Abstract
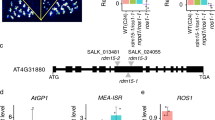
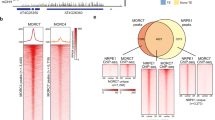
DNA methylation is an epigenetic mark affecting genes and transposons. Screening for mutants that fail to establish DNA methylation yielded two we termed “involved in de novo” (idn) 1 and 2. IDN1 encodes DMS3, an SMC-related protein, and IDN2 encodes a previously unknown double-stranded RNA–binding protein with homology to SGS3. IDN1 and IDN2 control de novo methylation and small interfering RNA (siRNA)-mediated maintenance methylation and are components of the RNA-directed DNA methylation pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Chan, S.W. et al. Science 303, 1336 (2004).
Cao, X. & Jacobsen, S.E. Curr. Biol. 12, 1138–1144 (2002).
Henderson, I.R. & Jacobsen, S.E. Nature 447, 418–424 (2007).
Li, C.F. et al. Cell 126, 93–106 (2006).
Pontes, O. et al. Cell 126, 79–92 (2006).
Wierzbicki, A.T., Haag, J.R. & Pikaard, C.S. Cell 135, 635–648 (2008).
Soppe, W.J. et al. Mol. Cell 6, 791–802 (2000).
Henderson, I.R. & Jacobsen, S.E. Genes Dev. 22, 1597–1606 (2008).
Cao, X. & Jacobsen, S.E. Proc. Natl. Acad. Sci. USA 99 Suppl 4, 16491–16498 (2002).
Kanno, T. et al. Nat. Genet. 40, 670–675 (2008).
Bateman, A. BMC Bioinformatics 3, 21 (2002).
Mourrain, P. et al. Cell 101, 533–542 (2000).
Zhang, D. & Trudeau, V.L. Cell Cycle 7, 2268–2270 (2008).
Fukunaga, R. & Doudna, J.A. EMBO J. 28, 545–555 (2009).
Acknowledgements
We thank J. Goodrich and D. Schubert for kindly providing T-DNA homozygous lines. We thank members of the Jacobsen laboratory for supportive discussions. Jacobsen lab research was supported by US National Institutes of Health grant GM60398. I.A. was supported by a postdoctoral fellowship from the Ministerio de Educacion y Ciencia. S.E.J. and J.C. are investigators of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.C.M. and J.C. provided the majority of the insertion mutagenized population. I.A. and S.E.J. designed the experiments. I.A. performed the experiments and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 170 kb)
Rights and permissions
About this article
Cite this article
Ausin, I., Mockler, T., Chory, J. et al. IDN1 and IDN2 are required for de novo DNA methylation in Arabidopsis thaliana. Nat Struct Mol Biol 16, 1325–1327 (2009). https://doi.org/10.1038/nsmb.1690
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1690
This article is cited by
-
Multi-omics data integration reveals link between epigenetic modifications and gene expression in sugar beet (Beta vulgaris subsp. vulgaris) in response to cold
BMC Genomics (2022)
-
Dynamics and function of DNA methylation in plants
Nature Reviews Molecular Cell Biology (2018)
-
RNA-directed DNA methylation involves co-transcriptional small-RNA-guided slicing of polymerase V transcripts in Arabidopsis
Nature Plants (2018)
-
AtMBD6, a methyl CpG binding domain protein, maintains gene silencing in Arabidopsis by interacting with RNA binding proteins
Journal of Biosciences (2017)
-
Two domain-disrupted hda6 alleles have opposite epigenetic effects on transgenes and some endogenous targets
Scientific Reports (2015)