency on oxidative phosphorylation for energy production. Proc Natl Acad Sci USA 104: 6223–6228.
Giangrande P, Zhu W, Schlisio S, Sun XH, Mori S, Gaubatz S et al. (2004). Genes Dev 18: 2941–2951.
Guda C, Subramaniam S . (2005). pTARGET [corrected] a new method for predicting protein subcellular localization in eukaryotes. Bioinformatics 21: 3963–3969.
Guo W, Giancotti FG . (2004). Integrin signalling during tumour progression. Nat Rev Mol Cell Biol 5: 816–826.
Hackett AJ, Smith HS, Springer EL, Owens RB, Nelson-Rees WA, Riggs JL et al. (1977). Two syngeneic cell lines from human breast tissue: the aneuploid mammary epithelial (Hs578T) and the diploid myoepithelial (Hs578Bst) cell lines. J Natl Cancer Inst 58: 1795–1806.
Hanahan D, Weinberg RA . (2000). The hallmarks of cancer. Cell 100: 57–70.
Johns ME, Mills SE . (1983). Cloning efficiency. A possible prognostic indicator in squamous cell carcinoma of the head and neck. Cancer 52: 1401–1404.
Johnson WE, Li C, Rabinovic A . (2007). Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8: 118–127.
Kang D, Hamasaki N . (2005). Ann NY Acad Sci 1042: 101–108.
Li F, Wang Y, Zeller KI, Potter JJ, Wonsey DR, O'Donnell KA et al. (2005). Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol Cell Biol 25: 6225–6234.
Lopez-Otin C, Matrisian LM . (2007). Emerging roles of proteases in tumour suppression. Nat Rev Cancer 7: 800–808.
Mattox DE, Von Hoff DD . (1980). in vitro stem cell assay in head and neck squamous carcinoma. Am J Surg 140: 527–530.
Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD et al. (2005). Genes that mediate breast cancer metastasis to lung. Nature 436: 518–524.
Minn AJ, Gupta GP, Padua D, Bos P, Nguyen DX, Nuyten D et al. (2007). Lung metastasis genes couple breast tumor size and metastatic spread. Proc Natl Acad Sci USA 104: 6740–6745.
Mukhopadhyay R, Theriault RL, Price JE . (1999). Clin Exp Metastasis 17: 325–332.
Nevins JR, Potti A . (2007). Mining gene expression profiles: expression signatures as cancer phenotypes. Nat Rev Genet 8: 601–609.
Nicolson GL, Lembo TM, Welch DR . (1988). Growth of rat mammary adenocarcinoma cells in semisolid clonogenic medium not correlated with spontaneous metastatic behavior: heterogeneity in the metastatic, antigenic, enzymatic, and drug sensitivity properties of cells from different sized colonies. Cancer Res 48: 399–404.
Nomura Y, Tashiro H, Hisamatsu K . (1989). in vitro clonogenic growth and metastatic potential of human operable breast cancer. Cancer Res 49: 5288–5293.
Price JE . (1986). Clonogenicity and experimental metastatic potential of spontaneous mouse mammary neoplasms. J Natl Cancer Inst 77: 529–535.
Samuels Y, Ericson K . (2006). Oncogenic PI3K and its role in cancer. Curr Opin Oncol 18: 77–82.
Scott MS, Thomas DY, Hallett MT . (2004). Predicting subcellular localization via protein motif co-occurrence. Genome Res 14: 1957–1966.
Shafie SM, Liotta LA . (1980). Formation of metastasis by human breast carcinoma cells (MCF-7) in nude mice. Cancer Lett 11: 81–87.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gilletee MA et al. (2005). Proc Natl Acad Sci USA 102: 15545–15550.
Sutherland CM, Mather FJ, Carter RD, Cerise EJ, Krementz ET . (1983). Breast cancer as analyzed by the human tumor stem cell assay. Surgery 94: 370–375.
Telang S, Lane AN, Nelson KK, Arumugam S, Chesney J . (2007). The oncoprotein H-RasV12 increases mitochondrial metabolism. Mol Cancer 6: 77.
Thompson EW, Paik S, Brünner N, Sommers CL, Zugmaier G, Clarke R et al. (1992). Association of increased basement membrane invasiveness with absence of estrogen receptor and expression of vimentin in human breast cancer cell lines. J Cell Physiol 150: 534–544.
Tong X, Zhao F, Thompson CB . (2009). The molecular determinants of de novo nucleotide biosynthesis in cancer cells. Curr Opin Genet Dev 19: 32–37.
van Slooten HJ, Bonsing BA, Hiller AJ, Colbern GT, van Dierendonck JH, Cornelisse CJ et al. (1995). Outgrowth of BT-474 human breast cancer cells in immune-deficient mice: a new in vivo model for hormone-dependent breast cancer. Br J Cancer 72: 22–30.
Vogelstein B, Kinzler KW . (2004). Cancer genes and the pathways they control. Nat Med 10: 789–799.
Warburg O . (1956). On respiratory impairment in cancer cells. Science 124: 269–270.
Weinberg RA . (2007). The biology of cancer. Garland Science: New York.
Yin JJ, Mohammad KS, Kakonen SM, Harris S, Wu-Wong JR, Wessale JL et al. (2003). A causal role for endothelin-1 in the pathogenesis of osteoblastic bone metastases. Proc Natl Acad Sci USA 100: 10954–10959.
Zhang RD, Fidler IJ, Price JE . (1991). Relative malignant potential of human breast carcinoma cell lines established from pleural effusions and a brain metastasis. Invasion Metastasis 11: 204–215.
Zu XL, Guppy M . (2004). Cancer metabolism: facts, fantasy, and fiction. Biochem Biophys Res Commun 313: 459–465.
Acknowledgements
We thank Drs J-T Chi, T Hallstrom, L Kong, R Rempel, J Freedman, M Gatza, Dr B Balakumaran, H Dressman, Y Yokota, S Akiyama, T Inoue, M Oshimura, M Araki, H Saya, Aburatani, A Niida, K Araki, Y Taya and Y Ito for helpful discussions; A Bild, T Kitamura, M Eilers and H Hanafusa for providing materials; L Jakoi for help with experiments; and T Henry and K Culler for assistance with the preparation of the article. This work was supported by a grant (5U54CA-112952-05) from the NIH/NCI to JRN. SM was a research fellow of the Uehara Memorial Foundation and is a visiting scholar of Riken Genome Science Center.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Mori, S., Chang, J., Andrechek, E. et al. Anchorage-independent cell growth signature identifies tumors with metastatic potential. Oncogene 28, 2796–2805 (2009). https://doi.org/10.1038/onc.2009.139
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.139