Abstract
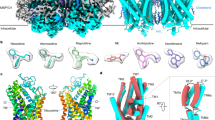
The gene coding for urate oxidase, an enzyme that catalyzes the oxidation of uric acid to allantoin, is inactivated in humans. Consequently, urate oxidase is used as a protein drug to overcome severe disorders induced by uric acid accumulation. The structure of the active homotetrameric enzyme reveals the existence of a small architectural domain that we call T-fold (for tunnelling-fold) domain. It assembles to form a perfect unusual dimeric α8β16 barrel. Urate oxidase may be the archetype of an expanding new family of tunnel-shaped proteins that now has three members; tetrahydropterin synthase, GTP cyclohydrolase I and urate oxidase. The structure of the active site of urate oxidase around the 8-azaxanthine inhibitor reveals an original mechanism of oxidation that does not require any ions or prosthetic groups.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Ames, B.N., Cathcart, R., Schwiers, E. & Hochstein, P. Uric acid provides an antioxidant defense in humans against oxidant- and radical-caused aging and cancer: a hypothesis. Proc Natl. Acad. Sci. USA 78, 6858–6862 (1981).
Varela-Echavarria, A. Montes de Oca-Luna, R. & Barrera-Saldana, H.A. Uricase protein sequences: conserved during vertebrate evolution but absent in humans. FASEB J. 2, 3092–3096 (1988).
Wu, X., Lee, C.C., Muzny, D.M. & Caskey, C.T. Urate oxidase: primary structure and evolutionary implications. Proc. Natl. Acad. Sci. USA 86, 9412–9416 (1989).
Wu, X., Muzny, D.M., Lee, C.C. & Caskey, C.T. Two independent mutational events in the loss of urate oxidase during hominoid evolution. J. Mol. Evol. 34, 78–84 (1992).
Legoux, R. et al. Cloning and expression in Escherichia coli of the gene encoding Aspergillus flavus urate oxidase. J. Biol. Chem. 267, 8565–8570 (1992).
Leplatois, P., Le Douarin, B. & Loison, G. High-level production of a peroxisomal enzyme: Aspergillus flavus uricase accumulates intracellularly and is active in Saccharomyces cerevisiae. Gene 122, 139–145 (1992).
Alvares, K. et al. Rat urate oxidase produced by recombinant baculovirus expression: formation of peroxisome crystalloid core-like structures. Proc. Natl. Acad. Sci. USA 89, 4908–4912 (1992).
Gould, S.J., Keller, G.A. & Subramani, S. Identification of peroxisomal targeting signals located at the carboxy terminus of four peroxisomal proteins. J. Cell Biol. 107, 897–905 (1988).
Orengo, C.A., Jones, D.T. & Thornton, J.M. Protein super-families and domain superfolds. Nature 372, 631–634 (1994).
Nar, H. et al. Atomic structure of GTP cyclohydrolase I. Structure 3, 459–466 (1995).
Nar, H., Huber, R., Heizmann, C.W., Thöny, B. & Bürgisser, D. Three-dimensional structure of 6-pyruvol tetrahydropterin synthase, an enzyme involved in tetrahydrobiopterin biosynthesis. EMBO J. 13, 1255–1262 (1994).
Murzin, A.G., Brenner, S.E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Bürgisser, D.M. et al. 6-pyruvoyl tetrahydropterin synthase, an enzyme with a novel type of active site involving both zinc binding and an intersubunit catalytic triad motif; site-directed mutagenesis of the proposed active center, characterization of the metal binding site and modelling of substrate binding. J. Mol. Biol. 253, 358–369 (1995).
Nar, H. et al. Active site topology and reaction mechanism of GTP cyclohydrolase I. Proc. Natl. Acad. Sci. USA 92, 12120–12125 (1995).
Bourne, H.R., Sanders, D.A. & McCormick, F. The GTPase superfamily: conserved structure and molecular mechanism. Nature 349, 117–127 (1991).
Bongaerts, G.P.A. & Vogels, G.D. Mechanism of uricase action. Biochim. Biophys. Acta 567, 295–308 (1979).
Bentley, R. & Neuberger, A. The mechanism of action of uricase. Biochem. J. 52, 694–699 (1952).
Pitts, O.M. & Priest, D.G. Uricase reaction intermediate. Mechanism of borate and hydroxide ion catalyst. Biochemistry 12, 1358–1363 (1973).
Sokolic, L., Modric, N. & Poje, M. Regiochemical course of chemical and enzymic uricolysis to allantoin. A non-degradative 13C-NMR evidence. Tetrahedron Lett. 32, 7477–7480 (1991).
Modric, N., Derome, A.E., Ashcroft, S.J.H. & Poje, M. Tracing and identification of uricase reaction intermediates. A direct 13C-NMR/isotope-labelling evidence. Tetrahedron Lett. 33, 6691–6694 (1992).
Kahn, K. & Tipton, P.A. Kinetic mechanism and cofactor content of soybean root nodule urate oxidase. Biochemistry 36, 4731–4738 (1997).
Kahn, K., Serfozo, P. & Tipton, P.A. Identification of the true product of the urate oxidase reaction. J. Am. Chem. Soc. 119, 5435–5442 (1997).
Messerschmidt, A. & Pflugrath, J.W. Crystal orientation and X-ray pattern prediction routines for area-detector diffractometer systems in macromolecular crystallography. J. Appl. Crystallogr. 20, 306–315 (1987).
Leslie, A.G.W. Joint CCP4 and ESF-EACMB Newsletter on Protein Crystallography (SERC Daresbury Laboratory, Warrington; 1992).
Schiltz, M., Prangé, T. & Fourme, R. On the preparation and X-ray data collection of isomorphous xenon derivatives. J. Appl. Crystallogr. 27, 950–960 (1994).
Prangé, T. et al. Exploring hydrophobic sites in proteins with xenon or krypton. Proteins, Struct, Funct, and Genetics, in the press.
CCP4. The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D50, 760–763 (1994).
Cowtan, K.D. & Main, P. Improvement of macromolecular electron-density maps by the simultaneous application of real and reciprocal space constraints. Acta Crystallogr. D49, 148–157 (1993).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Read, R.J. Structure-factor probabilities for related structures. Acta Crystallogr. A46, 900–912 (1990).
Brünger, A.T., Krukowski, A. & Erickson, J.W. Slow-cooling protocols for crystallographic refinement by simulated annealing. Acta Crystallogr. A46, 585–593 (1990).
Labesse, G., Colloc'h, N., Pothier, J. & Mornon, J.P. P-SEA: a new efficient assignment of secondary structure from Cα trace of proteins.CABIOS 13, 291–295 (1997).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures.J. Appl. Crystallogr. 24, 946–950 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Colloc'h, N., Hajji, M., Bachet, B. et al. Crystal Structure of the protein drug urate oxidase-inhibitor complex at 2.05 Å resolution. Nat Struct Mol Biol 4, 947–952 (1997). https://doi.org/10.1038/nsb1197-947
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1197-947
This article is cited by
-
Exploring the impact of taurine on the biochemical properties of urate oxidase: response surface methodology and molecular dynamics simulation
Journal of Biological Engineering (2024)
-
Mathematical modeling for bioprocess optimization of a protein drug, uricase, production by Aspergillus welwitschiae strain 1–4
Scientific Reports (2019)
-
Crystal structure of Bacillus fastidious uricase reveals an unexpected folding of the C-terminus residues crucial for thermostability under physiological conditions
Applied Microbiology and Biotechnology (2015)
-
Geometric consequences of electron delocalization for adenine tautomers in aqueous solution
Journal of Molecular Modeling (2014)
-
Spectroscopic investigations and hydrogen bond interactions of 8-aza analogues of xanthine, theophylline and caffeine: a theoretical study
Journal of Molecular Modeling (2013)