Abstract
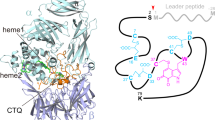
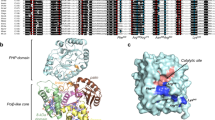
The PutA flavoprotein from Escherichia coli plays multiple roles in proline catabolism by functioning as a membrane-associated bi-functional enzyme and a transcriptional repressor of proline utilization genes. The human homolog of the PutA proline dehydrogenase (PRODH) domain is critical in p53-mediated apoptosis and schizophrenia. Here we report the crystal structure of a 669-residue truncated form of PutA that shows both PRODH and DNA-binding activities, representing the first structure of a PutA protein and a PRODH enzyme from any organism. The structure is a domain-swapped dimer with each subunit comprising three domains: a helical dimerization arm, a 120-residue domain containing a three-helix bundle similar to that in the helix-turn-helix superfamily of DNA-binding proteins and a β/α-barrel PRODH domain with a bound lactate inhibitor. Analysis of the structure provides insight into the mechanism of proline oxidation to pyrroline-5-carboxylate, and functional studies of a mutant protein suggest that the DNA-binding domain is located within the N-terminal 261 residues of E. coli PutA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brown, E. & Wood, J.M. Redesigned purification yields a fully functional PutA protein dimer from Escherichia coli. J. Biol. Chem. 267, 13086–13092 (1992).
Menzel, R. & Roth, J. Purification of the putA gene product. J. Biol. Chem. 256, 9755–9761 (1981).
Scarpulla, R.C. & Soffer, R.L. Membrane-bound proline dehydrogenase from Escherichia coli. J. Biol. Chem. 253, 5997–6001 (1978).
Maloy, S. & Roth, J.R. Regulation of proline utilization in Salmonella typhimurium: characterization of put:Mu L(Ap, lac) operon fusions. J. Bacteriol. 154, 561–568 (1983).
Menzel, R. & Roth, J. Regulation of genes for proline utilization in Salmonella typhimurium: autogenous repression by the putA gene product. J. Mol. Biol. 148, 21–44 (1981).
Wood, J.M. Genetics of L-proline utilization in Eschericia coli. J. Bacteriol. 146, 895–901 (1981).
Becker, D.F. & Thomas, E.A. Redox properties of the PutA protein from Escherichia coli and the influence of the flavin redox state on PutA-DNA interactions. Biochemistry 40, 4714–4722 (2001).
Ostrovsky de Spicer, P. & Maloy, S. PutA protein, a membrane-associated flavin dehydrogenase, acts as a redox-dependent transcriptional regulator. Proc. Natl. Acad. Sci. USA. 90, 4295–4298 (1993).
Surber, M.W. & Maloy, S. Regulation of flavin dehydrogenase compartmentalization: requirements for PutA-membrane association in Salmonella typhimurium. Biochim. Biophys. Acta 1421, 5–18 (1999).
Wood, J. Membrane association of proline dehydrogenase in Escherichia coli is redox dependent. Proc. Natl. Acad. Sci. USA 84, 373–377 (1987).
Polyak, K., Xia, Y., Zweier, J.L., Kinzler, K.W. & Vogelstein, B. A model for p53-induced apoptosis. Nature 389, 300–305 (1997).
Donald, S.P. et al. Proline oxidase, encoded by p53-induced gene-6, catalyzes the generation of proline-dependent reactive oxygen species. Cancer Res. 61, 1810–1815 (2001).
Liu, H. et al. Genetic variation at the 22q11 PRODH2/DGCR6 locus presents an unusual pattern and increases susceptibility to schizophrenia. Proc. Natl. Acad. Sci. USA 99, 3717–3722 (2002).
Chakravarti, A. A compelling genetic hypothesis for a complex disease: PRODH2/DGCR6 variation leads to schizophrenia susceptibility. Proc. Natl. Acad. Sci. USA 99, 4755–4756 (2002).
Vinod, M.P., Bellur, P. & Becker, D.F. Electrochemical and functional characterization of the proline dehydrogenase domain of the PutA flavoprotein from Escherichia coli. Biochemistry 41, 6525–6532 (2002).
Ling, M., Allen, S.W. & Wood, J.M. Sequence analysis edentifies the proline dehydrogenase and pyrroline-5-carboxylate dehydrogenase domains of the multifunctional Escherichia coli PutA protein. J. Mol. Biol. 245, 950–956 (1994).
Bennett, M.J., Schlunegger, M.P. & Eisenberg, D. 3D domain swapping: a mechanism for oligomer assembly. Protein Sci. 4, 2455–2468 (1995).
Guenther, B.D. et al. The structure and properties of methylenetetrahydrofolate reductase from Escherichia coli suggest how folate ameliorates human hyperhomocysteinemia. Nat. Struct. Biol. 6, 359–365 (1999).
Umhau, S. et al. The X-ray structure of D-amino acid oxidase at very high resolution identifies the chemical mechanism of flavin-dependent substrate dehydrogenation. Proc. Natl. Acad. Sci. USA 97, 12463–12468 (2000).
Mattevi, A. et al. Crystal structure of D-amino acid oxidase: a case of active site mirror-image convergent evolution with flavocytochrome b2. Proc. Natl. Acad. Sci. USA 93, 7496–7501 (1996).
Pollegioni, L., Blodig, W. & Ghisla, S. On the mechanism of D-amino acid oxidase. Structure/linear free energy correlations and deuterium kinetic isotope effects using substituted phenylglycines. J. Biol. Chem. 272, 4924–4934 (1997).
Kurtz, K.A., Rishavy, M.A., Cleland, W.W. & Fitzpatrick, P.F. Nitrogen isotope effects as probes of the mechanism of D-amino acid oxidase. J. Am. Chem. Soc. 122, 12896–12897 (2000).
Surber, M.W. & Maloy, S. The PutA protein of Salmonella typhimurium catalyzes the two steps of proline degradation via a leaky channel. Arch. Biochem. Biophys. 354, 281–287 (1998).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Lewis, R.J. et al. The trans-activation domain of the sporulation response regulator Spo0A revealed by X-ray crystallography. Mol. Microbiol. 38, 198–212 (2000).
Baikalov, I. et al. Structure of the Escherichia coli response regulator NarL. Biochemistry 35, 11053–11061 (1996).
Liu, J. et al. Structure and function of Cdc6/Cdc18: implications for origin recognition and checkpoint control. Mol. Cell. 6, 637–648 (2000).
Brennan, R.G. & Matthews, B.W. Structural basis of DNA-protein recognition. Trends. Biochem. Sci. 1989, 286–290 (1989).
Nadaraia, S., Lee, Y.H., Becker, D.F. & Tanner, J.J. Crystallization and preliminary crystallographic analysis of the proline dehydrogenase domain of the multifunctional PutA flavoprotein from Escherichia coli. Acta Crystallogr. D 57, 1925–1927 (2001).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. X-PLOR Version 3.1: A System for X-ray Crystallography and NMR (Yale University Press, New Haven; 1992).
Noll, F. L-(+)-Lactate. in Methods of Enzymatic Analysis, Vol. 6 (eds. Bergmeyer, J. & Grassl, M.) Chap. 3.12, 582–588 (Verlag Chemie, Weinheim; Deerfield Beach; 1983).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D version 2.0: a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Read, R.J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42, 140–149 (1986).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. Model. 15, 132–134 (1997).
Acknowledgements
This research was supported by grants from the NIH (J.J.T. and D.F.B) and the NSF (D.F.B.). We thank L. Beamer for collecting the NaBr derivative data, L. Beamer and P. Tipton for helpful comments on the manuscript, and the beamline personnel at APS SBC19-ID, NSLS X8C and NSLS X12B for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lee, YH., Nadaraia, S., Gu, D. et al. Structure of the proline dehydrogenase domain of the multifunctional PutA flavoprotein. Nat Struct Mol Biol 10, 109–114 (2003). https://doi.org/10.1038/nsb885
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb885
This article is cited by
-
N-Propargylglycine: a unique suicide inhibitor of proline dehydrogenase with anticancer activity and brain-enhancing mitohormesis properties
Amino Acids (2021)
-
Proline dehydrogenase in cancer: apoptosis, autophagy, nutrient dependency and cancer therapy
Amino Acids (2021)
-
The Janus-like role of proline metabolism in cancer
Cell Death Discovery (2020)
-
Proline dehydrogenase from Thermus thermophilus does not discriminate between FAD and FMN as cofactor
Scientific Reports (2017)
-
Metabolic network capacity of Escherichia coli for Krebs cycle-dependent proline hydroxylation
Microbial Cell Factories (2015)