Abstract
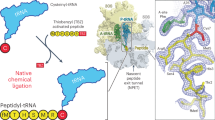
Nascent proteins emerge out of ribosomes through an exit tunnel, which was assumed to be a firmly built passive path. Recent biochemical results, however, indicate that the tunnel plays an active role in sequence-specific gating of nascent chains and in responding to cellular signals. Consistently, modulation of the tunnel shape, caused by the binding of the semi-synthetic macrolide troleandomycin to the large ribosomal subunit from Deinococcus radiodurans, was revealed crystallographically. The results provide insights into the tunnel dynamics at high resolution. Here we show that, in addition to the typical steric blockage of the ribosomal tunnel by macrolides, troleandomycin induces a conformational rearrangement in a wall constituent, protein L22, flipping the tip of its highly conserved β-hairpin across the tunnel. On the basis of mutations that alleviate elongation arrest, the tunnel motion could be correlated with sequence discrimination and gating, suggesting that specific arrest motifs within nascent chain sequences may induce a similar gating mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Milligan, R.A. & Unwin, P.N. Location of exit channel for nascent protein in 80S ribosome. Nature 319, 693–695 (1986).
Yonath, A., Leonard, K.R. & Wittmann, H.G. A tunnel in the large ribosomal subunit revealed by three-dimensional image reconstruction. Science 236, 813–816 (1987).
Gabashvili, I.S. et al. The polypeptide tunnel system in the ribosome and its gating in erythromycin resistant mutants of L4 and L22. Mol. Cell 8, 181–188 (2001).
Tenson, T. & Ehrenberg, M. Regulatory nascent peptides in the ribosomal tunnel. Cell 108, 591–594 (2002).
Morris, D.R. & Geballe, A.P. Upstream open reading frames as regulators of mRNA translation. Mol. Cell. Biol. 20, 8635–8642 (2000).
Sarker, S., Rudd, K.E. & Oliver, D. Revised translation start site for secM defines an atypical signal peptide that regulates E. coli secA expression. J. Bacteriol. 182, 5592–5595 (2000).
Nakatogawa, H. & Ito, K. The ribosomal exit tunnel functions as a discriminating gate. Cell 108, 629–636 (2002).
Gong, F. & Yanofsky, C. Instruction of translating ribosome by nascent peptide. Science 297, 1864–1867 (2002).
Stroud, R.M. & Walter, P. Signal sequence recognition and protein targeting. Curr. Opin. Struct. Biol. 9, 754–759 (1999).
Nissen, P., Hansen, J., Ban, N., Moore, P.B. & Steitz, T.A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Harms, J. et al. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell 107, 679–688 (2001).
Unge, J. et al. The crystal structure of ribosomal protein L22 from Thermus thermophilus: insights into the mechanism of erythromycin resistance. Structure 6, 1577–1586 (1998).
Schluenzen, F. et al. Structural basis for the interaction of antibiotics with the peptidyl transferase centre in eubacteria. Nature 413, 814–821 (2001).
Hansen, J.L. et al. The structures of four macrolide antibiotics bound to the large ribosomal subunit. Mol. Cell 10, 117–128 (2002).
Schluenzen, F. et al. Structural basis for the antibiotic activity of ketolides and azalides. Structure 11, 329–338 (2003).
Periti, P., Tonelli, F., Mazzei, T. & Ficari, F. Antimicrobial chemoimmunoprophylaxis in colorectal surgery with cefotetan and thymostimulin: prospective, controlled multicenter study. Italian Study Group on Antimicrobial Prophylaxis in Abdominal Surgery. J. Chemother. 5, 37–42 (1993).
Chepkwony, H.K., Roets, E. & Hoogmartens, J. Liquid chromatography of troleandomycin. J. Chromatogr. A 914, 53–58 (2001).
Tenson, T. & Mankin, A.S. Short peptides conferring resistance to macrolide antibiotics. Peptides 22, 1661–1668 (2001).
Verdier, L., Gharbi-Benarous, J., Bertho, G., Mauvais, P. & Girault, J.P. Antibiotic resistance peptides: interaction of peptides conferring macrolide and ketolide resistance with Staphylococcus aureus ribosomes: conformation of bound peptides as determined by transferred NOE experiments. Biochemistry 41, 4218–4229 (2002).
Douthwaite, S., Hansen, L.H. & Mauvais, P. Macrolide-ketolide inhibition of MLS-resistant ribosomes is improved by alternative drug interaction with domain II of 23S rRNA. Mol. Microbiol. 36, 183–193 (2000).
Weisblum, B. Erythromycin resistance by ribosome modification. Antimicrob. Agents Chemother. 39, 577–585 (1995).
Hansen, L.H., Mauvais, P. & Douthwaite, S. The macrolide-ketolide antibiotic binding site is formed by structures in domains II and V of 23S ribosomal RNA. Mol. Microbiol. 31, 623–631 (1999).
Xiong, L., Shah, S., Mauvais, P. & Mankin, A.S. A ketolide resistance mutation in domain II of 23S rRNA reveals the proximity of hairpin 35 to the peptidyl transferase centre. Mol. Microbiol. 31, 633–639 (1999).
Goldman, R.C., Fesik, S.W. & Doran, C.C. Role of protonated and neutral forms of macrolides in binding to ribosomes from Gram-positive and Gram-negative bacteria. Antimicrob. Agents Chemother. 34, 426–431 (1990).
Davydova, N., Streltsov, V., Wilce, M., Liljas, A. & Garber, M. L22 ribosomal protein and effect of its mutation on ribosome resistance to erythromycin. J. Mol. Biol. 322, 635–644 (2002).
Liao, S., Lin, J., Do, H. & Johnson, A.E. Both lumenal and cytosolic gating of the aqueous ER translocon pore are regulated from inside the ribosome during membrane protein integration. Cell 90, 31–41 (1997).
Stern, S., Moazed, D. & Noller, H.F. Structural analysis of RNA using chemical and enzymatic probing monitored by primer extension. Methods Enzymol. 164, 481–489 (1988).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Bailey, S. The CCP4 suite — programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Flocco, M.M. & Mowbray, S.L. Cα-based torsion angles: a simple tool to analyze protein conformational changes. Protein Sci. 4, 2118–2122 (1995).
Carson, M. Ribbons. Methods Enzymol. 277, 493–505 (1997).
Wallace, A.C., Laskowski, R.A. & Thornton, J.M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995).
Acknowledgements
We thank G. Glaser, R. Zarivach and P. Fucini for critical discussions; I. Agmon, R. Albrecht, C. Liebe, A. Wolff, H. Bartels, W.S. Bennett, E. Ben-Zeev, C. Glotz, H.A.S. Hansen, M. Kessler and A. McLeod for participating in this work; and the staff at ID19/SBC/APS and ID29/ESRF/EMBL. The Max-Planck-Society, the US National Institutes of Health, the German Science & Technology Ministry and the Kimmelman Center for Macromolecular Assembly provided support. A.Y. holds the Hellen and Martin Kimmel Professorial Chair.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Berisio, R., Schluenzen, F., Harms, J. et al. Structural insight into the role of the ribosomal tunnel in cellular regulation. Nat Struct Mol Biol 10, 366–370 (2003). https://doi.org/10.1038/nsb915
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb915
This article is cited by
-
A comparative genomics study on the effect of individual amino acids on ribosome stalling
BMC Genomics (2015)
-
Chemical and structural biology of nucleic acids and protein-nucleic acid complexes for novel drug discovery
Science China Chemistry (2011)