Abstract
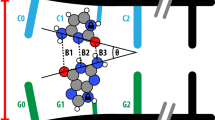
Cytosine methylation or bromination of the DNA sequence d(GGCGCC)2 is shown here to induce a novel extended and eccentric double helix, which we call E-DNA. Like B-DNA, E-DNA has a long helical rise and bases perpendicular to the helix axis. However, the 3′-endo sugar conformation gives the characteristic deep major groove and shallow minor groove of A-DNA. Also, if allowed to crystallize for a period of time longer than that yielding E-DNA, the methylated sequence forms standard A-DNA, suggesting that E-DNA is a kinetically trapped intermediate in the transition to A-DNA. Thus, the structures presented here chart a crystallographic pathway from B-DNA to A-DNA through the E-DNA intermediate in a single sequence. The E-DNA surface is highly accessible to solvent, with waters in the major groove sitting on exposed faces of the stacked nucleotides. We suggest that the geometry of the waters and the stacked base pairs would promote the spontaneous deamination of 5-methylcytosine in the transition mutation of dm5C-dG to dT-dA base pairs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Noyer-Weidner, M. & Trautner, T.A. In DNA methylation: molecular biology and biological significance. (eds, Jost, J.P. & Saluz, H.P.), 39–108 (Birkhäuser Verlag, Boston; 1993).
Singer-Sam, J. & Riggs, A.D. In DNA methylation: molecular biology and biological significance. (eds, Jost, J.P. & Saluz, H.P.) 358–384 (Birkhäuser Verlag, Boston; 1993).
Sasaki, H., Allen, N.D. & Surani, M.A. In DNA Methylation: Molecular Biology and Biological Significance. (eds Jost, J.P. & Saluz, H.P.) 469–486 (Birkhäuser Verlag, Boston, 1993).
Antequera, F. & Bird, A. In DNA methylation: molecular biology and biological significance. (eds, Jost, J.P. & Saluz, H.P.), 169–185 (Birkhäuser Verlag, Boston; 1993).
Mooers, B.H.M., Schroth, G.P., Baxter, W.W. & Ho, P.S. J. Mol. Biol. 249, 772–784 (1995).
Frederick, C.A. et al. Biopolymers 26, S145–S160 (1987).
Behe, M. & Felsenfeld, G. Proc. Natl Acad. Sci. USA 78, 1619–1623 (1981).
Fujii, S., Wang, A.H.-J., van der Marel, G., van Boom, J.H. & Rich, A. Nucleic Acids Res. 10, 7879–7892 (1982).
Timsit, Y. & Moras, D. Methods Enzymol. 211, 409–429 (1992).
Xu, Q., Shoemaker, R.K. & Braunlin, W.H. Biophys. J. 65, 1039–1049 (1993).
Takahara, P.M., Rosenzweig, A.C., Frederick, C.A. & Lippard, S.J. Nature 377, 649–652 (1995).
Arnott, S., Chandrasekaran, R., Millane, R.P. & Park, H.-S. J. Mol. Biol. 188, 631–640 (1986).
Doucet, J., Benoit, J.-P., Cruse, W.B.T., Prange, T. & Kennard, O. Nature 337, 190–192 (1989).
Mayer-Jung, C., Moras, D. & Timsit, Y. EMBO J. 17, 2709–2718 (1998).
Timsit, Y. & Moras, D. EMBO J. 13, 2737–2746 (1994).
Wang, A.H.-J., Fujii, S., van Boom, J.H. and Rich, A. Proc. Natl. Acad. Sci. USA 79, 3968–3972 (1982).
Ng, H.-L., Kopka, M.L., & Dickerson, R.E. Proc. Natl. Acad. Sci. USA 97, 2035–2039 (2000).
Zhang, X. & Mathews, C.K. J. Biol. Chem. 269, 7066–7069 (1994).
Brünger, A.T. X-PLOR version 3.1: a system for X-ray crystallography and NMR. (Yale University Press, New Haven, Connecticut; 1992).
Dickerson, R.E. Methods Enzymol. 211, 67–111 (1992).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Lavery, R. & Sklenar, H. J. Biomol. Struct. Dyn. 6, 655–667 (1989).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Carter, C.W. Jr., Biochimie 77, 92–98 (1995).
Weiner, S.J. et al. J. Am. Chem. Soc. 106, 765–784 (1984).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Pflugrath, J.W. Acta Crystallogr. D 55, 1718–1725 (1999).
Parkinson, G., Vojtechovsky, J., Clowney, L., Brünger, A.T. & Berman, H.M. Acta Crystallogr. D 52, 57–64 (1996).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Brünger, A.T. Nature 355, 472–475 (1992).
Acknowledgements
We thank B.H.M. Mooers and the P.A. Karplus laboratory for helpful discussion, and K.E. van Holde, C.K. Mathews, and W.C. Johnson, Jr. for reading this manuscript. This work was supported by the National Science Foundation, the Oregon American Cancer Society, and the Environmental Heath Science Center at OSU. X-ray facilities were funded in part by the M.J. Murdock Charitable Trust.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vargason, J., Eichman, B. & Ho, P. The extended and eccentric E-DNA structure induced by cytosine methylation or bromination. Nat Struct Mol Biol 7, 758–761 (2000). https://doi.org/10.1038/78985
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/78985
This article is cited by
-
Demethylation of (Cytosine-5-C-methyl) DNA and regulation of transcription in the epigenetic pathways of cancer development
Cancer and Metastasis Reviews (2008)
-
Rational Vector Design for Efficient Non-viral Gene Delivery: Challenges Facing the Use of Plasmid DNA
Molecular Biotechnology (2008)
-
Cobalt(II), nickel(II) and zinc(II) do not bind to intra-helical N(7) guanine positions in the B-form crystal structure of d(GGCGCC)
JBIC Journal of Biological Inorganic Chemistry (2003)