Abstract
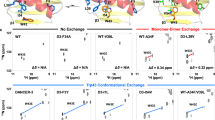
The synchronization (correlation) of conformational fluctuations in folded proteins may influence the rates of enzyme catalysis and ligand binding as well as the stabilities of native proteins and their complexes. However, experimental detection of correlated motions remains difficult. Herein, we present an analysis of the covariation of NMR-derived backbone dynamical parameters among a family of ten mutants of a small protein. Both the spatial restriction and the time scales of backbone motions exhibit a higher degree of covariation than would be expected if the internal motions of each group were independent, providing experimental support for correlated dynamics. Application of this approach to other proteins may reveal dynamical correlations that influence catalysis, ligand-binding and/or protein stability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Knapp, M.J., Rickert, K. & Klinman, J.P. Temperature-dependent isotope effects in soybean lipoxygenase-1: correlating hydrogen tunneling with protein dynamics. J. Am. Chem. Soc. 124, 3865–3874 (2002).
Balabin, I.A. & Onuchic, J.N. Dynamically controlled protein tunneling paths in photosynthetic reaction centers. Science 290, 114–117 (2000).
Bruice, T.C. & Benkovic, S.J. Chemical basis for enzyme catalysis. Biochemistry 39, 6267–6274 (2000).
Osborne, M.J., Schnell, J., Benkovic, S.J., Dyson, H.J. & Wright, P.E. Backbone dynamics in dihydrofolate reductase complexes: role of loop flexibility in the catalytic mechanism. Biochemistry 40, 9846–9859 (2001).
Matko, J. et al. Correlation between activity and dynamics of the protein matrix of phosphorylase b. Biochemistry 19, 5782–5786 (1980).
Wand, A.J. Dynamic activation of protein function: a view emerging from NMR spectroscopy. Nat. Struct. Biol. 8, 926–931 (2001).
Wand, A.J. On the dynamic origins of allosteric activation. Science 293, 1395 (2001).
Viswanathan, M., Linthicum, D.S. & Subramaniam, S. Analysis of correlated motion in antibody combining sites from molecular dynamics simulations. Methods 20, 362–371 (2000).
Palmer, A.G. III. NMR probes of molecular dynamics: overview and comparison with other techniques. Annu. Rev. Biophys. Biomol. Struct. 30, 129–155 (2001).
Kay, L.E. Protein dynamics from NMR. Nat. Struct. Biol. Suppl. 5, 513–517 (1998).
Fischer, M.W.F., Majumdar, A. & Zuiderweg, E.R.P. Protein NMR relaxation: theory, applications and outlook. Prog. Nucl. Magn. Reson. Spectros. 33, 207–272 (1998).
Stone, M.J. NMR relaxation studies of the role of conformational entropy in protein stability and ligand binding. Acc. Chem. Res. 34, 379–388 (2001).
Kay, L.E., Muhandiram, D.R., Wolf, G., Shoelson, S.E. & Forman-Kay, J.D. Correlation between binding and dynamics at SH2 domain interfaces. Nat. Struct. Biol. 5, 156–163 (1998).
Maler, L., Blankenship, J., Rance, M. & Chazin, W.J. Site-site communication in the EF-hand Ca2+-binding protein calbindin D9k. Nat. Struct. Biol. 7, 245–250 (2000).
Lee, A.L. & Wand, A.J. Microscopic origins of entropy, heat capacity and the glass transition in proteins. Nature 411, 501–504 (2001).
Li, Z., Raychaudhuri, S. & Wand, A.J. Insights into the local residual entropy of proteins provided by NMR relaxation. Protein Sci. 5, 2647–2650 (1996).
Akke, M., Bruschweiler, R. & Palmer, A.G. III. NMR order parameters and free energy: an analytical approach and its application to cooperative Ca2+ binding by calbindin D9k . J. Am. Chem. Soc. 115, 9832–9833 (1993).
Yang, D.W. & Kay, L.E. Contributions to conformational entropy arising from bond vector fluctuations measured from NMR-derived order parameters: application to protein folding. J. Mol. Biol. 263, 369–382 (1996).
Boyle, M. Bacterial Immunoglobulin-Binding Proteins (Academic, San Diego, California, USA, 1990).
Smith, C.K., Withka, J.M. & Regan, L. A thermodynamic scale for the β-sheet forming tendencies of the amino acids. Biochemistry 33, 5510–5517 (1994).
Stone, M.J., Gupta, S., Snyder, N. & Regan, L. Comparison of protein backbone entropy and β-sheet stability: NMR-derived dynamics of protein G B1 domain mutants. J. Am. Chem. Soc. 123, 185–186 (2001).
Sheinerman, F.B. & Brooks, C.L. Calculations on folding of segment B1 of Streptococcal protein G. J. Mol. Biol. 278, 439–456 (1998).
Blanco, F.J. & Serrano, L. Folding of protein G B1 domain studied by the conformational characterization of fragments comprising its secondary structure elements. Eur. J. Biochem. 230, 634–649 (1998).
Lee, A.L., Sharp, K.A., Kranz, J.K., Song, X.-J. & Wand, A.J. Temperature dependence of the internal dynamics of a calmodulin-peptide complex. Biochemistry 41, 13814–13825 (2002).
Daragan, V.A. & Mayo, K.H. A simple approach to analyzing protein side-chain dynamics from 13C NMR relaxation data. J. Magn. Reson. 130, 329–334 (1998).
Lemaster, D.M. NMR relaxation order parameter analysis of the dynamics of protein side chains. J. Am. Chem. Soc. 121, 1726–1742 (1999).
Goodman, J.L., Pagel, M.D. & Stone, M.J. Relationships between protein structure and dynamics from a database of NMR-derived backbone order parameters. J. Mol. Biol. 295, 963–978 (2000).
Fischer, M.W., Zeng, L., Majumdar, A. & Zuiderweg, E.R. Characterizing semilocal motions in proteins by NMR relaxation studies. Proc. Natl. Acad. Sci. USA 95, 8016–8019 (1998).
Prompers, J.J. & Bruschweiler, R. General framework for studying the dynamics of folded and nonfolded proteins by NMR relaxation spectroscopy and MD simulation. J. Am. Chem. Soc. 124, 4522–4534 (2002).
Seewald, M.J. et al. The role of backbone conformational heat capacity in protein stability: temperature dependent dynamics of the B1 domain of Streptococcal protein G. Protein Sci. 9, 1177–1193 (2000).
Gronenborn, A.M. et al. A novel, highly stable fold of the immunoglobulin binding domain of Streptococcal protein G. Science 253, 657–661 (1991).
Kuszewski, J., Gronenborn, A.M. & Clore, G.M. Improving the packing and accuracy of NMR structures with a pseudopotential for the radius of gyration. J. Am. Chem. Soc. 121, 2337–2338 (1999).
Clore, G.M., Driscoll, P.C., Wingfield, P.T. & Gronenborn, A.M. Analysis of the backbone dynamics of interleukin-1β using two-dimensional inverse detected heteronuclear 15N-1H NMR spectroscopy. Biochemistry 29, 7387–7401 (1990).
Acknowledgements
We thank J.L. Vaughn for discussions. This work was supported by grants from the US National Science Foundation and the American Chemical Society Petroleum Research Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Mayer, K., Earley, M., Gupta, S. et al. Covariation of backbone motion throughout a small protein domain. Nat Struct Mol Biol 10, 962–965 (2003). https://doi.org/10.1038/nsb991
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb991
This article is cited by
-
Globally correlated conformational entropy underlies positive and negative cooperativity in a kinase’s enzymatic cycle
Nature Communications (2019)
-
Understanding biomolecular motion, recognition, and allostery by use of conformational ensembles
European Biophysics Journal (2011)