Abstract
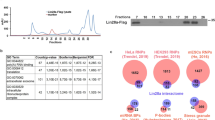
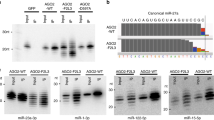
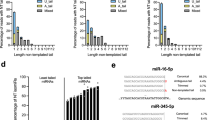
Lin28 and Lin28B, two developmentally regulated RNA-binding proteins and likely proto-oncogenes, selectively inhibit the maturation of let-7 family microRNAs (miRNAs) in embryonic stem cells and certain cancer cell lines. Moreover, let-7 precursors (pre–let-7) were previously found to be terminally uridylated in a Lin28-dependent fashion. Here we identify Zcchc11 (zinc finger, CCHC domain containing 11) as the 3′ terminal uridylyl transferase (TUTase) responsible for Lin28-mediated pre–let-7 uridylation and subsequent blockade of let-7 processing in mouse embryonic stem cells. We demonstrate that Zcchc11 activity is UTP-dependent, selective for let-7 and recruited by Lin28. Furthermore, knockdown of either Zcchc11 or Lin28, or overexpression of a catalytically inactive TUTase, relieves the selective inhibition of let-7 processing and leads to the accumulation of mature let-7 miRNAs and repression of let-7 target reporter genes. Our results establish a role for Zcchc11-catalyzed pre–let-7 uridylation in the control of miRNA biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Winter, J., Jung, S., Keller, S., Gregory, R.I. & Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 11, 228–234 (2009).
Blakaj, A. & Lin, H. Piecing together the mosaic of early mammalian development through microRNAs. J. Biol. Chem. 283, 9505–9508 (2008).
Esquela-Kerscher, A. & Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006).
Hagan, J.P. & Croce, C.M. MicroRNAs in carcinogenesis. Cytogenet. Genome Res. 118, 252–259 (2007).
Büssing, I., Slack, F.J. & Grosshans, H. let-7 microRNAs in development, stem cells and cancer. Trends Mol. Med. 14, 400–409 (2008).
Viswanathan, S.R., Daley, G.Q. & Gregory, R.I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre–let-7 maturation during neural stem-cell commitment. Nat. Cell Biol. 10, 987–993 (2008).
Newman, M.A., Thomson, J.M. & Hammond, S.M. Lin-28 interaction with the let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008).
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor microRNA. Mol. Cell 32, 276–284 (2008).
Piskounova, E. et al. Determinants of microRNA processing inhibition by the developmentally regulated RNA-binding protein Lin28. J. Biol. Chem. 283, 21310–21314 (2008).
Ambros, V. & Horvitz, H.R. Heterochronic mutants of the nematode Caenorhabditis elegans. Science 226, 409–416 (1984).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
West, J.A. et al. A role for Lin28 in primordial germ-cell development and germ-cell malignancy. Nature 460, 909–913 (2009).
Lettre, G. et al. Identification of ten loci associated with height highlights new biological pathways in human growth. Nat. Genet. 40, 584–591 (2008).
Sulem, P. et al. Genome-wide association study identifies sequence variants on 6q21 associated with age at menarche. Nat. Genet. 41, 734–738 (2009).
Ong, K.K. et al. Genetic variation in LIN28B is associated with the timing of puberty. Nat. Genet. 41, 729–733 (2009).
Stolk, L. et al. Loci at chromosomes 13, 19 and 20 influence age at natural menopause. Nat. Genet. 41, 645–647 (2009).
He, C. et al. Genome-wide association studies identify loci associated with age at menarche and age at natural menopause. Nat. Genet. 41, 724–728 (2009).
Perry, J.R. et al. Meta-analysis of genome-wide association data identifies two loci influencing age at menarche. Nat. Genet. 41, 648–650 (2009).
Dangi-Garimella, S. et al. Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let-7. EMBO J. 28, 347–358 (2009).
Guo, Y. et al. Identification and characterization of lin-28 homolog B (LIN28B) in human hepatocellular carcinoma. Gene 384, 51–61 (2006).
Chang, T.C. et al. Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation. Proc. Natl. Acad. Sci. USA 106, 3384–3389 (2009).
Viswanathan, S.R. et al. LIN28 promotes transformation and is associated with advanced human malignancies. Nat. Genet. 41, 843–848 (2009).
Martin, G. & Keller, W. RNA-specific ribonucleotidyl transferases. RNA 13, 1834–1849 (2007).
Wilusz, C.J. & Wilusz, J. New ways to meet your (3′) end oligouridylation as a step on the path to destruction. Genes Dev. 22, 1–7 (2008).
Trippe, R. et al. Identification, cloning, and functional analysis of the human U6 snRNA-specific terminal uridylyl transferase. RNA 12, 1494–1504 (2006).
Mullen, T.E. & Marzluff, W.F. Degradation of histone mRNA requires oligouridylation followed by decapping and simultaneous degradation of the mRNA both 5′ to 3′ and 3′ to 5′. Genes Dev. 22, 50–65 (2008).
Kwak, J.E. & Wickens, M. A family of poly(U) polymerases. RNA 13, 860–867 (2007).
Tomecki, R., Dmochowska, A., Gewartowski, K., Dziembowski, A. & Stepien, P.P. Identification of a novel human nuclear-encoded mitochondrial poly(A) polymerase. Nucleic Acids Res. 32, 6001–6014 (2004).
Lehrbach N.J. et al. LIN-28 and the poly(U) polymerase PUP-2 regulate let-7 microRNA processing in Caenorhabditis elegans. Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.1675 (27 August 2009).
Rissland, O.S. & Norbury, C.J. Decapping is preceded by 3′ uridylation in a novel pathway of bulk mRNA turnover. Nat. Struct. Mol. Biol. 16, 616–623 (2009).
Acknowledgements
We are grateful to S. Viswanathan and G. Daley (Children's Hospital Boston) for providing Flag-Lin28–expressing ES cells, H. Rusk and R. LaPierre for technical assistance and the Proteomics Center at Children's Hospital Boston for expertise in the microcapillary HPLC and MS. We thank E. Miska (University of Cambridge) for helpful discussions. R.I.G. is supported by laboratory start-up funds from The Children's Hospital Boston and grants from the US National Institute of General Medical Sciences (NIGMS) (1R01GM086386-01A1), The Harvard Stem Cell Institute, The March of Dimes Basil O'Conner award and the Emerald Foundation. R.I.G. is supported by The Pew Scholars Program in the Biomedical Sciences.
Author information
Authors and Affiliations
Contributions
J.P.H. and E.P. performed all experiments; J.P.H., E.P. and R.I.G. designed all experiments, analyzed data and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–4 (PDF 1178 kb)
Rights and permissions
About this article
Cite this article
Hagan, J., Piskounova, E. & Gregory, R. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nat Struct Mol Biol 16, 1021–1025 (2009). https://doi.org/10.1038/nsmb.1676
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1676
This article is cited by
-
Lin28 affects the proliferation and osteogenic differentiation of human dental pulp stem cells by directly inhibiting let-7b maturation
BDJ Open (2024)
-
TENT2, TUT4, and TUT7 selectively regulate miRNA sequence and abundance
Nature Communications (2022)
-
Overexpression of Lin28A in neural progenitor cells in vivo does not lead to brain tumor formation but results in reduced spine density
Acta Neuropathologica Communications (2021)
-
The TUTase URT1 connects decapping activators and prevents the accumulation of excessively deadenylated mRNAs to avoid siRNA biogenesis
Nature Communications (2021)
-
BCDIN3D RNA methyltransferase stimulates Aldolase C expression and glycolysis through let-7 microRNA in breast cancer cells
Oncogene (2021)