Abstract
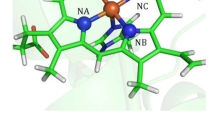
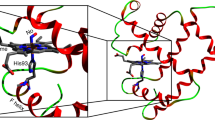
The distribution of carbon monoxide after photodissociation in the myoglobin haem pocket has been investigated using molecular dynamics simulations at 300 K. The results show that both intermediates (one close to the haem iron and one further away) observed in recent low temperature X-ray studies of photodissociated CO have a high probability of occurrence, even at ambient temperatures. The fact that the O of CO is oriented toward the haem iron in the closer intermediate provides an explanation for the slow rate of CO geminate rebinding. A refinement against X-ray data generated from the molecular dynamics simulations indicates that the CO has a broader distribution in the haem pocket than is apparent from the experimental electron density. This effect is likely to be general for systems containing highly mobile groups.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Antonini, E. & Brunori, M. Haemoglobin and myoglobin in their reactions with ligands. (North-Holland, London, 1971).
Springer, B.A., Sligar, S.G., Olson, J.S. & Phillips, Jr. G.N. Mechanisms of ligand recognition in myoglobin. J. Chem. Rev. 94, 699–714 (1994).
Collman, J.P., Brauman, J.I., Halbert, T.R. & Suslick, K.S. Nature of O2 and CO binding to metalloporphyrins and haem proteins. Proc. Natl. Acad. Sci. USA 73, 3333–3337 (1976).
Ansari, A. et al. Rebinding and relaxation in the myoglobin pocket. Biophys. Chem. 26, 337–355 (1987).
Elber, R. & Karplus, M. Enhanced sampling in molecular dynamics: Use of the time-dependent Hartree approximation for a simulation of carbon monoxide diffusion through myoglobin J. Am. Chem. Soc. 112, 9161–9175 (1990).
Straub, J. & Karplus, M. Molecular dynamics study of the photodissociation of carbon monoxide from myoglobin: ligand dynamics in the first 10 ps. Chem. Phys. 158, 221–248 (1991).
Gibson, Q.E., Regan, R., Elber, R., Olson, J.S. & Carver, T.E. Distal pocket residues affect picosecond ligand recombination in myoglobin. J. Biol. Chem. 267, 22022–22034 (1992).
Carlson, M.L. et al. Nitric oxide recombination to double mutants of myoglobin:role of ligand diffusion in a fluctuating haem pocket. Biochemistry 33, 10597–10606 (1994).
Li, H., Elber, R. & Straub, J.E. Molecular dynamics simulation of NO recombination to myoglobin mutants. J. Biol. Chem. 268, 17908–17916 (1993).
Schlichting, I., Berendzen, J., Phillips, Jr. G.N. & Sweet, R.M. Crystal structure of photolyzed carbonmonoxymyoglobin. Nature 371, 808–812 (1994).
Teng, T.-Y., _rajer, V. & Moffat, K. Photolysis-induced structural changes in single crystals of carbonmonoxymyoglobin at 40K. Nature Struct. Biol. 1, 701–705 (1994).
Hartmann, H. et al. X-ray structure determination of a metastable state of carbonmonoxy myoglobin after photodissociation. Proc. Natl. Acad. Sci. USA 93, 7013–7016 (1996).
Lim, M., Jackson, T.A. & Anfinrud, P.A. Binding of CO to myoglobin from a haem pocket docking site to form nearly linear Fe-C-O. Science 269, 962–966 (1995).
Lim, M., Jackson, T.A. & Anfinrud, P.A. Mid-infrared vibrational spectrum of CO after photodissociation from haem: Evidence for a ligand docking site in the haem pocket of haemoglobin and myoglobin. J. Chem. Phys. 102, 4355–4366 (1995).
Petrich, J.W., Poyart, C. & Martin, J.L. Photophysics and reactivity of haem proteins: A femtosecond absorption study of haemoglobin, myoglobin, and protohaem. Biochem. 27, 4049–4060 (1988).
Anfinrud, P.A., Han, C. & Hochstrasser, R.M. Direct observation of ligand dynamics in haemoglobin by subpicosecond infrared spectroscopy. Proc. Natl. Acad. Sci.USA 86, 8387–8391 (1989).
Alben, J.O. et al. Infrared spectroscopy of photodissociated carboxymyoglobm at low temperatures. Proc. Natl. Acad. Sci. USA 79, 3744–3748 (1982).
Kuriyan, J., Wilz, S., Karplus, M. & Petsko, G.A. X-ray structure and refinement of carbon-monoxy myoglobin at 1.5 Å resolution. J. Mol. Biol. 192, 133–154 (1986).
Teng, T.-Y., Huang, H.W. & Olah, G.A. 5K extended X-ray absorption fine structure and 40 K 10-s resolved extended X-ray absorption fine structurestudies of photolyzed carboxymyoglobin. Biochemistry 26, 8066–8072 (1987).
Powers, L. et al. Kinetic, structural, and spectroscopic identification of geminatestates of myoglobin: A ligand binding site on the reaction pathway. Biochemistry 26, 4785–4796 (1987).
Alben, J.O. et al. Isotope effect in molecular tunneling. Phys. Rev. Letts. 44, 1157–1160 (1980).
Jongeward, K.A. et al. Picosecond and nanosecond geminate recombination of myoglobin with CO, O2, NO, and Isocyamides. J. Am. Chem. Soc. 110, 380–387 (1988).
Kuriyan, J., Petsko, G.A., Levy, R.M. & Karplus, M. Effect of anisotropy and anharmonicity on protein crystallographic refinement. J. Mol. Biol. 190, 227–254 (1986).
Ichiye, T. & Karplus, M. Anisotropy and anharmonicity of atomic fluctuations inproteins: implications for X-ray analysis. Biochemistry 27, 3487–3497 (1988).
Sassaroli, M. & Rousseau, D.L. Simulation of carboxymyoglobin photodissociation. J. biol. Chem. 261, 16292–16294 (1986).
Hajdu, J. & Andersson, I. Fast Crystallography and time-resolved structures. Annu. Rev. Biopys. Biomol. Struct. 22, 467–497 (1993).
Bourgeois, D. et al. Single pulse Laue images from macromolecular crystals recorded at ESRF. SPIE 2521, 178–181 (1996).
Levitt, M. & Park, B.H. Water: now you see it, now you do not. Structure 1, 223–226 (1993).
Brooks, B.R. et al. CHARMM: A program for macromolecular energy, minimization, and dynamic calculations. J. Comp. Chem. 4, 187–217 (1983).
Petrich, J.W. et al. Ligand binding and protein relaxation in haem proteins: A room temperature analysis of NO geminate recombination. Biochemistry 30, 3975–3987 (1991).
Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W. & Klein, M.L. Comparison of simple potential functions for simulating liquid water. J Chem. Phys. 79, 926–935 (1983).
Loncharich, R.J. & Brooks, B.R. The effects of truncating long-range forces on protein dynamics. Proteins 6, 32–45 (1989).
Van Gunsteren, W.F. & Berendsen, H.J.C. Algorithms for molecular dynamics and constraint dynamics. Mol. Phys. 34, 1311–1327 (1977).
Brünger, A.T. & Karplus, M. Polar hydrogen in proteins:empirical energyplacement and neutron diffraction comparison. Proteins 4, 148–156 (1988).
Steinbach, P.J. & Brooks, B.R. Protein hydration elucidated by molecular dynamics simulation. Proc. Natl. Acad. Sci. USA 90, 9135–9139 (1993).
Brünger, A.T., Kuriyan, J. & Karplus, M. Crystallograpic R factor refinement by molecular dynamics. Science 235, 458–460 (1987)
Srajer, V. et al. Monoxide of myoglobin: nanosecond time-resolved crystallography. Science 274, 1726–1729 (1996).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vitkup, D., Petsko, G. & Karplus, M. A comparison between molecular dynamics and X-ray results for dissociated CO in myoglobin. Nat Struct Mol Biol 4, 202–208 (1997). https://doi.org/10.1038/nsb0397-202
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0397-202
This article is cited by
-
Oxygen diffusion in minihemoglobin from Cerebratulus lacteus: a locally enhanced sampling study
Theoretical Chemistry Accounts (2007)
-
Ligand binding and conformational motions in myoglobin
Nature (2000)
-
Ultrafast rotation and trapping of carbon monoxide dissociated from myoglobin
Nature Structural Biology (1997)