Abstract
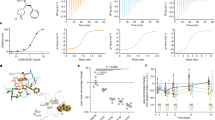
Members of the serum paraoxonase (PON) family have been identified in mammals and other vertebrates, and in invertebrates. PONs exhibit a wide range of physiologically important hydrolytic activities, including drug metabolism and detoxification of nerve agents. PON1 and PON3 reside on high-density lipoprotein (HDL, 'good cholesterol') and are involved in the prevention of atherosclerosis. We describe the first crystal structure of a PON family member, a variant of PON1 obtained by directed evolution, at a resolution of 2.2 Å. PON1 is a six-bladed β-propeller with a unique active site lid that is also involved in HDL binding. The three-dimensional structure and directed evolution studies permit a detailed description of PON1's active site and catalytic mechanism, which are reminiscent of secreted phospholipase A2, and of the routes by which PON family members diverged toward different substrate and reaction selectivities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Draganov, D.I. & La Du, B.N. Pharmacogenetics of paraoxonases: a brief review. Naunyn Schmiedebergs Arch. Pharmacol. 369, 78– 88 (2004).
Lusis, A.J. Atherosclerosis. Nature 407, 233– 241 (2000).
Shih, D.M. et al. Mice lacking serum paraoxonase are susceptible to organophosphate toxicity and atherosclerosis. Nature 394, 284– 287 (1998).
Mackness, M.I., Arrol, S. & Durrington, P.N. Paraoxonase prevents accumulation of lipoperoxides in low-density-lipoprotein. FEBS Lett. 286, 152– 154 (1991).
Reddy, S.T. et al. Human paraoxonase-3 is an HDL-associated enzyme with biological activity similar to paraoxonase-1 protein but is not regulated by oxidized lipids. Arterioscler. Thromb. Vasc. Biol. 21, 542– 547 (2001).
Rodrigo, L., Mackness, B., Durrington, P.N., Hernandez, A. & Mackness, M.I. Hydrolysis of platelet-activating factor by human serum paraoxonase. Biochem. J. 354, 1– 7 (2001).
Ahmed, Z. et al. Apolipoprotein A-I promotes the formation of phosphatidylcholine core aldehydes that are hydrolyzed by paraoxonase (PON-1) during high density lipoprotein oxidation with a peroxynitrite donor. J. Biol. Chem. 276, 24473– 24481 (2001).
Jakubowski, H. Calcium-dependent human serum homocysteine thiolactone hydrolase. A protective mechanism against protein N-homocysteinylation. J. Biol. Chem. 275, 3957– 3962 (2000).
Lund-Katz, S., Liu, L.J., Thuahnai, S.T. & Phillips, M.C. High density lipoprotein structure. Front. Biosci. 8, D1044– D1054 (2003).
Fuhrman, B., Volkova, N. & Aviram, M. Oxidative stress increases the expression of the CD36 scavenger receptor and the cellular uptake of oxidized low-density lipoprotein in macrophages from atherosclerotic mice: protective role of antioxidants and of paraoxonase. Atherosclerosis 161, 307– 316 (2002).
Rozenberg, O., Shih, D.M. & Aviram, M. Human serum paraoxonase 1 decreases macrophage cholesterol biosynthesis: possible role for its phospholipase-A(2)-like activity and lysophosphatidylcholine formation. Arterioscler. Thromb. Vasc. Biol. 23, 461– 467 (2003).
Sorenson, R.C. et al. Human serum paraoxonase/arylesterase's retained hydrophobic N-terminal leader sequence associates with HDLs by binding phospholipids: apolipoprotein A-I stabilizes activity. Arterioscler. Thromb. Vasc. Biol. 19, 2214– 2225 (1999).
Aharoni, A. et al. Directed evolution of mammalian paraoxonases PON1 and PON3 for bacterial expression and catalytic specialization. Proc. Natl. Acad. Sci. USA 101, 482– 487 (2004).
Fokine, A. et al. Direct phasing at low resolution of a protein copurified with human paraoxonase (PON1). Acta Crystallogr. D. 59, 2083– 2087 (2003).
Josse, D. et al. Oligomeric states of the detergent-solubilized human serum paraoxonase (PON1). J. Biol. Chem. 277, 33386– 33397 (2002).
Kuo, C.L. & La Du, B.N. Comparison of purified human and rabbit serum paraoxonases. Drug Metab. Dispos. 23, 935– 944 (1995).
Jawad, Z. & Paoli, M. Novel sequences propel familiar folds. Structure 10, 447– 454 (2002).
Kuo, C.L. & La Du, B.N. Calcium binding by human and rabbit serum paraoxonases. Structural stability and enzymatic activity. Drug Metab. Dispos. 26, 653– 60 (1998).
Scharff, E.I., Koepke, J., Fritzsch, G., Lucke, C. & Ruterjans, H. Crystal structure of diisopropylfluorophosphatase from Loligo vulgaris . Structure 9, 493– 502 (2001).
Josse, D. et al. Identification of residues essential for human paraoxonase (PON1) arylesterase/organophosphatase activities. Biochemistry 38, 2816– 2825 (1999).
Jonas, A. Lecithin cholesterol acyltransferase. Biochim. Biophys. Acta 1529, 245– 256 (2000).
Josse, D. et al. The active site of human paraoxonase (PON1). J. Appl. Toxicol. 21, S7– S11 (2001).
Josse, D., Xie, W.H., Masson, P., Schopfer, L.M. & Lockridge, O. Tryptophan residue(s) as major components of the human serum paraoxonase active site. Chem. Biol. Interact. 120, 79– 84 (1999).
Sekar, K. et al. Phospholipase A2 engineering. Structural and functional roles of the highly conserved active site residue aspartate-99. Biochemistry 36, 3104– 3114 (1997).
Aviram, M. et al. Paraoxonase active site required for protection against LDL oxidation involves its free sulfhydryl group and is different from that required for its arylesterase/paraoxonase activities: selective action of human paraoxonase allozymes Q and R. Arterioscler. Thromb. Vasc. Biol. 18, 1617– 1624 (1998).
Kobayashi, M., Shinohara, M., Sakoh, C., Kataoka, M. & Shimizu, S. Lactone-ring-cleaving enzyme: genetic analysis, novel RNA editing, and evolutionary implications. Proc. Natl. Acad. Sci. USA 95, 12787– 12792 (1998).
Chelur, D.S. et al. The mechanosensory protein MEC-6 is a subunit of the C. elegans touch-cell degenerin channel. Nature 420, 669– 73 (2002).
Navab, M. et al. High density associated enzymes: their role in vascular biology. Curr. Opin. Lipidol. 9, 449– 456 (1998).
Borhani, D.W., Rogers, D.P., Engler, J.A. & Brouillette, C.G. Crystal structure of truncated human apolipoprotein A-I suggests a lipid-bound conformation. Proc. Natl. Acad. Sci. USA 94, 12291– 12296 (1997).
Segrest, J.P., Harvey, S.C. & Zannis, V. Detailed molecular model of apolipoprotein A-I on the surface of high-density lipoproteins and its functional implications. Trends Cardiovasc. Med. 10, 246– 252 (2000).
Killian, J.A. & von Heijne, G. How proteins adapt to a membrane-water interface. Trends Biochem. Sci. 25, 429– 434 (2000).
Waldo, G.S. Genetic screens and directed evolution for protein solubility. Curr. Opin. Chem. Biol. 7, 33– 38 (2003).
Bencharit, S., Morton, C.L., Xue, Y., Potter, P.M. & Redinbo, M.R. Structural basis of heroin and cocaine metabolism by a promiscuous human drug-processing enzyme. Nat. Struct. Biol. 10, 349– 356 (2003).
Millard, C.B., Lockridge, O. & Broomfield, C.A. Organophosphorus acid anhydride hydrolase activity in human butyrylcholinesterase: synergy results in a somanase. Biochemistry 37, 237– 247 (1998).
Greenblatt, H.M., Dvir, H., Silman, I. & Sussman, J.L. Acetylcholinesterase: a multifaceted target for structure-based drug design of anticholinesterase agents for the treatment of Alzheimer's disease. J. Mol. Neurosci. 20, 369– 383 (2003).
Leviev, I., Deakin, S. & James, R.W. Decreased stability of the M54 isoform of paraoxonase as a contributory factor to variations in human serum paraoxonase concentrations. J. Lipid Res. 42, 528– 535 (2001).
Oda, M.N., Bielicki, J.K., Berger, T. & Forte, T.M. Cysteine substitutions in apolipoprotein A-I primary structure modulate paraoxonase activity. Biochemistry 40, 1710– 1718 (2001).
Lee, B. & Richards, F.M. The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55, 379– 389 (1971).
Acknowledgements
We acknowledge financial support by the Minerva Foundation and the Israel Science Foundation to D.S.T. and US army Medical Research & Materiel Command to I.S. and J.L.S. The structure was determined in collaboration with the Israel Structural Proteomics Center, supported by the Israel Ministry of Science & Technology, the European Commission Structural Proteomics Project (SPINE) and the Divadol Foundation. J.L.S. is the Morton and Gladys Pickman Professor of Structural Biology.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Harel, M., Aharoni, A., Gaidukov, L. et al. Structure and evolution of the serum paraoxonase family of detoxifying and anti-atherosclerotic enzymes. Nat Struct Mol Biol 11, 412–419 (2004). https://doi.org/10.1038/nsmb767
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb767
This article is cited by
-
Ca2+ efflux facilitated by co-transport of inorganic phosphate anion in the H+/Ca2+ antiporter YfkE
Communications Biology (2023)
-
A targeted multi-omics approach reveals paraoxonase-1 as a determinant of obesity-associated fatty liver disease
Clinical Epigenetics (2021)
-
Association between serum paraoxonase 1 activity and its polymorphisms with multiple sclerosis: a systematic review
Neurological Sciences (2021)
-
WTAP and BIRC3 are involved in the posttranscriptional mechanisms that impact on the expression and activity of the human lactonase PON2
Cell Death & Disease (2020)
-
Structural and Biochemical Characterization of AaL, a Quorum Quenching Lactonase with Unusual Kinetic Properties
Scientific Reports (2018)