Abstract
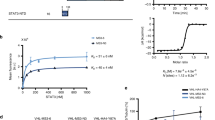
Interactions between Src homology 2 (SH2) domains and phosphotyrosine sites regulate tyrosine kinase signaling networks. Selective perturbation of these interactions is challenging due to the high homology among the 120 human SH2 domains. Using an improved phage-display selection system, we generated a small antibody mimic (or 'monobody'), termed HA4, that bound to the Abelson (Abl) kinase SH2 domain with low nanomolar affinity. SH2 protein microarray analysis and MS of intracellular HA4 interactors showed HA4's specificity, and a crystal structure revealed how this specificity is achieved. HA4 disrupted intramolecular interactions of Abl involving the SH2 domain and potently activated the kinase in vitro. Within cells, HA4 inhibited processive phosphorylation activity of Abl and also inhibited STAT5 activation. This work provides a design guideline for highly specific and potent inhibitors of a protein interaction domain and shows their utility in mechanistic and cellular investigations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Pawson, T. & Nash, P. Assembly of cell regulatory systems through protein interaction domains. Science 300, 445–452 (2003).
Rual, J.F. et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature 437, 1173–1178 (2005).
Kocher, T. & Superti-Furga, G. Mass spectrometry-based functional proteomics: from molecular machines to protein networks. Nat. Methods 4, 807–815 (2007).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Weiss, W.A., Taylor, S.S. & Shokat, K.M. Recognizing and exploiting differences between RNAi and small-molecule inhibitors. Nat. Chem. Biol. 3, 739–744 (2007).
Visintin, M., Melchionna, T., Cannistraci, I. & Cattaneo, A. In vivo selection of intrabodies specifically targeting protein-protein interactions: a general platform for an “undruggable” class of disease targets. J. Biotechnol. 135, 1–15 (2008).
Campbell, S.J. & Jackson, R.M. Diversity in the SH2 domain family phosphotyrosyl peptide binding site. Protein Eng. 16, 217–227 (2003).
Bradshaw, J.M., Mitaxov, V. & Waksman, G. Investigation of phosphotyrosine recognition by the SH2 domain of the Src kinase. J. Mol. Biol. 293, 971–985 (1999).
Ladbury, J.E. & Arold, S. Searching for specificity in SH domains. Chem. Biol. 7, R3–R8 (2000).
Taylor, J.D. et al. Structure, dynamics, and binding thermodynamics of the v-Src SH2 domain: implications for drug design. Proteins 73, 929–940 (2008).
Gan, W. & Roux, B. Binding specificity of SH2 domains: insight from free energy simulations. Proteins 74, 996–1007 (2009).
Shakespeare, W.C. SH2 domain inhibition: a problem solved? Curr. Opin. Chem. Biol. 5, 409–415 (2001).
Machida, K. & Mayer, B.J. The SH2 domain: versatile signaling module and pharmaceutical target. Biochim. Biophys. Acta 1747, 1–25 (2005).
Mandine, E. et al. High-affinity Src-SH2 ligands which do not activate Tyr(527)-phosphorylated Src in an experimental in vivo system. Biochem. Biophys. Res. Commun. 298, 185–192 (2002).
Garcia-Echeverria, C. Antagonists of the Src homology 2 (SH2) domains of Grb2, Src, Lck and ZAP-70. Curr. Med. Chem. 8, 1589–1604 (2001).
Reichert, J.M., Rosensweig, C.J., Faden, L.B. & Dewitz, M.C. Monoclonal antibody successes in the clinic. Nat. Biotechnol. 23, 1073–1078 (2005).
Koide, A., Bailey, C.W., Huang, X. & Koide, S. The fibronectin type III domain as a scaffold for novel binding proteins. J. Mol. Biol. 284, 1141–1151 (1998).
Koide, A., Abbatiello, S., Rothgery, L. & Koide, S. Probing protein conformational changes in living cells by using designer binding proteins: application to the estrogen receptor. Proc. Natl. Acad. Sci. USA 99, 1253–1258 (2002).
Gilbreth, R.N., Esaki, K., Koide, A., Sidhu, S.S. & Koide, S. A dominant conformational role for amino acid diversity in minimalist protein-protein interfaces. J. Mol. Biol. 381, 407–418 (2008).
Daley, G.Q., Van Etten, R.A. & Baltimore, D. Induction of chronic myelogenous leukemia in mice by the P210bcr/abl gene of the Philadelphia chromosome. Science 247, 824–830 (1990).
Nagar, B. et al. Organization of the SH3–SH2 unit in active and inactive forms of the c-Abl tyrosine kinase. Mol. Cell 21, 787–798 (2006).
Hantschel, O. & Superti-Furga, G. Regulation of the c-Abl and Bcr-Abl tyrosine kinases. Nat. Rev. Mol. Cell Biol. 5, 33–44 (2004).
Filippakopoulos, P. et al. Structural coupling of SH2-kinase domains links Fes and Abl substrate recognition and kinase activation. Cell 134, 793–803 (2008).
Hackel, B.J., Kapila, A. & Wittrup, K.D. Picomolar affinity fibronectin domains engineered utilizing loop length diversity, recursive mutagenesis, and loop shuffling. J. Mol. Biol. 381, 1238–1252 (2008).
Tokonzaba, E., Capelluto, D.G., Kutateladze, T.G. & Overduin, M. Phosphoinositide, phosphopeptide and pyridone interactions of the Abl SH2 domain. Chem. Biol. Drug Des. 67, 230–237 (2006).
Porter, C.J. et al. Grb7 SH2 domain structure and interactions with a cyclic peptide inhibitor of cancer cell migration and proliferation. BMC Struct. Biol. 7, 58 (2007).
Burckstummer, T. et al. An efficient tandem affinity purification procedure for interaction proteomics in mammalian cells. Nat. Methods 3, 1013–1019 (2006).
Brehme, M. et al. Charting the molecular network of the drug target Bcr-Abl. Proc. Natl. Acad. Sci. USA 106, 7414–7419 (2009).
Basu, D., El-Assal Sel, D., Le, J., Mallery, E.L. & Szymanski, D.B. Interchangeable functions of Arabidopsis PIROGI and the human WAVE complex subunit SRA1 during leaf epidermal development. Development 131, 4345–4355 (2004).
Ewing, R.M. et al. Large-scale mapping of human protein-protein interactions by mass spectrometry. Mol. Syst. Biol. 3, 89 (2007).
Wu, C. et al. Systematic identification of SH3 domain-mediated human protein-protein interactions by peptide array target screening. Proteomics 7, 1775–1785 (2007).
MacPartlin, M., Smith, A.M., Druker, B.J., Honigberg, L.A. & Deininger, M.W. Bruton's tyrosine kinase is not essential for Bcr-Abl–mediated transformation of lymphoid or myeloid cells. Leukemia 22, 1354–1360 (2008).
Pendergast, A.M. et al. BCR-ABL-induced oncogenesis is mediated by direct interaction with the SH2 domain of the GRB-2 adaptor protein. Cell 75, 175–185 (1993).
Hantschel, O. et al. The chemokine interleukin-8 and the surface activation protein CD69 are markers for Bcr-Abl activity in chronic myeloid leukemia. Mol. Oncol. 2, 272–281 (2008).
El Hader, C. et al. HCaRG increases renal cell migration by a TGF-α autocrine loop mechanism. Am. J. Physiol. Renal Physiol. 289, F1273–F1280 (2005).
Nagar, B. et al. Structural basis for the autoinhibition of c-Abl tyrosine kinase. Cell 112, 859–871 (2003).
Overduin, M., Rios, C.B., Mayer, B.J., Baltimore, D. & Cowburn, D. Three-dimensional solution structure of the src homology 2 domain of c-abl. Cell 70, 697–704 (1992).
Waksman, G., Shoelson, S.E., Pant, N., Cowburn, D. & Kuriyan, J. Binding of a high affinity phosphotyrosyl peptide to the Src SH2 domain: crystal structures of the complexed and peptide-free forms. Cell 72, 779–790 (1993).
Eck, M.J., Shoelson, S.E. & Harrison, S.C. Recognition of a high-affinity phosphotyrosyl peptide by the Src homology-2 domain of p56lck. Nature 362, 87–91 (1993).
Lawrence, M.C. & Colman, P.M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Hantschel, O. et al. A myristoyl/phosphotyrosine switch regulates c-Abl. Cell 112, 845–857 (2003).
Patwardhan, P. & Miller, W.T. Processive phosphorylation: mechanism and biological importance. Cell. Signal. 19, 2218–2226 (2007).
Schaller, M.D. & Schaefer, E.M. Multiple stimuli induce tyrosine phosphorylation of the Crk-binding sites of paxillin. Biochem. J. 360, 57–66 (2001).
Mayer, B.J., Hirai, H. & Sakai, R. Evidence that SH2 domains promote processive phosphorylation by protein-tyrosine kinases. Curr. Biol. 5, 296–305 (1995).
Ye, D., Wolff, N., Li, L., Zhang, S. & Ilaria, R.L. Jr. STAT5 signaling is required for the efficient induction and maintenance of CML in mice. Blood 107, 4917–4925 (2006).
Nieborowska-Skorska, M. et al. Signal transducer and activator of transcription (STAT)5 activation by BCR/ABL is dependent on intact Src homology (SH)3 and SH2 domains of BCR/ABL and is required for leukemogenesis. J. Exp. Med. 189, 1229–1242 (1999).
Schoepfer, J. et al. Highly potent inhibitors of the Grb2–SH2 domain. Bioorg. Med. Chem. Lett. 9, 221–226 (1999).
Poy, F. et al. Crystal structures of the XLP protein SAP reveal a class of SH2 domains with extended, phosphotyrosine-independent sequence recognition. Mol. Cell 4, 555–561 (1999).
Bae, J.H. et al. The selectivity of receptor tyrosine kinase signaling is controlled by a secondary SH2 domain binding site. Cell 138, 514–524 (2009).
Fellouse, F.A. et al. High-throughput generation of synthetic antibodies from highly functional minimalist phage-displayed libraries. J. Mol. Biol. 373, 924–940 (2007).
Koide, A., Gilbreth, R.N., Esaki, K., Tereshko, V. & Koide, S. High-affinity single-domain binding proteins with a binary-code interface. Proc. Natl. Acad. Sci. USA 104, 6632–6637 (2007).
Fellouse, F.A. et al. Molecular recognition by a binary code. J. Mol. Biol. 348, 1153–1162 (2005).
Fellouse, F.A., Barthelemy, P.A., Kelley, R.F. & Sidhu, S.S. Tyrosine plays a dominant functional role in the paratope of a synthetic antibody derived from a four amino acid code. J. Mol. Biol. 357, 100–114 (2006).
Hoppe-Seyler, F., Crnkovic-Mertens, I., Tomai, E. & Butz, K. Peptide aptamers: specific inhibitors of protein function. Curr. Mol. Med. 4, 529–538 (2004).
Mayer, B.J., Jackson, P.K., Van Etten, R.A. & Baltimore, D. Point mutations in the abl SH2 domain coordinately impair phosphotyrosine binding in vitro and transforming activity in vivo. Mol. Cell. Biol. 12, 609–618 (1992).
Skorski, T. et al. Transformation of hematopoietic cells by BCR/ABL requires activation of a PI-3k/Akt-dependent pathway. EMBO J. 16, 6151–6161 (1997).
Koide, A. & Koide, S. Monobodies: antibody mimics based on the scaffold of the fibronectin type III domain. Methods Mol. Biol. 352, 95–109 (2007).
Machida, K. et al. High-throughput phosphotyrosine profiling using SH2 domains. Mol. Cell 26, 899–915 (2007).
Acknowledgements
We thank R. Gilbreth and E. Duguid for assistance with X-ray structure determination, E. Duguid and C. He for assistance with DNA synthesis, M. Ciaccio, R. Gilbreth, P. Nash, B. Liu and A.A. Kossiakoff for discussion, the staff of the Life Sciences Collaborative Access Team (LS-CAT) beamline at the Advanced Photon Source and the University of Chicago DNA Sequencing Core facility for technical support, A.A. Kossiakoff for access to the BIAcore instrument, A. Müller, N. Venturini and M. Planyavsky for expert assistance with the MS analysis and the Structural Genomics Consortium for making the SH2 vectors available through Open Biosystems. This work was supported by US National Institutes of Health grants R01-GM72688, U54-GM74946 and R21-CA132700 to S.K., by the University of Chicago Cancer Research Center and by the Austrian Academy of Sciences. J.W. was supported in part by US National Institutes of Health grant 5T32GM07281-33 and European Molecular Biology Organization fellowship ASTF 293.00-2009. Use of the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-06CH11357. Use of the LS-CAT Sector 21 was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor for the support of this research program (grant 085P1000817).
Author information
Authors and Affiliations
Contributions
J.W., A.K. and S.K. designed phage-display library, selection and biophysical characterization; A.K. and J.W. optimized phage-display methods; J.W. performed selections, biophysical characterization, crystallization and structure determination; J.B. and R.B.J. designed and made SH2 microarrays; J.W. and J.B. conducted microarray experiments; J.W., F.G., O.H. and G.S.F. designed cellular experiments and kinase assays; J.W., O.H., F.G. and I.K. conducted cellular studies and kinase assays; K.L.B. conducted MS and data analysis. J.W., O.H., F.G., R.B.J., G.S.F. and S.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Tables 1–3, Supplementary Data and Supplementary Methods (PDF 626 kb)
Rights and permissions
About this article
Cite this article
Wojcik, J., Hantschel, O., Grebien, F. et al. A potent and highly specific FN3 monobody inhibitor of the Abl SH2 domain. Nat Struct Mol Biol 17, 519–527 (2010). https://doi.org/10.1038/nsmb.1793
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1793
This article is cited by
-
Adaptable, turn-on maturation (ATOM) fluorescent biosensors for multiplexed detection in cells
Nature Methods (2023)
-
Monobody adapter for functional antibody display on nanoparticles for adaptable targeted delivery applications
Nature Communications (2022)
-
Tuning SAS-6 architecture with monobodies impairs distinct steps of centriole assembly
Nature Communications (2021)
-
Selective and noncovalent targeting of RAS mutants for inhibition and degradation
Nature Communications (2021)
-
Development of light-responsive protein binding in the monobody non-immunoglobulin scaffold
Nature Communications (2020)