Abstract
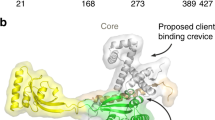
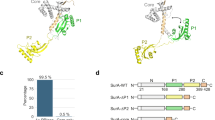
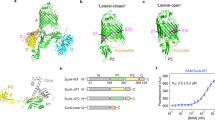
The 17-kDa protein (Skp) of Escherichia coli is a homotrimeric periplasmic chaperone for newly synthesized outer-membrane proteins. Here we present its X-ray structure at a resolution of 2.35 Å. Three hairpin-shaped α-helical extensions reach out by ∼60 Å from a trimerization domain, which is composed of three intersubunit β-sheets that wind around a central axis. The α-helical extensions approach each other at their distal turns, resulting in a fold that resembles a 'three-pronged grasping forceps'. The overall shape of Skp is reminiscent of the cytosolic chaperone prefoldin, although it is based on a radically different topology. The peculiar architecture, with apparent plasticity of the prongs and distinct electrostatic and hydrophobic surface properties, supports the recently proposed biochemical mechanism of this chaperone: formation of a Skp3–Omp complex protects the outer membrane protein from aggregation during passage through the bacterial periplasm.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Danese, P.N. & Silhavy, T.J. Targeting and assembly of periplasmic and outer-membrane proteins in Escherichia coli. Annu. Rev. Genet. 32, 59–94 (1998).
Thome, B.M. & Müller, M. Skp is a periplasmic Escherichia coli protein requiring SecA and SecY for export. Mol. Microbiol. 5, 2815–2821 (1991).
Chen, R. & Henning, U. A periplasmic protein (Skp) of Escherichia coli selectively binds a class of outer membrane proteins. Mol. Microbiol. 19, 1287–1294 (1996).
Schäfer, U., Beck, K. & Müller, M. Skp, a molecular chaperone of gram-negative bacteria, is required for the formation of soluble periplasmic intermediates of outer membrane proteins. J. Biol. Chem. 35, 24567–24574 (1999).
De Cock, H. et al. Affinity of the periplasmic chaperone Skp of Escherichia coli for phospholipids, lipopolysaccharides and non-native outer membrane proteins. Role of Skp in the biogenesis of outer membrane protein. Eur. J. Biochem. 259, 96–103 (1999).
Harms, N. et al. The early interaction of the outer membrane protein PhoE with the periplasmic chaperone Skp occurs at the cytoplasmic membrane. J. Biol. Chem. 276, 18804–18811 (2001).
Pautsch, A. & Schulz, G.E. Structure of the outer membrane protein A transmembrane domain. Nat. Struct. Biol. 5, 1013–1017 (1998).
Cowan, S.W. et al. Crystal structures explain functional properties of two E. coli porins. Nature 358, 727–733 (1992).
Dutzler, R., Wang, Y.F., Rizkallah, P., Rosenbusch, J.P. & Schirmer, T. Crystal structures of various maltooligosaccharides bound to maltoporin reveal a specific sugar translocation pathway. Structure 4, 127–134 (1996).
Bulieris, P.V., Behrens, S., Holst, O. & Kleinschmidt, J.H. Folding and insertion of the outer membrane protein OmpA is assisted by the chaperone Skp and by lipopolysaccharide. J. Biol. Chem. 278, 9092–9099 (2003).
Voulhoux, R., Bos, M.P., Geurtsen, J., Mols, M. & Tommassen, J. Role of a highly conserved bacterial protein in outer membrane protein assembly. Science 299, 262–265 (2003).
Schlapschy, M. et al. The periplasmic E. coli chaperone Skp is a trimer in solution: biophysical and preliminary crystallographic characterization. Biol. Chem. 385, 137–143 (2004).
Crick, F.H.C. The packing of α-helices: simple coiled-coils. Acta Crystallogr. 6, 689–697 (1953).
Kleywegt, G.J., Zou, J.Y., Kjeldgaard, M. & Jones, T.A. Around O. In International Tables for Crystallography Vol. F (eds. Rossmann, M.G. & Arnold, E.) 353–356, 366–367 (Kluwer Academic, Dordrecht, The Netherlands, 2001).
Bansal, M., Kumar, S. & Velavan, R. HELANAL—a program to characterize helix geometry in proteins. J. Biomol. Struct. Dyn. 17, 811–819 (2000).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 223, 123–138 (1993).
Siegert, R., Leroux, M.R., Scheufler, C., Hartl, F.U. & Moarefi, I. Structure of the molecular chaperone prefoldin: unique interaction of multiple coiled coil tentacles with unfolded proteins. Cell 103, 621–632 (2000).
Martín-Benito, J. et al. Structure of eukaryotic prefoldin and of its complexes with unfolded actin and the cytosolic chaperonin CCT. EMBO J. 21, 6377–6386 (2002).
Sha, B., Lee, S. & Cyr, D.M. The crystal structure of the peptide-binding fragment from the yeast Hsp40 protein Sis1. Structure 8, 799–807 (2000).
Devereux, J., Haeberli, P. & Smithies, O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids. Res. 12, 387–395 (1984).
Xu, Z., Horwich, A.L. & Sigler, P.B. The crystal structure of the asymmetric GroEL–GroES–(ADP)7 chaperonin complex. Nature 388, 741–750 (1997).
Zavialov, A.V. et al. Structure and biogenesis of the capsular F1 antigen from Yersinia pestis: preserved folding energy drives fiber formation. Cell 113, 587–596 (2003).
Creighton, T.E. Proteins: Structures and Molecular Properties (W.H. Freeman, New York, 1993).
Koppenol, W.H. & Margoliash, E. The asymmetric distribution of charges on the surface of horse cytochrome c. Functional implications. J. Biol. Chem. 257, 4426–4437 (1982).
Neidhardt, F.C., Ingraham, J.L. & Schaechter, M. Physiology of the Bacterial Cell: A Molecular Approach (Sinauer Associates, Sunderland, Massachusetts, USA, 1990).
Skerra, A. Lipocalins as a scaffold. Biochim. Biophys. Acta 1482, 337–350 (2000).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002).
De la Fortelle, E., Irwin, J. & Bricogne, G. SHARP: a maximum-likelihood heavy atom parameter refinement and phasing program for the MIR and MAD methods. In Crystallographic Computing Vol. 7 (eds. Bourne, P.E. & Watenpaugh, K.D.) (Oxford Univ. Press, Oxford, 1997).
Oldfield, T.J. A number of real-space torsion-angle refinement techniques for proteins, nucleic acids, ligands and solvent. Acta Crystallogr. D 57, 82–94 (2001).
Ramachandran, G.N. & Sasisekharan, V. Conformation of polypeptides and proteins. Adv. Prot. Chem. 23, 283–437 (1968).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallog. 24, 946–950 (1991).
Merrit, E.A. & Bacon, D.J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Barton, G.J. ALSCRIPT. A tool to format multiple sequence alignments. Protein Eng. 6, 37–40 (1993).
Glaser, F. et al. ConSurf: identification of functional regions in proteins by surface-mapping of phylogenetic information. Bioinformatics 19, 163–164 (2003).
Tao, P., Wang, R. & Pai, L. Calculating partition coefficients of peptides by the addition method. J. Mol. Model. 5, 189–195 (1999).
Acknowledgements
We thank J. Danzer for help with the preparation and crystallization of Skp, G. Reil for mass spectrometric analysis of SeMet derivatives, and the European Molecular Biology Laboratory Grenoble Outstation, in particular M. Walsh, for supporting measurements at the European Synchrotron Radiation Facility under the European Union Improving Human Potential Programme. This work was supported by the Fonds der Chemischen Industrie.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
Representative 2Fo − Fc electron density from the prong of chain C contoured at 1.0 σ. (PDF 114 kb)
Rights and permissions
About this article
Cite this article
Korndörfer, I., Dommel, M. & Skerra, A. Structure of the periplasmic chaperone Skp suggests functional similarity with cytosolic chaperones despite differing architecture. Nat Struct Mol Biol 11, 1015–1020 (2004). https://doi.org/10.1038/nsmb828
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb828
This article is cited by
-
Engineering sigma factors and chaperones for enhanced heterologous lipoxygenase production in Escherichia coli
Biotechnology for Biofuels and Bioproducts (2022)
-
Overproducing the BAM complex improves secretion of difficult-to-secrete recombinant autotransporter chimeras
Microbial Cell Factories (2021)
-
Molecular chaperones and their denaturing effect on client proteins
Journal of Biomolecular NMR (2021)
-
Regulation of α-synuclein by chaperones in mammalian cells
Nature (2020)
-
Affinity of Skp to OmpC revealed by single-molecule detection
Scientific Reports (2020)