Abstract
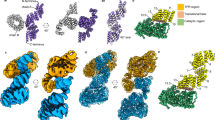
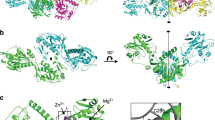
The high resolution crystal structure of human lysosomal aspartylglucosaminidase (AGA) has been determined. This lysosomal enzyme is synthesized as a single polypeptide precursor, which is immediately post-translationally cleaved into α- and β-subunits. Two α- and β-chains are found to pack together forming the final heterotetrameric structure. The catalytically essential residue, the N-terminal threonine of the β-chain is situated in the deep pocket of the funnel-shaped active site. On the basis of the structure of the enzyme–product complex we present a catalytic mechanism for this lysosomal enzyme with an exceptionally high pH optimum. The three-dimensional structure also allows the prediction of the structural consequences of human mutations resulting in aspartylglucosaminuria (AGU), a lysosomal storage disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Mononen, I., Fisher, K.J., Kaartinen, V. & Aronson, Jr, N. Aspartylglucosaminuria: protein chemistry and molecular biology of the most common lysosomal storage disorder of glycoprotein degradation. FASEB J. 7, 1247–1256 (1993).
Ikonen, E., Julkunen, I., Tollersrud, O.-K., Kalkkinen, N. & Peltonen, L. Lysosomal aspartylglucosaminidase is processed to the active subunit in the endoplasmic reticulum. EMBO J. 12, 295–302 (1993).
Fisher, K.J., Klein, M., Park, H., Vettese, M.B. & Aronson Jr, N.N. Post-translational processing and Thr206 are required for glycosylasparaginase activity. FEBS Lett. 323, 271–275 (1993).
Riikonen, A., Tikkanen, R., Jalanko, A. & Peltonen, L. Immediate interaction between the nascent subunits and two conserved amino acids Trp34 and Thr206 are needed for the catalytic activity of aspartylglucosaminidase. J. biol. Chem. 270, 4903–4907 (1995).
Ikonen, E. et al. Spectrum of mutations in aspartylglucosaminuria. Proc. natn. Acad. Sci. U.S.A. 88, 11222–11226 (1991).
Tikkanen, R., Rouvinen, J., Törrönen, A., Kalkkinen, N. & Peltonen, L. Large scale purification and preliminary X-ray diffraction studies of glycosylated human aspartylglucosaminidase. Proteins in the press.
Swain, A.L., Jaskólski, M., Housset, D., Rao, J.K.M. & Wlodawer, A. Crystal structure of Eschericia coli L-asparaginase, an enzyme used in cancer therapy. Proc. natn. Acad. Sci. U.S.A. 90, 1474–1478 (1993).
Miller, M., Rao, J.K.M., Wlodawer, A. & Gribskov, M.R. A left-handed crossover involved in amidohydrolase catalysis. Crystal structure of Erwinia chrysanthemi L-asparaginase with bound L-aspartate. FEBS Lett. 328, 275–279 (1993).
Norris, G.E., Stillman, T.J., Anderson, B.F. & Baker, E.N. The three-dimensional structure of PNGase F, a glycosylasparaginase from Flavobacterium meningosepticum. Structure 2, 1049–1059 (1994).
Kuhn, P., Tarentino, A.L., Plummer, Jr, T.H. & Van Roey, P. Crystal structure of peptide N4-(N-acetyl-β-D-glucosaminyl)asparagine amidase F at 2.2-Å resolution. Biochemistry 33, 11699–11706 (1994).
Smith, J.L. et al. Structure of the allosteric regulatory enzyme of purine biosynthesis. Science 264, 1427–1433 (1994).
Duggleby, H.J. et al. Penicillin acylase has a single-amino-acid catalytic centre. Nature 373, 264–268 (1995).
Löwe, J. et al. Crystal structure of the 205 proteasome from the archaeon T. acidophilum at 3.4 Å resolution. Science 268, 533–539 (1995).
Ikonen, E. et al. Aspartylglucosaminuria: cDNA encoding human aspartylglucosaminidase and the missense mutation causing the disease. EMBO J. 10, 51–58 (1991).
Park, H., Fisher, K.J. & Aronson, Jr, N.N. Genomic structure of human lysosomal glycosylasparaginase. FEBS Lett. 288, 168–172 (1991).
Tenhunen, K., Laan, M., Manninen, T., Palotie, A., Peltonen, L. & Jalanko, A. Molecular cloning, chromosomal assignment and expression of the mouse aspartylglucosaminidase gene. Genomics in the press.
Tarentino, A.L., Quinones, G., Hauer, C.R., Changchien, L.-M. & Plummer Jr, T.H. Molecular cloning and sequence analysis of Flavobacterium meningosepticum glycosylasparaginase: a single gene encodes the α and β subunits. Archs Biochem. Biophys. 316, 399–406 (1995).
Lough, T.J. et al. The isolation and characterisation of a cDNA clone encoding L-asparaginase from developing seeds of lupin (Lupinus arboreus). Pl. molec. Biol. 19, 391–399 (1992).
Kaartinen, V., Mononen, T., Laatikainen, R. & Mononen, I. Substrate specificity and reaction mechanism of human glycosasparaginase. J. biol. Chem. 267, 6855–6858 (1992).
Vyas, N.K. Atomic features of protein-carbohydrate interactions. Curr. Opin struct. Biol. 1, 732–740.
Kaartinen, V. et al. Glycoasparaginase from human leukocytes. J. biol. Chem. 266, 5860–5869 (1991).
Tikkanen, R., Enomaa, N., Riikonen, A., Ikonen, E. & Peltonen, L. Intracellular sorting of aspartylglucosaminidase: the role of N-linked oligosaccharides and evidence of Man-6-P-independent lysosomal targeting. DNA Cell Biol. 14, 305–312 (1995).
Pfeffer, S.R. Targeting of proteins to the lysosomes. Curr. Top. Microbiol. Immun. 170, 43–65 (1991).
Musil, D. et al. The refined 2.15 Å X-ray crystal structure of human liver cathepsin B: the structural basis for its specificity. EMBO J. 10, 2321–2330 (1991).
Jia, Z. et al. Crystal structures of recombinant rat cathepsin B and a cathepsin B-inhibitor complex. J. biol. Chem. 270, 5527–5533 (1995).
Metcalf, P. and Fusek, M. Two crystal structures for cathepsin D: the lysosomal targeting signal and active site. EMBO J. 12, 1293–1302 (1993).
Baldwin, E.T. et al. Crystal structures of native and inhibited forms of human cathepsin D: implications for lysosomal targeting and drug design. Proc. natn. Acad. Sci. U.S.A. 90, 6796–6800 (1993).
Pollitt, R.J., Jenner, F.A. & Merskey, H. Aspartylglycosaminuria: an inborn error of metabolism associated with mental defect. Lancet ii, 253–255 (1968).
Ikonen, E., Enomaa, N., Ulmanen, I. & Peltonen, L. In vitro mutagenesis helps to unravel the biological consequences of aspartylglucosaminuria mutation. Genomics 11, 206–211 (1991).
McRee, D.E. A visual protein crystallographic software system for X11/XView. J. molec. Graph. 10, 44–46 (1992).
Steigemann, W. Protein, User's Guide (München 1992).
Zhang, K.Y.J. SQUASH - combining constraints for macromolecular phase refinement and extension. Acta Crystallogr. D49, 213–222 (1993).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density and the location of errors in these models. Ada Crystallogr. A47, 110–119 (1991).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallographic R factor refinement by molecular dynamics. Science 235, 458–460 (1987).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. appl. Crystallogr. 24, 946–950 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Oinonen, C., Tikkanen, R., Rouvinen, J. et al. Three-dimensional structure of human lysosomal aspartylglucosaminidase. Nat Struct Mol Biol 2, 1102–1108 (1995). https://doi.org/10.1038/nsb1295-1102
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1295-1102
This article is cited by
-
A plant-type L-asparaginase from Pyrobaculum calidifontis undergoes temperature dependent autocleavage
Biologia (2022)
-
Temperature dependent autocleavage and applications of recombinant L-asparaginase from Thermococcus kodakarensis for acrylamide mitigation
3 Biotech (2022)
-
Pcal_0970: an extremely thermostable l-asparaginase from Pyrobaculum calidifontis with no detectable glutaminase activity
Folia Microbiologica (2019)
-
Identification of Small Molecule Compounds for Pharmacological Chaperone Therapy of Aspartylglucosaminuria
Scientific Reports (2016)
-
Role of asparaginase variable loop at the carboxyl terminal of the alpha subunit in the determination of substrate preference in plants
Planta (2012)