Abstract
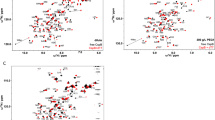
The solution structure of the DNA-binding domain of the Drosophila heat shock transcription factor, as determined by multidimensional multinuclear NMR, resembles that of the helix-turn-helix class of DNA-binding proteins. The domain comprises a four-stranded antiparallel β-sheet, packed against a three-helix bundle. The second helix is significantly distorted and is separated from the third helix by an extended turn which is subject to conformational averaging on an intermediate time scale. Helix 3 forms a classical amphipathic helix with polar and charged residues exposed to the solvent. Upon titration with DNA, resonance shifts in the backbone and Asn and Gln side-chain amides indicate that helix 3 acts as the recognition helix of the heat shock transcription factor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Lindquist, S. & Craig, E.A. The heat-shock proteins. A. Rev. Genet. 22, 631–677 (1988).
Morimoto, R.I., Tissieres, A. &. Georgopoulos, C. The stress response, function of the proteins, and perspectives. In Stress proteins in Biology and Medicine (Eds, Morimoto, R.I., Tissieres, A. & Georgopoulos, C.,). 1–36 (Cold Spring Harbor Laboratory Press, New York, 1990).
Hendrick, J.P. & Hartl, F.U. Molecular chaperone functions of heat-shock proteins. A. Rev. Biochem. 62, 349–384 (1993).
Sorger, P.K. Heat shock factor and the heat shock response. Cell 65, 363–366 (1991).
Lis, J.T. & Wu, C. Heat Shock Factor. In Transcriptional Regulation, (Eds, McKnight, S.L. & Yamamoto, K.R.,) 907–930 (Cold Spring Harbor Laboratory Press, New York, 1992).
Lis, J.T. & Wu, C. Protein traffic on the heat shock promoter: parking, stalling and trucking along. Cell, 74, 1–20 (1993).
Kim, S.-J., Tsukiyama, T., Lewis, M.S. & Wu, C. The interaction of the DNA binding domain of Drosophila heat shock factor with its cognate DNA site: a thermodynamic analysis using analytical ultracentrifugation. Prot. Sci. 3, 1040–1051 (1994).
Vuister, G.W., Kim, S.-J., Wu, C. & Bax, A. NMR evidence for similarities between the DNA-binding regions of Drosophila melanogaster heat shock factor and the helix-turn-helix and HNF3/fork head families of transcription factors. Biochemistry 33, 10–16 (1994).
Vuister, G.W., Kim, S.-J., Wu, C. & Bax, A. 2D and 3D NMR study of phenylalanine residues in proteins by reverse isotopic labeling. J. Am. chem. Soc. (in the press).
Nilges, M., Clore, G.M. & Gronenborn, A.M. Determination of three-dimensional structures of proteins from interproton distance data by hybrid distance geometry-dynamical simulated annealing calculations. FEBS Lett. 229, 317–324 (1988).
Brunger, A.T. X-PLOR Version 3.1: A System for X-ray Crystallography and NMR, Yale University, New Haven, CT, USA (1992).
Wüthrich, K. NMR of Proteins and Nucleic Acids, (John Wiley, New York, 1986).
Harrison, C.J., Bohm, A.A. & Nelson, H.C.M. Crystral structure of the DNA binding domain of the heat shock transcription factor. Science 263, 224–227 (1994).
Weber, I.T. & Steitz, T.A. Structure of a complex of catabolite gene activator protein and cyclic AMP refined at 2.5 Å resolution. J. molec. Biol. 198, 311–326 (1987).
Clark, K.L., Halay, E.D., Lai, E. & Burley, S.K. Co-crystal structure of the HNF-3/fork head DNA-recognition motif resembles histone H5. Nature 364, 412–420 (1993).
Perisic, O., Xiao, H. & Lis, J.T. Stable binding of Drosophila heat shock factor to head-to-head and tail-to-tail repeats of a conserved 5 bp recognition unit. Cell 59, 797–806 (1989).
Fernandes, M., Xiao, H. & Lis, J.T. Fine structure analyses of the Drosophila and Saccharomyces heat shock factor – heat shock element interactions. Nucleic Acids Res. 22, 167–173 (1994).
Harrison, S.C. A structural taxonomy of DNA-binding domains. Nature 353, 715–719 (1991).
Dekker, N., Cox, M., Boelens, R., Verrijzer, C.P. var der Vliet, P.C. & Kaptein, R. Solution structure of the POU-specific DNA-binding domain of Oct-1. Nature 362, 852–855 (1993).
Pabo, C.O., Aggarwal, A.K., Jordan, S.R., Beamer, L.J., Obeysekare, U.R. & Harrison, S.C. Conserved residues make similar contacts in two represser-operator complexes. Science 247, 1210–1213 (1990).
Assa-Munt, N., Mortishire-Smith, R.J., Aurora, R., Herr, W. & Wright, P.E. The solution structure of the Oct-1 POU-specific domain reveals a striking similarity to the bacteriophage I repressor DNA-binding domain. Cell 73, 193–205 (1993).
Clos, J. et al. Molecular cloning and expression of a hexameric Drosophila heat shock factor subject to negative regulation. Cell 63, 1085–1097 (1990).
Clore, G.M. & Gronenborn, A.M. Applications of three- and four-dimensional heteronuclear NMR spectroscopy to protein structure determination. Progr. NMR Spectr. 23, 43–92 (1991).
Bax, A., et al. Measurement of homo- and heteronuclear J couplings from quantitative J correlation. Meth. Enzymol., (Eds, James, T. L. & Oppenheimer, N.) 239 79–125 (Academic Press, San Diego, 1994).
Grzesiek, S. & Bax, A. The importance of not saturating H2O in protein NMR. Application to sensitivity enhancement and NOE measurements. J. Am. chem. Soc. 115, 12593–12594 (1993).
Muhandiram, D.R., Xu, G.Y. & Kay, L.E. An enhanced-sensitivity pure absorption gradient 4D 13C-edited NOESY experiment. J. biomol. NMR 3, 463–470 (1993).
Vuister, G.W. et al. Increased resolution and improved spectral quality in four-dimensional 13C/13C separated HMQC-NOESY-HMQC spectra using pulsed field gradients. J. magn. Reson. B 101, 210–213 (1993).
Clore, G.M. et al. The three-dimensional structure of α1-purothionin in solution: combined use of nuclear magnetic resonance, distance geometry and restrained molecular dynamics. EMBO J. 5, 2729–2735 (1986).
Wüthrich, K., Billeter, M. & Braun, W. Pseudo-structures for the 20 common amino acids for use in studies of protein conformations by measurements of intramolecular proton-proton distance constraints with nuclear magnetic resonance. J. molec. Biol. 169, 949–961 (1983).
Clore, G.M., Gronenborn, A.M., Nilges, M. & Ryan, C.A. Three-dimensional structure of potato carboxypeptidase inhibitor in solution. A study using nuclear magnetic resonance, distance geometry, and restrained molecular dynamics. Biochemistry 26, 8012–8023 (1987).
Huebel, A. & Schoeffl, F. Arabidopsis thaliana HSF gene sequence (submitted to the EMBL Data Library, November 1993).
Scharf, K.D., Rose, S., Zott, W., Schoef, F. & Nover, L. Three tomato genes code for heat stress transcription factors with a region of remarkable homology to the DNA-binding domain of the yeast HSF. EMBO J. 9, 4495–4501 (1990).
Jakobsen, B.K. & Pelham, H.R.B. A conserved heptapeptide restrains the activity of the yeast heat shock transcription factor. EMBO J. 10, 369–375 (1991).
Sorger, P.K. & Pelham, H.R. Yeast heat shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell 54, 855–864 (1988).
Gallo, G.J., Prentice, H. & Kingston, R.E. Heat shock factor is required for growth at normal temperatures in the fission yeast Schizosaccharomyces pombe. Molec. cell. Biol. 13, 749–761 (1993).
Nakai, A. & Morimoto, R.I. Characterization of a novel chicken heat shock transcription factor 3, suggests a new regulatory pathway. Molec. cell. Biol. 13, 1983–1997 (1993).
Sarge, K.D., Zimarino, V., Holm, K., Wu, C. & Morimoto, R.I. Cloning and characterization of two mouse heat shock factors with distinct inducible and constitutive DNA binding ability. Genes Dev. 5, 1902–1911 (1991).
Rabindran, S.K., Giorgi, G., Clos, J. & Wu, C. Molecular cloning and expression of a human heat shock factor, HSF1. Proc. natn. Acad. Sci. U.S.A. 88, 6906–6910 (1991).
Schuetz, T.J., Sheldon, L., Gallo, G.J., Tempst, P. & Kingston, R.E. Isolation of a cDNA for HSF2: evidence for two heat shock factor genes in humans. Proc. natn. Acad. Sci. U.S.A. 88, 6911–6915 (1991).
Higgins, D.G. & Sharp, P.M. Fast and sensitive multiple sequence alignments on a microcomputer. Cabios Comm. 5, 151–153 (1989).
Carson, M. Ribbon models for macromolecules. J. molec. Graphics 5, 103–106 (1987).
Brooks, B.R. et al. CHARMM: a program for macromolecular energy, minimization and dynamics calculations. J. comput. Chem. 74, 187–217 (1983).
Schultz, S.C., Shields, G.C. & Steitz, T.A. Crystal structure of CAP-DNA is bent by 90°. Science 253, 1001–1007 (1991).
Klemm, J.D., Rould, M.A., Aurora, R., Herr, W. & Pabo, C.O. Crystal structure of the Oct-1 POU domain bound to an octamer site: DNA recognition with tethered DNA-binding modules. Cell 77, 21–32 (1994).
Beamer, L.J. & Pabo, C.O. Refined 1.8 Å crystal structure of the λ repressor-operator complex. J. molec. Biol. 227, 177–196 (1992).
Aggarwal, A.K., Rodgers, D.W., Drottar, M., Ptashne, M. & Harrison, S.C. Recognition of a DNA operator by the repressor of phage 434: a view at high resolution. Science 242, 899–907 (1988).
Mondragon, A. & Harrison, S.C. The phage 434 Cro/Or1 complex at 2.5 Å resolution. J. molec. Biol. 219, 321–334 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vuister, G., Kim, SJ., Orosz, A. et al. Solution structure of the DNA-binding domain of Drosophila heat shock transcription factor. Nat Struct Mol Biol 1, 605–614 (1994). https://doi.org/10.1038/nsb0994-605
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0994-605
This article is cited by
-
Molecular characterization of Hsf1 as a master regulator of heat shock response in the thermotolerant methylotrophic yeast Ogataea parapolymorpha
Journal of Microbiology (2021)
-
Over-expression of LlHsfA2b, a lily heat shock transcription factor lacking trans-activation activity in yeast, can enhance tolerance to heat and oxidative stress in transgenic Arabidopsis seedlings
Plant Cell, Tissue and Organ Culture (PCTOC) (2017)
-
Structure of human heat-shock transcription factor 1 in complex with DNA
Nature Structural & Molecular Biology (2016)
-
Metabolic control of the proteotoxic stress response: implications in diabetes mellitus and neurodegenerative disorders
Cellular and Molecular Life Sciences (2016)
-
Genome-wide analysis of HSF family transcription factors and their responses to abiotic stresses in two Chinese cabbage varieties
Acta Physiologiae Plantarum (2014)