Abstract
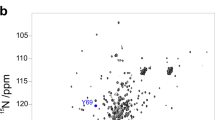
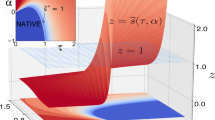
Molten globules are partially folded forms of proteins that are thought to be general intermediates in protein folding. Nonetheless, there is limited structural information about such species because they possess conformational heterogeneity and complex dynamical properties that lead to extreme line broadening in NMR spectra. Here we use a 2-D NMR approach that overcomes this difficulty by detecting the unfolding of individual residues in a molten globule in increasing concentrations of denaturant. The results show that the structure in the low pH form of α-lactalbumin (α-LA) is not formed cooperatively. Moreover, a core region remains collapsed under extremely denaturing conditions, even when the majority of the polypeptide chain is completely unfolded. Our results support a model for protein folding in which the core provides a template for correct assembly of the remainder of the structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kuwajima, K. The molten globule state as a clue for understanding the folding and cooperativity of globular-protein structure. Proteins 6, 87–103 (1989).
Ptitsyn, O.B. The molten globule state. in Protein Folding (ed. Creighton, T.E.) 243–300 (W.H. Freeman and Co., New York; 1992).
Dobson, C.M. Solid evidence for molten globules. Curr. Biol. 4, 636–640 (1994).
Peng, Z-y. & Kim, P.S. A protein dissection study of a molten globule. Biochemistry 33, 2136–2141 (1994).
Wu, L.C., Peng, Z-y. & Kim, P.S. Bipartite structure of the α-lactalbumin molten globule. Nature Struct. Biol. 2, 281–286 (1995).
Hughson, F.M., Barrick, D. & Baldwin, R.L. Probing the stability of a partly folded apomyoglobin intermediate by site-directed mutagenesis. Biochemistry 30, 4113–4118 (1991).
Lin, L., Pinker, R.J., Forde, K., Rose, G.D. & Kallenbach, N.R. Molten globular characteristics of the native state of apomyoglobin. Nature Struct. Biol. 1, 447–452 (1994).
Kay, M.S. & Baldwin, R.L. Packing interactions in the apomyoglobin folding intermediate. Nature Struct. Biol. 3, 439–445 (1996).
Kuwajima, K. The molten globule state of α-lactalbumin. The FASEB Journal 10, 102–109 (1996).
Kay, L.E., Keifer, P. & Saarinen, T. Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J. Am. Chem. Soc. 114, 1063–1065 (1992).
Baum, J., Dobson, C.M., Evans, P.A. & Hanley, C. Characterization of a partly-folded protein by NMR methods: studies of the molten globule state of guinea pig α-lactalbumin. Biochemistry 28, 7–13 (1989).
Wang, Y. & Shortle, D. The equilibrium folding pathway of staphylococcal nuclease: identification of the most stable chain-chain interactions by NMR and CD spectroscopy. Biochemistry 34, 15895–15905 (1995).
Wang, Y., Alexandrescu, A.T. & Shortle, D. Initial studies of the equilibrium folding pathway of staphylococcal nuclease. Phil. Trans. R. Soc. tond. B 348, 27–34 (1995).
Eliezer, D. & Wright, P.E. Is apomyoglobin a molten globule? Structural characterization by NMR. J. Mol. Biol. 263, 531–538 (1996).
Wishart, D.S., Sykes, B.D. & Richards, F.M. Relationship between nuclear magnetic resonance chemical shift and protein secondary structure. J. Mol. Biol. 222, 311–333 (1991).
Fiebig, K.M., Schwalbe, H., Buck, M., Smith, L.J. & Dobson, C.M. Toward a description of the conformation of denatured states of proteins. Comparison of a random coil model with NMR measurements J. Phys. Chem. 100, 2661–2666 (1996).
Wüthrich, K. NMR of Proteins and Nucleic Acids, John Wiley & Sons, New York (1986).
Peng, Z-y, Wu, L.C. & Kim, P.S. Local structural preferences in the α-lactalbumin molten globule. Biochemistry 34, 3248–3252 (1995).
Alexandrescu, A.T., Evans, P.A., Pitkeathly, M., Baum, J. & Dobson, C.M. Structure and dynamics of the acid-denatured molten globule state of α-lactalbumin: a two-dimensional NMR study. Biochemistry 32, 1707–1718 (1993).
Chyan, C.L., Wormald, C., Dobson, C.M., Evans, P.A. & Baum, J. Structure and stability of the molten globule state of guinea-pig α-lactalbumin: a hydrogen exchange study. Biochemistry 32, 5681–5691 (1993).
Schulman, B.A., Redfield, C., Peng, Z-y., Dobson, C.M. & Kim, P.S. Different subdomains are most protected from hydrogen exchange in the molten globule and native states of human α-lactalbumin. J. Mol. Biol. 253, 651–657 (1995).
Schulman, B.A. & Kim, P.S. Proline scanning mutagenesis of a molten globule reveals non-cooperative formation of protein's overall topology. Nature Struct. Biol. 3, 682–687 (1996).
Morozova, L.A., Haynie, D.T., Arico-Muendel, C., Van Dael, H. & Dobson, C.M. Structural basis of the stability of a lysozyme molten globule. Nature Struct. Biol. 2, 871–875 (1995).
Fontana, A., de Laureto, P. P., de Filippis, V., Scaramella, E. & Zambonin, M. Probing the partly folded states of proteins by limited proteolysis. Folding & Design 2, R17–R26 (1997).
Uversy, V.N. & Ptitsyn, O.B. All-or-none solvent induced transitions between native, molten globule and unfolded states in globular proteins. Folding & Design 1, 117–122 (1996).
Jennings, P.A. & Wright, P.E. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science 262, 892–896 (1993).
Loh, S.N., Kay, M.S. & Baldwin, R.L. Structure and stability of a second molten globule intermediate in the apomyoglobin folding pathway. Proc. Natal. Acad. Sci. USA 92, 5446–5450 (1995)
Kay, M.S. & Baldwin, R.L. Packing interactions in the apomyoglobin folding intermediate. Nature Struct. Biol. 3, 439–445 (1996).
Jamin, M. & Baldwin, R.L. Refolding and unfolding kinetics of the equilibrium folding intermediate of apomyoglobin. Nature Struct. Biol. 3, 613–618 (1996).
Shimizu, A., Ikeguci, M. & Sugai, S. Unfolding of the molten globule state of α-lactalbumin studied by 1H NMR. Biochemistry 32, 13198–13203 (1993).
Xie, D., Bhakuni, V. & Freire, E. Calorimetric determination of the energetics of the molten globule intermediate in protein folding: apo-α-lactalbumin. Biochemistry 30, 10673–10678 (1991).
Yutani, K., Ogasahar, K. & Kuwajima, K. Absence of the thermal transition in apo-α-lactalbumin in the molten globule state. A study by differential scanning microcalorimetry. J. Mol. Biol. 228, 347–350 (1992).
Jeng, M.-F. & Englander, S.W. Stable submolecular folding units in a non-compact formofcytochromec. J.Mol.Biol. 221, 1045–1061 (1991).
Balbach, J., Forge, V., van Nuland, N.A.J., Winder, S., Hore, P.J. & Dobson, C.M. Following protein folding in real time using NMR spectroscopy. Nature Struct. Biol. 2, 865–870 (1995).
Arai, M. & Kuwajima, K. Rapid formation of a molten globule intermediate in refolding of α-lactalbumin. Folding and Design 1, 275–287 (1996).
Marion, D., Driscoll, P.C., Kay, L.E., Wingfield, P.T., Bax, A., Gronenborn, A.M. & Clore, G.M. Overcoming the overlap problem in the assignment of 1H NMR spectra of larger proteins by use of three-dimensional heteronuclear 1H-15N Hartmann-Hahn multiple quantum coherence and nuclear Overhauser multiple quantum coherence spectroscopy: application to interleukin-1β. Biochemistry 28, 6150–6156 (1989).
Frenkiel, T., Bauer, C., Carr, M.D., Birdsall, B. & Feeney, J. HMQC-NOESY-HMQC, a three-dimensional NMR experiment which allows detection of nuclear Overhauser effects between protons with overlapping signals. J. Magn. Reson. 90, 420–425 (1990).
Driscoll, P.C., Clore, G.M., Marion, D., Wingfield, P.T. & Gronenborn, A.M. Complete resonance assignment for the polypeptide backbone of interleukin 1β using three-dimensional heteronuclear NMR spectroscopy. Biochemistry 29, 3542–3556 (1990).
Pace, C.N. Determination and analysis of urea and guanidine hydrochloride denaturation curves. Meth. Enz. 131, 266–280 (1986).
Acharya, K.R., Ren, J.S., Stuart, D.I., Phillips, D.C. & Fenna, R.E. Crystal structure of human α-lactalbumin at 1. 7 Å resolution. J. Mol. Biol. 221, 571–581 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schulman, B., Kim, P., Dobson, C. et al. A residue-specific NMR view of the non-cooperative unfolding of a molten globule. Nat Struct Mol Biol 4, 630–634 (1997). https://doi.org/10.1038/nsb0897-630
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0897-630
This article is cited by
-
New structural insights into Golgi Reassembly and Stacking Protein (GRASP) in solution
Scientific Reports (2016)
-
Cold denaturation of a protein dimer monitored at atomic resolution
Nature Chemical Biology (2013)
-
The client protein p53 adopts a molten globule–like state in the presence of Hsp90
Nature Structural & Molecular Biology (2011)
-
N-terminal myristoylation alters the calcium binding pathways in neuronal calcium sensor-1
JBIC Journal of Biological Inorganic Chemistry (2011)
-
Probing the urea dependence of residual structure in denatured human α-lactalbumin
Journal of Biomolecular NMR (2009)