Abstract
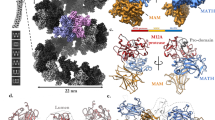
The three–dimensional structure of the catalytic domain of stromelysin–1 complexed with an N–carboxyl alkyl inhibitor has been determined by NMR methods. The global fold consists of three helices, a five stranded β–sheet and a methionine located in a turn near the catalytic histidines, classifying stromelysin–1 as a metzincin. Stromelysin–1 is unique in having two independent zinc binding sites: a catalytic site and a structural site. The inhibitor binds in an extended conformation. The S1′ subsite is a deep hydrophobic pocket, whereas S2′ appears shallow and S3′ open.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Woessner, Jr., J.F. Matrix metalloproteinases and their inhibitors in connective tissue remodeling. FASEB J. 5, 2145–2154 (1991).
Saus, J., Quinones, S., Otani, Y., Nagase, H., Harris Jr., E.D. & Kurkinen, M. The Complete Primary Structure of Human Matrix Metalloproteinase-3. J. biol. Chem. 263, 6742–6745 (1988).
MacNaul, K.L., Chartrain, N., Lark, M., Tocci, M.J. & Hutchinson, N.I., Expression of Stromelysin, Collagenase, and Tissue Inhibitor of Metalloproteinases-1 in Rheumatoid Human Synovial Fibroblasts. J. biol. Chem. 265, 17238–17245 (1990).
Marcy, A.I. et al. Human Fibroblast Stromelysin Catalytic Domain: Expression, Purification, and Characterization of a C-Terminally Truncated Form. Biochemistry 30, 6476–6483 (1991).
Salowe, S.P. et al. Characterization of Zinc-Binding Sites in Human Stromelysin-1: Stoichiometry of the Catalytic Domain and Identification of a Cysteine Ligand in the Proenzyme. Biochemistry 31, 4535–4540 (1992).
Holz, R.C., Salowe, S.P., Smith, C.K., Cuca, G.C. & Que, Jr., L. EXAFS Evidence for a “Cysteine Switch” in the Activation of Prostromelysin. J. Am. chem. Soc. 114, 9611–9614 (1992).
Gooley, P.R. et al. Secondary Structure and Zinc Ligation of Human Recombinant Short-Form Stromelysin by Multidimensional Heteronuclear NMR. Biochemistry 32, 13098–13108 (1993).
Bode, W., Gomis-Rüth, F.X., Huber, R., Zwilling, R. & Stöcker, W. Structure of astacin and implications for activation of astacin and zinc-ligation of collagenases. Nature 358, 164–167 (1992).
Bode, W., Gomis-Rüth, F.X. & Stöcker, W., Astacins, serralysins, snake venom and matrix metalloproteinases exhibit identical zinc-binding environments (HExxHxxGxxH and Met-turn) and topologies and should be grouped into a common family, the ‘metzincins’. FEBS Lett. 331, 134–140 (1993).
Chapman, K.T. et al. Inhibition of Matrix Metalloproteinases by N-Carboxyalkyl Peptides. J. med. Chem. 36, 4293–4301 (1993).
Nilges, M., Clore, G.M. & Gronenborn, A.M. Determination of three-dimensional structures of proteins from interproton distance data by hybrid distance geometry-dynamical simulated annealing calculations. FEBS Lett. 229, 317–324 (1988).
Brunger, A. X-PLOR Manual, Version 3.1 (Yale University Press, New Haven and London, 1992).
Van Doren, S.R. et al. Assignments for the Main-Chain Nuclear Magnetic Resonances and Delineation of the Secondary Structure of the Catalytic Domain of Human Stromelysin-1 As Obtained from Triple-Resonance 3D NMR Experiments. Biochemistry 32, 13109–13122 (1993).
Richardson, J.S. β-Sheet topology and the relatedness of proteins. Nature 268, 495–500 (1977).
Reynolds, W.F., Peat, I.R., Freedman, M.H. & Lyerla, Jr., J.H. Determination of theTautomeric Form of the Imidazole Ring of L-Histidine in Basic Solution by Carbon-13 Magnetic Resonance Spectroscopy. J. Am. chem. Soc. 95, 328–331 (1973).
Vallee, B.L. & Auld, D.S., and Structure of Zinc Enzymes and Other Proteins. Biochemistry 29, 5647–5659 (1990).
Schechter, I. & Berger, A. On the Size of the Active Site in Proteases. Biochem. Biophys. Res. Commun. 27, 157–162 (1967).
Mathews, B.W., Jansonius, J.N., Colman, P.M., Schoenborn, B.P. & Dupourque, D., Three-dimensional Structure of Thermolysin. Nature new Biol. 238, 37–41 (1972).
Pauptit, R.A. et al. Crystal Structure of Neutral Protease from Bacillus cereus Refined at 3.0 Å Resolution and Comparison with the Homologous But More Thermostable Enzyme Thermolysin. J. molec. Biol. 199, 525–537 (1988).
Thayer, M.M. Flaherty, K.M. & MacKay, D.B. Three-Dimensional Structure of the Elastase of Pseudomonas aeruginosa at 1.5-Å Resolution. J. biol. Chem. 266, 2864–2871 (1991).
Gomis-Rüth, F.X., Stöcker, W., Huber, R., Zwilling, R. & Bode, W. Refined 1.8 Å X-ray Crystal Structure of Astacin, a Zinc-endopeptidase from the Crayfish Asfacus astacus L. Structure Determination, Refinement, Molecular Structure and Comparison with Thermolysin. J. molec. Biol. 229, 945–968 (1993).
Gomis-Rüth, F.X., Kress, L.F. & Bode, W. First structure of a snake venom metalloproteinase: a prototype for matrix metalloproteinases/collagenases. EMBO J. 12, 4151–4157 (1993).
Murphy, G.J.P., Murphy, G. & Reynolds, J.J. The origin of matrix metalloproteinases and their familial relationships. FEBS Lett. 289, 4–7 (1991).
Dumermuth, E. et al. The Astacin Family of Metalloendopeptidases. J. biol. Chem. 266, 21381–21385 (1991).
Duong, F., Lazdunski, A., Cami, B. & Murgier, M. Sequence of a cluster of genes controlling synthesis and secretion of alkaline protease in Pseudomonas aeruginosa: relationships to other secretory pathways. Gene 121, 47–54 (1992).
Weaver, L.H., Kester, W.R. & Mathews, B.W. A Crystallographic Study of the Complex of Phosphoramidon with Thermolysin. A model for the Presumed Catalytic Transition State and for the Binding of Extended Substrates. J. molec. Biol. 114, 119–132 (1977).
Mathews, B.W. Structural Basis of the Action of Thermolysin and Related Zinc Peptidases. Acc. chem. Res. 21, 333–340 (1988).
Ikura, M., Kay, L.E., Tschudin, R. & Bax, A. Three-Dimensional NOESY-HMQC Spectroscopy of a 13C-labelled Protein. J. magn. Reson. 86, 204–209 (1990).
Zuiderweg, E.R.P. & Fesik, S . Heteronuclear Three-Dimensional NMR Spectroscopy of the Inflammatory Protein C5a. Biochemistry 28, 2387–2391 (1989).
Clore, G.M., Kay, L.E., Bax, A. & Gronenborn, A.M. Four-Dimensional 13C/13C-Edited Nuclear Overhauser Enhancement Spectroscopy of a Protein in Solution: Application to Interleukin 1β. Biochemistry 30, 12–18 (1991).
Suri, A.K. & Levy, R.M. Estimation of Interatomic Distances in Proteins from NOE Spectra at Longer Mixing Times Using an Empirical Two-Spin Equation. J. magn. Reson. 101, 320–324 (1993).
Kay, L.E. & Bax, A., Methods for the Measurement of NH-CαH Coupling Constants in 15N-Labeled Proteins. J. magn. Reson. 86, 110–126 (1990).
Powers, R., Gronenborn, A.M., Clore, G.M. & Bax, A., Triple-Resonance NMR of 13C/15N-Enriched Proteins Using Constant-Time Evolution. J. magn. Reson. 94, 209–213 (1991).
Vuister, G.W., Delaglio, F. & Bax, A., An Empirical Correlation between 1JCαHα and Protein Backbone Conformation. J. Am. chem. Soc. 114, 9674–9675 (1992).
Spera, S. & Bax, A., Correlation between Protein backbone Conformation and Cα and Cβ 13C Nuclear Magnetic resonance Chemical Shifts. J. Am. chem. Soc. 113, 5490–5492 (1991).
Neri, D., Szperski, T., Otting, G., Senn, H. & Wuthrich, K. Stereospecific Nuclear Magnetic Resonance Assignments of the Methyl Groups of Valine and Leucine in the DNA-Binding Domain of the 434 Represser by Biosynthetically Directed Fractional 13C Labeling. Biochemistry 28, 7510–7516 (1989).
Otting, G. & Wüthrich, K. Heteronuclear filters in two-dimensional [1H,1H]-NMR Spectroscopy: combined use with isotope labelling for studies of macromolecular conformation and intermolecular interactions. Q. Rev. Biophys. 23, 39–96 (1990).
Petros, A.M., Kawai, M., Luly, J.R. & Fesik, S.W. Conformation of two non-immunosuppressive FK506 analogs when bound to FKBP by isotope-filtered NMR. FEBS Lett. 308, 309–314 (1992).
Horrocks, Jr., W.D., Ishley, J.N. & Whittle, R.R. Models for Cobalt(II)-Substituted Zinc Metalloenzymes. 1. Comparison of the Crystal Structures of Complexes of the Type {M(RCOO)2(Im)2] (Im = Imidazole; M = Co, Zn; R = CH3, C2H5). Inorg. Chem. 21, 3265–3269 (1982).
Bear, C.A., Duggan, K.A. & Freeman, H.C., Tetraimidazolezinc(II) Pechlorate. Acta. Crystallog. 31, 2713–2715 (1975).
Garrett, T.P.J., Guss, J.M. & Freeman, H.C. Hexakis(imidazole)manganese(II) Dichloride Tetrahydrate, [Mn(C3H4N2)6]Cl2.4H2O, and Hexakis(imidazole)zinc(II) DichlorideTetrahydrate,[Zn(C3H4N2)6]Cl2.4H2O. Acta. Crystallog. 39, 1027–1031 (1983).
Ashby, C.I.H., Cheng, C.P., Duesler, E.N. & Brown, T.L., Structure and 14N Nuclear Quadrupole Resonance Spectrum of Catena-m-imidazolato-bis(imidazole)zinc Nitrate. Donor Characteristics of Coordinated Imidazolate. J. Am. chem. Soc. 100, 6063–6067 (1978).
Brooks, B.R., Bruccoleri, R.E., States, D.J., Swaminathan, S. & Karplus, M. CHARMM: a program for macromolecular energy,minimization, and dynamics calculations. J. comput. Chem. 4, 187–217 (1983).
Holmes, M.A. & Mathews, B.W. Structure of Thermolysin Refined at 1.6 Å Resolution. J. molec. Biol. 160, 623–639 (1982).
Berstein, F.C. et al. The Protein Data Bank: A Computer-based Archival File for Macromolecular Structures. J. molec. Biol. 112,, 535–542 (1977).
Hasty, K.A. et al. Human Neutrophil Collagenase. A Distinct gene Product with Homology To Other Metalloproteinases. J. biol. Chem. 265, 11421–11424 (1990).
Colman, P.M., Jansonius, J.N. & Mathews, B.W., Structure of Thermolysin: An Electron Density Map at 2.3 Å Resolution. J. molec. Biol. 70, 701–724 (1972).
Niedzwiecki, L., Teahan, J., Harrison, R.K. & Stein, R.L., Specificity of the Human Matrix Metalloproteinase Stromelysin and the Development of Continuous Flurometric Assays. Biochemistry 31, 12618–12623 (1992).
Netzel-Arnett, S., Fields, G., Birkedal-Hansen, H. & Van Wart, H.E., Specificities of Human Fibroblast and Neutrophil Collagenases. J. biol. Chem. 266, 6747–6755 (1991).
Netzel-Arnett, S., Sang, Q.-X., Moore, W.G.I., Navre, M., Birkedal-Hansen, H. & Van Wart, H.E. Specificities of Human 72- and 92-kDa Gelatinases (Type IV Collagenases) and PUMP (Matrilysin). Biochemistry 32, 6427–6432 (1993).
Krauhs, E., Dörsam, H., Little, M., Zwilling, R. & Ponstingl, H. A Protease from Astacus fluviatilis as an Aid in Protein Sequencing. Analyt. Biochem. 119, 153–157 (1982).
Fox, J.W., Campbell, R., Beggerly, L., Bjarnason, J.B. Substrate specificities and inhibition of two hemorrhagic zinc proteases Ht-c and Ht-d from Crotalus atrox venom. Eur. J. Biochem. 156, 65–72 (1986).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gooley, P., O'Connell, J., Marcy, A. et al. The NMR structure of the inhibited catalytic domain of human stromelysin–1. Nat Struct Mol Biol 1, 111–118 (1994). https://doi.org/10.1038/nsb0294-111
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0294-111
This article is cited by
-
A non-synonymous coding SNP Lys45Glu of mmp3 associated with ESCC genetic susceptibility in population of Henan, China
The Chinese-German Journal of Clinical Oncology (2009)
-
Matrix Metalloproteinase (MMP)-3 Polymorphism in Patients with HBV Related Chronic Liver Disease
Digestive Diseases and Sciences (2008)