Abstract
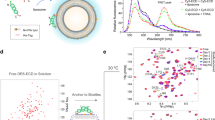
TRAIL, an apoptosis inducing ligand, has at least four cell surface receptors including the death receptor DR5. Here we report the crystal structure at 2.2 Å resolution of a complex between TRAIL and the extracellular region of DR5. TRAIL forms a central homotrimer around which three DR5 molecules bind. Radical differences in the surface charge of the ligand, together with variation in the alignment of the two receptor domains confer specificity between members of these ligand and receptor families. The existence of a switch mechanism allowing variation in receptor domain alignment may mean that it is possible to engineer receptors with multiple specificities by exploiting contact positions unique to individual receptor–ligand pairs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Nagata, S. Cell 88, 355–365 (1997).
Golstein, P. Curr-Biol 7, R750–R753 (1997).
Wiley, S. R., et al. Immunity 3, 673–682 (1995).
Mongkolsapaya, J. et al. J-Immunol 160, 3–6 (1998).
Walczak, H. et al. 5, 157–163 (1999).
Jones, E. Y., Stuart, D. I. & Walker, N. P. Nature 338, 225–228 (1989).
Banner, D. W. et al. Cell 73, 431–445 (1993).
Karpusas, M. et al. Structure 3, 1031–1039 (1995).
Cha, S. S. et al. 11, 253–261 (1999).
Naismith, J. H., Devine, T. Q., Kohno, T. & Sprang, S. R. Structure 4, 1251–1262. (1996).
Yamagishi, J. et al. Protein Eng. 3, 713–719 (1990).
Goh, C. R., Loh, C. S. & Porter, A. G. Protein Eng. 4, 785–791 (1991).
Schneider, P. et al. J. Biol. Chem. 272, 18827–12833 (1997).
Brojatsch, J., Naughton, J., Rolls, M. M., Zingler, K. & Young, J. A. Cell 87, 845–855 (1996).
Screaton, G. R., Mongkolsapaya, J., Xu, X. N., Cowper, A. E., McMichael, A. J. & Bell, J. I. Curr. Biol. 7, 693–696 (1997).
Park, Y. C., Burkitt, V., Villa, A. R., Tong, L. & Wu, H Nature 398, 533–538 (1999).
Gao, G. F. et al. Protein Sci. 7, 1245–1249 (1998).
Cordingley, M. G., Callahan, P. L., Sardana, V. V., Garsky, V. M. & Colonno, R. J. J. Biol. Chem. 265, 9062–9065 (1990).
Otwinowski, Z. O. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Navaza, J. Acta. Crystallogr. A 50, 157–163 (1994).
Brünger, A. T. XPLOR Version 3.1: A system for X-ray Crystallography and NMR. (Yale University Press New Haven, Connecticut; 1992).
Brunger, A. T. et al. Acta. Crystallogr. D 54, 905–921 (1998).
Collaborative Computational Project, No. 4 Acta Crystallogr D 50: 760–763 (1994).
De La Fortelle, E. & Bricogne, G. Meth. Enzymol. 276, 442–494 (1997).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Acta. Crystallogr. A 47, 110–119 (1991).
Stuart, D. I., Levine, M., Muirhead, H. & Stammers, D.K. J. Mol. Biol 134, 109–142 (1979).
Hubbard, S. J. & Thornton, J. M. Naccess, computer program. (University College, London; 1993).
Esnouf, R. M. Acta. Crystallogr. D 55, 938–940 (1999).
Merritt, E. A. & Bacon, D. J. Methods Enzymol. 277, 505–524 (1997).
Van Ostade, X., Tavernier, J. & Fiers, W. Protein Eng. 7, 5–22 (1994).
Nicholls, A., Sharp, K. A. & Honig, B. Proteins 11, 281–296 (1991).
Acknowledgements
We thank R. Esnouf, P. Gouet and R. Bryan for computing facilities and programs, K. Harlos for help with heavy atom soaks, G. Gao for advice on refolding protocols and crystallisation trials, K. Hudson and J. Heath for the rhinovirus 3c protease methodology, M. Gross for circular dichroism analysis, A. van der Merwe and A. McMichael for discussion. We also thank the staff at the European Molecular Biology Laboratory outstation, ID2 (ESRF, Grenoble) and at 9.6 (SRS, Daresbury). JM is funded by Siriraj Hospital Mahidol University Thailand, EYJ is funded by the Royal Society and the Cancer Research Campaign, GRS and DIS are funded by the MRC.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mongkolsapaya, J., Grimes, J., Chen, N. et al. Structure of the TRAIL–DR5 complex reveals mechanisms conferring specificity in apoptotic initiation. Nat Struct Mol Biol 6, 1048–1053 (1999). https://doi.org/10.1038/14935
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/14935
This article is cited by
-
Characterizing the regulatory Fas (CD95) epitope critical for agonist antibody targeting and CAR-T bystander function in ovarian cancer
Cell Death & Differentiation (2023)
-
Autoinhibitory structure of preligand association state implicates a new strategy to attain effective DR5 receptor activation
Cell Research (2023)
-
Programming cell-surface signaling by phase-separation-controlled compartmentalization
Nature Chemical Biology (2022)
-
Disulfide bond-disrupting agents activate the tumor necrosis family-related apoptosis-inducing ligand/death receptor 5 pathway
Cell Death Discovery (2019)
-
Inflammatory bowel disease-associated ubiquitin ligase RNF183 promotes lysosomal degradation of DR5 and TRAIL-induced caspase activation
Scientific Reports (2019)