Abstract
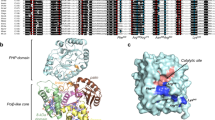
The crystal structure of Serratia endonuclease has been solved to 2.1 Å by multiple isomorphous replacement. This magnesium-dependent enzyme is equally active against single- and double-stranded DNA, as well as RNA, without any apparent base preference. The Serratia endonuclease fold is distinct from that of other nucleases that have been solved by X-ray diffraction. The refined structure consists of a central layer containing six antiparallel β-strands which is flanked on one side by a helical domain and on the opposite side by one dominant helix and a very long coiled loop. Electrostatic calculations reveal a strongly polarized molecular surface and suggest that a cleft between this long helix and loop, near His 89, may contain the active site of the enzyme.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Saito, H., Elting, L., Bodey, G.P. & Berkey, P. Serratia bacteremia: review of 118 cases. Rev. infect. Dis. 11, 912–920 (1989).
Acar, J.F. Serratia marcescens infections. Infect. Control. 7, 273–278 (1986).
Ball, T.K., Saurugger, P.N. & Benedik, M.J. The extracellular nuclease gene of Serratia marcescens and its secretion from Escherichia coli. Gene 57, 183–192 (1987).
Molla, A., Matsumura, Y., Yamamoto, T., Okamura, R. & Maeda, H. Pathogenic capacity of proteases from Serratia marcescens and Pseudomonas aeruginosa and their suppression by chicken egg white ovomacroglobulin. Infect. Immun. 55, 2509–2517 (1987).
Heller, K. Lipolytic activity copurified with the outer membrane of Serratia marcescens. J. Bact. 140, 1120–1122 (1979).
Givskov, M., Olsen, L. & Molin, S. Cloning and expression in Escherichia coil of the gene for extracellular phospholipase A1 from Serratia liquefaciens. J. Bact. 170, 5855–5862 (1988).
Jones, J.D.G., Grady, K.L., Suslow, T.V. & Bedbrook, J.R. Isolation and characterization of genes encoding two chitinase enzymes from Serratia marcescens. EMBO J. 5, 496–473 (1986).
Muro-Pastor, A.M., Flores, E., Herrero, A. & Wolk, C.P. Identification, genetic analysis and characterization of a sugar-nonspecific nuclease from the cyanobacterium Anabaena sp. PCC 7120. Molec. Microbiol. 6, 3021–3030 (1992).
Eaves, G.N. & Jeffries, C.D. Isolation and properties of an exocellular nuclease of Serratia marcescens. J. Bact. 85, 273–278 (1963).
Nestle, M. & Roberts, W.K. An extracellular nuclease from Serratia marcescens: II. specificity of the enzyme. J. biol. Chem. 244, 5219–5225 (1969).
Weston, S.A., Lahm, A. & Suck, D. X-ray structure of the DNase I-d(GGTATACC)2 complex at 2.3 Å resolution. J. molec. Biol. 226, 1237–1256 (1992).
Miller, M.D., Benedik, M.J., Sullivan, M.C., Shipley, N.S. & Krause, K.L. Crystallization and preliminary crystallographic analysis of a novel nuclease from Serratia Marcesens. J. molec. Biol. 222, 27–30 (1991).
Filimonova, M.N., Balaban, N.P., Sharipova, F.R. & Leshchinskaya, I.B. Production of Serratia marcescens nuclease in a homogeneous state and study of the physicochemical properties of the enzyme. Biokhimiya 45, 2096–2103 (1980).
Product Specifications for Benzonase, The first industrial endonuclease. (Benzon Pharma A/S, Helseholmen 1, P.O. Box 1185, DK-2650 Hvidovre Denmark, 1993).
Kurinenko, B.M., Belyaeva, M.I., Cherepneva, I.E. & Viesture, Z.A. The antitumor effect of Serratia marcescens nuclease bound covalently to soluble dextran. Vopr. Onkol. 23, 94–98 (1977).
Panfilova, Z.I., Batalina, T.A. & Salganik, R.I. Summaries of Reports at the Second All-Union Conference on the Enzymes of Microorganisms 1–67 (Moscow, 1978).
Suck, D. Nuclease structure and catalytic function. Curr. Opin. struct. Biol. 2, 84–92 (1992).
Tan, R.C., Truong, T.N., McCammon, J.A. & Sussman, J.L. Acetylcholinesterase: electrostatic steering increases the rate of ligand binding. Biochemistry 32, 401–403 (1993).
Sines, J.J., Allison, S.A. & McCammon, J.A. Point charge distributions and electrostatic steering in enzyme/substrate encounter: Brownian dynamics of modified copper/zinc superoxide dismutases. Biochemistry 29, 9403–9412 (1990).
Knowles, J.R. Enzyme catalysis: not different, just better. Nature. 350, 121–124 (1991).
Brändén, C. Relation between structure and function of α/β proteins. Q. Rev. Biophys. 13, 317–338. (1980).
Suck, D. & Oefner, C. Structure of DNase I at 2.0 Å resolution suggests a mechanism for binding to and cutting DNA. Nature 321, 620–625 (1986).
Messerschmidt, A. & Pflugrath, J.W. Crystal orientation and X-ray pattern prediction routines for area-detector diffractometer systems in macromolecular crystallography. J. appl. Cryst. 20, 306–315 (1987).
Kabsch, W. Evaluation of single-crystal X-ray diffraction data from a position-sensitive detector. J. appl. Cryst. 21, 916–924 (1988).
Steigemann, W. PROTEIN: A program system for the crystal structure analysis of proteins. (Max-Plank Institute for Biochemistry, Munich, Germany, 1992).
Jones, T.A. A graphics model building and refinement system for macromolecules. J. appl. Cryst. 11, 268–272 (1978).
Brünger, A.T. X-PLOR version 3.1 A System for X-ray Crystallography and NMR 1–382 (Yale University Press, New Haven, 1992).
Hendrickson, W.A. Stereochemically restrained refinement of macromolecular structures. Meth. Enzym. 115, 252–270, 1985.
Nicholls, A. & Honig, B. GRASP: Graphical Representation and Analysis of Surface Properties. (Columbia University, New York, 1992).
Davis, M.E., Madura, J.D., Luty, B.A. & McCammon, J.A. Electrostatics and diffusion of molecules in solution: simulations with the University of Houston brownian dynamics program. Comput. Phys. Commun. 62, 187–197 (1991).
QUANTA, version 3.3 (Molecular Simulations Inc., Waltham, MA, 1992)
Carson, M. RIBBONS 2.0 J. appl. Crystallogr. 24, 958–961 (1991).
Jones, T.A., Zou, J.-Y. & Cowan, S.W. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta crystallogr. A47, 110–119 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Miller, M., Tanner, J., Alpaugh, M. et al. 2.1 Å structure of Serratia endonuclease suggests a mechanism for binding to double-stranded DNA. Nat Struct Mol Biol 1, 461–468 (1994). https://doi.org/10.1038/nsb0794-461
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0794-461
This article is cited by
-
Endonuclease increases efficiency of osteoblast isolation from murine calvariae
Scientific Reports (2021)
-
An integrated transcriptomic and proteomic approach to identify the main Torymus sinensis venom components
Scientific Reports (2021)
-
A novel cell permeability assay for macromolecules
BMC Molecular and Cell Biology (2020)
-
Study of ChiR function in Serratia marcescens and its application for improving 2,3-butanediol from crystal chitin
Applied Microbiology and Biotechnology (2017)
-
Analysis of the Mechanism of Mg2+ Action on the RNase Activity of Serratia marcescens Endonuclease
BioNanoScience (2017)