Abstract
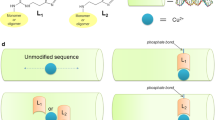
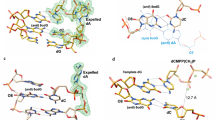
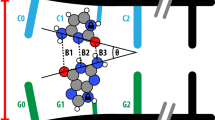
Intercalating complexes of rhodium(III) are strong photo-oxidants that promote DNA strand cleavage or electron transfer through the double helix. The 1.2 Å resolution crystal structure of a sequence-specific rhodium intercalator bound to a DNA helix provides a rationale for the sequence specificity of rhodium intercalators. It also explains how intercalation in the center of an oligonucleotide modifies DNA conformation. The rhodium complex intercalates via the major groove where specific contacts are formed with the edges of the bases at the target site. The phi ligand is deeply inserted into the DNA base pair stack. The primary conformational change of the DNA is a doubling of the rise per residue, with no change in sugar pucker from B-form DNA. Based upon the five crystallographically independent views of an intercalated DNA helix observed in this structure, the intercalator may be considered as an additional base pair with specific functional groups positioned in the major groove.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lerman, L.S. J. Mol. Biol. 3, 18–30 ( 1961).
Wilson, W.D. & Jones, R.L. Adv. Pharmacol. Chemother. 18, 177–222 (1981).
W. Saenger Principles of nucleic acid structure. (Springer-Verlag, New York, New York; 1984).
Johann, T.W. & Barton, J.K. Phil. Trans. R. Soc. Lond. A 354, 299–324 (1996).
Dandliker, P.J., Holmlin, R.E. & Barton, J.K. Science 275, 1465– 1468 (1997).
Sitlani, A., Long, E.C., Pyle, A.M. & Barton, J.K. J. Am. Chem. Soc. 114, 2303–2312 ( 1992).
Hall, D.B., Holmlin, R.E. & Barton, J.K. Nature 382, 731– 735 (1996).
Murphy, C.J. & Barton, J.K. Methods Enzymol. 226 , 576–594 (1993).
Jackson, B.A., Alekseyev, V.Y. & Barton, J.K. Biochemistry 38, 4655– 4662 (1999).
Chow, C.S. & Barton, J..K. Methods Enzymol. 212 , 219–242 (1992).
Holmlin, R.E., Dandliker, P.J. & Barton, J.K. Angew. Chemie Intl. Ed. 36, 2715–2730 (1997).
Odom, D.T., Parker, C.S. & Barton, J.K. Biochemistry 38, 5155– 5163 (1999).
Krotz, A.H., Hudson, B.P. & Barton, J.K. J. Am. Chem. Soc. 115, 12577 –12578 (1993).
Hudson, B.P. & Barton, J.K. J. Am. Chem. Soc. 120 , 6877–6888 (1998).
Sobell, H.M., Tsai, C.C., Jain, S.C. & Gilbert, S.G. J. Mol. Biol. 114, 333–365 ( 1977).
Wang, A.H.-J. et al. Nature 276, 471–474 (1978).
Wang, A.H.-J. et al. J. Biomolec. Struct. Dynamics 4, 319 –342 (1986).
Wang, A.H.-J., Ughetto, G., Quigley, G.J. & Rich, A. Biochemistry 26, 1152–1163 ( 1987).
Kamitori, S. & Takusagawa, F. J. Mol. Biol. 225 , 445–456 (1992).
Gasper, S. M. et al. J. Am. Chem. Soc. 120, 12402– 12409 (1998).
Bugg, C.E., Thomas, J.M. & Sundaralingam, M. & Rao, S.T. Biopolymers 10, 175–219 (1971).
Kelley, S.O. & Barton, J.K. Science 283, 375–381 (1999).
Egli, M. & Gessner, R.V. Proc. Natl. Acad. Sci. USA 92, 180–184 (1995).
Shields, T.P. & Barton, J.K. Biochemistry 34, 15037–15408 (1995).
Krotz, A.H., Kuo, L.Y., Shields, T.P. & Barton, J.K. J. Am. Chem. Soc. 115, 3877–3882 ( 1993).
Baker, E.N. & Hubbard, R.E. Prog. Biophys. Molec. Biol. 44, 97–179 (1984).
Schwabe, J.W.R. Curr. Opin. Struct. Biol. 7, 126–134 (1997).
Krotz, A.H. & Barton, J.K. Inorg. Chem. 33, 1940–1947 (1994).
Otwinowski, Z. In Data collection and processing (eds Sawyer, N.I.L. & Bailey, S.) 56–61(SERC Daresbury Laboratory, United Kingdom; 1993).
Terwilliger, T.C. SOLVE. http://www.solve.lanl.gov (1999).
Bailey, S. Acta Crystallogr. D 50, 760–763 (1994).
Sheldrick, G.M. & Schneider, T.R. Methods Enzymol. 277, 319–343 (1997).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277 , 505–524 (1997).
Esnouf, R.M. Acta Crystallogr. D 55, 938–940 (1999).
Lavery, R. & Sklenar, H. J. Biomolec. Struct. Dynamics 6, 63–91 (1988).
Acknowledgements
We are grateful to the NIH for research support and to the NSF and NIH for predoctoral fellowships to C.L.K and K.E.E. We thank L. Joshua-Tor, S.S. David, J.E. Wedekind, A.J. Chirino and R. Bau for assistance with the structure solution, and S. Horvath for oligonucleotide synthesis. The rotation camera facility at SSRL is supported by the U.S. Department of Energy and NIH.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Kielkopf, C., Erkkila, K., Hudson, B. et al. Structure of a photoactive rhodium complex intercalated into DNA. Nat Struct Mol Biol 7, 117–121 (2000). https://doi.org/10.1038/72385
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72385
This article is cited by
-
Two New Mononuclear Copper(II)-Dipeptide Complexes of 2-(2′-Pyridyl)Benzoxazole: DNA Interaction, Antioxidation and in Vitro Cytotoxicity Studies
Journal of Fluorescence (2017)
-
Crystal structure of Δ-[Ru(bpy)2dppz]2+ bound to mismatched DNA reveals side-by-side metalloinsertion and intercalation
Nature Chemistry (2012)
-
Double-helix disruption
Nature Chemistry (2012)
-
DNA base mismatch detection with bulky rhodium intercalators: synthesis and applications
Nature Protocols (2007)