Abstract
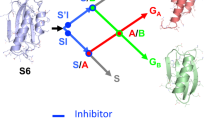
A lattice model with side chains was used to investigate protein folding with computer simulations. In this model, we rigorously demonstrate the existence of a specific folding nucleus. This nucleus contains specific interactions not present in the native state that, when weakened, slow folding but do not change protein stability. Such a decoupling of folding kinetics from thermodynamics has been observed experimentally for real proteins. From our results, we conclude that specific non-native interactions in the transition state would give rise to φ-values that are negative or larger than unity. Furthermore, we demonstrate that residue Ile 34 in src SH3, which has been shown to be kinetically, but not thermodynamically, important, is universally conserved in proteins with the SH3 fold. This is a clear example of evolution optimizing the folding rate of a protein independent of its stability and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Creighton, T.E. Proteins: structures and molecular properties, 2nd edition (W.H. Freeman and Company, New York; 1993).
Li, A. & Daggett, V. Molecular dynamics simulation of the unfolding of barnase: characterization of the major intermediate. J. Mol. Biol. 275, 677–694 (1998).
Duan, Y. & Kollman, P.A. Pathways to a protein folding intermediate observed in a 1-microsecond simulation in aqueous solution. Science 282, 740–744 ( 1998).
Lazaridis, T. & Karplus, M. ‘New view’ of protein folding reconciled with the old through multiple unfolding simulations. Science 278, 1928–1931 ( 1997).
Abkevich, V.I., Gutin, A.M. & Shakhnovich, E.I. Specific nucleus as the transition state for protein folding: evidence from lattice model. Biochemistry 33, 10026–10036 (1994).
Klimov, D.K. & Thirumalai, D. Cooperativity in protein folding, from lattice models with sidechains to real proteins. Folding Design 3, 127–139 ( 1998).
Socci, N.D., Onuchic, J.N. & Wolynes, P.G. Protein folding mechanisms and the multidimensional folding funnel. Proteins 32, 136– 158 (1998).
Hao, M-H & Scheraga, H.A. Molecular mechanisms for cooperative folding of proteins. J. Mol. Biol. 277, 973–983 (1998).
Kolinski, A. & Skolnick, J. Monte Carlo simulations of protein folding. I. Lattice model and interaction scheme. Proteins 18, 338–352 (1994).
Berriz, G.F., Gutin, A.M. & Shakhnovich, E.I. Cooperativity and stability in a Langevin model of proteinlike folding. J. Chem. Phys. 106, 9276–9285 (1997).
Guo, Z. & Brooks III, C.L. Thermodynamics of protein folding: a statistical mechanical study of a small all-β protein . Biopolymers 42, 745–757 (1997).
Dokholyan, N.V., Buldyrev, S.V., Stanley, H.E. & Shakhnovich, E.I. Identifying the protein folding nucleus using molecular dynamics. J. Mol. Biol. 296, 1183–1188 (2000).
Itzhaki, L.S., Otzen, D.E. & Fersht, A.R. The structure of the transition state for folding of chymotrypsin inhibitor 2 analysed by protein engineering methods: evidence for a nucleation-condensation mechanism for protein folding. J. Mol. Biol. 254, 260–288 (1995).
López-Hernández, E. & Serrano, L. Structure of the transition state for folding of the 129 aa protein CheY resembles that of a smaller protein, CI-2. Folding Design 1, 43–55 (1996).
Martinez, J.C., Pisabarro, M.T. & Serrano, L. Obligatory steps in protein folding and the conformational diversity of the transition state. Nature Struct. Biol. 5, 721–729 (1998).
Martinez, J.C. & Serrano, L. The folding transition state between SH3 domains is conformationally restricted and evolutionarily conserved. Nature Struct.Biol. 6, 1010– 1015 (1999).
Grantcharova, V.P., Riddle, D.S., Santiago, J.V. & Baker, D. Important role of hydrogen bonds in the structurally polarized transition state for folding of the src SH3 domain. Nature Struct. Biol. 5, 714–720 (1998).
Riddle, D.S. et al. Experiment and theory highlight role of native state topology in SH3 folding. Nature Struct. Biol. 6, 1016–1024 (1999).
Fulton, K.F., Main, E.R.G., Daggett, V. & Jackson, S.E. Mapping the interactions present in the transition state for unfolding/folding of FKBP12. J. Mol. Biol. 291, 445– 461 (1999).
Bromberg, S. & Dill, K.A. Side-chain entropy and packing in proteins. Protein Sci. 3, 997– 1009 (1994).
Hart, W.E. & Istrail, S. Lattice and off-lattice side chain models of protein folding: linear time structure prediction better the 86% of optimal. J. Comp. Biol. 4, 241– 259 (1997).
Villegas, V., Martínez, J.C., Avilés, F.X. & Serrano, L. Structure of the transition state in the folding process of human procarboxypeptidase A2 activation domain. J. Mol. Biol. 283, 1027–1036 (1998).
Dobson, C.M. et al. Mutational analysis of acylphosphatase suggests the importance of topology and contact order in protein folding. Nature Struct. Biol. 6, 1005–1009 ( 1999).
Daggett, V., Li, A. & Fersht, A.R. Combined molecular dynamics and φ-value analysis of structure-reactivity relationships in the transition state and unfolding pathway of barnase: structural basis of Hammond and anti-Hammond effects. J. Am. Chem. Soc. 120, 12740–12754 (1998).
Klimov, D.K. & Thirumalai, D. Lattice models for proteins reveal multiple folding nuclei for nucleation-collapse mechanism. J. Mol. Biol. 282, 471–492 ( 1998).
Gutin, A.M., Abkevich, V.I. & Shakhnovich, E.I. A protein engineering analysis of the transition state for protein folding: simulation in the lattice model. Folding Design 3, 183–194 ( 1998).
Matouschek, A., Kellis, J.T., Serrano L. & Fersht, A.R. Mapping the transition state and pathway of protein folding by protein engineering. Nature 340, 122–126 ( 1989).
Fersht, A.R., Itzhaki, L.S., ElMasry, N.F., Matthews, J.M. & Otzen, D.E. Single versus parallel pathways of protein folding and fractional formation of structure in the transition state. Proc. Natl Acad. Sci. USA 91, 10426 –10429 (1994).
_ali, A., Shakhnovich, E. & Karplus, M. How does a protein fold? Nature 369, 248–251 (1994).
Mirny, L.A., Abkevich, V.I. & Shakhnovich, E.I. How evolution makes proteins fold quickly. Proc. Natl Acad. Sci. USA 95, 4976– 4981 (1998).
Mirny, L.A. & Shakhnovich, E.I. Universally conserved positions in protein folds: reading evolutionary signals about stability, folding kinetics and function. J. Mol. Biol. 291, 177– 196 (1999).
Miyazawa, S. & Jernigan, R. Estimation of effective interresidue contact energies from protein crystal structures: quasi-chemical approximation . Macromolecules 18, 534– 552 (1985).
Metropolis, N., Rosenbluth, A.W., Rosenbluth, M.N., Teller, A.H. & Teller, E. Equation of state calculations by fast computing machines. J. Chem. Phys. 21, 1087–1092 (1953).
Abkevich, V.I., Gutin, A.M. & Shakhnovich, E.I. Improved design of stable and fast-folding model proteins . Folding Design 1, 221– 230 (1996).
Dodge, C., Schneider, R. & Sander, C. The HSSP database of protein structure-sequence alignments and family profiles. Nucleic Acids Res. 26, 313–315 (1998).
Holm, L. & Sander, C. The FSSP database: fold classification based on structure-structure alignment of proteins. Nucleic Acids Res. 24, 206–209 ( 1996).
Shakhnovich, E.I. Folding nucleus: specific or multiple? Insights from lattice models and experiments . Folding Design 3, R108– R111 (1998).
Per, J.K. MOLSCRIPT: A program to produce both detailed and schematic plots of protein structures. J. App. Crystallogr. 24, 946 –950 (1991).
Merrit, E.A. & Bacon, D.J. Raster3D: Photorealistic molecular graphics. Methods Enzymol. 277, 505– 524 (1997).
Acknowledgements
We thank the NSERC of Canada and NIH for financial support. L.L. would like to thank H. Angerman, G. Berriz, S. Choe, N.V. Dokholyan, L. Gutman, A. Ishchenko, E. Kussell, M. Morrissey and J. Shimada for computer assistance and stimulating discussions. He is particularly grateful to O. Clement for encouragement at a crucial point in this research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, L., Mirny, L. & Shakhnovich, E. Kinetics, thermodynamics and evolution of non-native interactions in a protein folding nucleus. Nat Struct Mol Biol 7, 336–342 (2000). https://doi.org/10.1038/74111
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/74111
This article is cited by
-
Native structure-based modeling and simulation of biomolecular systems per mouse click
BMC Bioinformatics (2014)
-
Structural characterization of a misfolded intermediate populated during the folding process of a PDZ domain
Nature Structural & Molecular Biology (2010)
-
Three key residues form a critical contact network in a protein folding transition state
Nature (2001)