Abstract
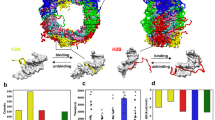
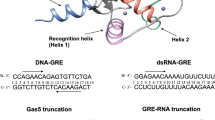
We have determined the solution structure of the complex between the 'winged-helix' enhancer binding domain of the Mu repressor protein and its cognate DNA site. The structure reveals an unusual use for the 'wing' which becomes immobilized upon DNA binding where it makes intermolecular hydrogen bond contacts deep within the minor groove. Although the wing is mobile in the absence of DNA, it partially negates the large entropic penalty associated with its burial by maintaining a small degree of structural order in the DNA-free state. Extensive contacts are also formed between the recognition helix and the DNA, which reads the major groove of a highly conserved region of the binding site through a single base-specific hydrogen bond and van der Waals contacts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Mizuuchi, K. Transpostional recombination: mechanistic insights from studies of Mu and other elements. Annu. Rev. Biochem. 61, 1011–1051 (1992).
Lavoie, B.D. & Chaconas, G. Transposition of phage Mu DNA. Curr. Topics Microbiol. Immunol. 204, 83–102 (1996).
Chaconas, G., Lavoie, B.D. & Watson, M.A. DNA transposition: jumping gene machine, some assembly required. Curr. Biol. 6, 817–820 (1996).
Mizuuchi, M. & Mizuuchi, K. Efficient Mu transposition requires interaction of transposase with a DNA sequence at the Mu operator: implications for regulation. Cell 58, 399–408 (1989).
Leung, P.C., Teplow, D.B. & Harshey, R.M. Interaction of distinct domains in Mu transposase with Mu DNA ends and an internal transpositional enhancer. Nature 338, 656–658 (1989).
Surette, M.G., Lavoie, B.D. & Chaconas, G. Action at a distance in Mu DNA transposition: an enhancer-like element is the site of action of supercoiling relief activity by integration host factor (IHF). EMBO J. 8, 3483–3489 (1989).
Surette, M.G. & Chaconas, G. The Mu transpositional enhancer can function in trans: requirement of the enhancer for synapsis but not strand cleavage. Cell 68, 1101–1108 (1992).
Harshey, R.M., Getzoff, E.D., Baldwin, D.L., Miller, J.L. & Chaconas, G. Primary structure of phage Mu transposase: homology to Mu repressor. Proc. Natl. Acad. Sci. USA 82, 7676–7680 (1985).
Clubb, R.T. et al. A novel class of winged helix-turn-helix protein: the DNA-binding domain of Mu transposase. Structure 2, 1041–1048 (1994).
Ilangovan, U., Wojciak, J.M., Connolly, K.M. & Clubb, R.T. NMR structure and functional studies of the Mu repressor DNA-binding domain. Biochemistry 38, 8367–8376 (1999).
Watson, M.A. & Chaconas, G. Three-site synapsis during Mu DNA transposition: a critical intermediate preceding engagement of the active site. Cell 85, 435–445 (1996).
Jiang, H., Yang, J.Y. & Harshey, R.M. Criss-crossed interactions between the enhancer and the att sites of phage Mu during DNA transposition. EMBO J. 18, 3845–3855 (1999).
Mizuuchi, M., Baker, T.A. & Mizuuchi, K. Assembly of the active form of the transposase-Mu DNA complex: a critical control point in Mu transposition. Cell 70, 303–311 (1992).
Baker, T.A. & Mizuuchi, K. DNA-promoted assembly of the active tetramer of the Mu transposase. Genes Dev. 6, 2221–2232 (1992).
Rousseau, P., Bétermier, M., Chandler, M. & Alazard, R. Interactions between the repressor and the early operator region of bacteriophage Mu. J. Biol. Chem. 271, 9739–9745 (1996).
Vogel, J.L., Li, Z.J., Howe, M.M., Toussaint, A. & Higgins, N.P. Temperature-sensitive mutations in the bacteriophage Mu c repressor locate a 63-amino-acid DNA-binding domain. J. Bacteriol. 173, 6568–6577 (1991).
Krause, H.M. & Higgins, N.P. Positive and negative regulation of the Mu operator by Mu repressor and Escherichia coli integration host factor. J. Biol. Chem. 261, 3744–3752 (1986).
Rice, P. & Mizuuchi, K. Structure of the bacteriophage Mu transposase core: a common structural motif for DNA transposition and retroviral integration. Cell 82, 209–220 (1995).
Clubb, R.T., Schumacher, S., Mizuuchi, K., Gronenborn, A.M. & Clore, G.M. Solution structure of the I gamma subdomain of the Mu end DNA-binding domain of phage Mu transposase. J. Mol. Biol. 273, 19–25 (1997).
Schumacher, S. et al. Solution structure of the Mu end DNA-binding ibeta subdomain of phage Mu transposase: modular DNA recognition by two tethered domains. EMBO J. 16, 7532–7541 (1997).
Tolman, J.R., Flanagan, J.M., Kennedy, M.A. & Prestegard, J.H. Nuclear magnetic dipole interactions in field-oriented proteins: information for structure determination in solution. Proc. Natl. Acad. Sci. USA 92, 9279–9283 (1995).
Tjandra, N., Omichinski, J.G., Gronenborn, A.M., Clore, G.M. & Bax, A. Use of dipolar 1H-15N and 1H-13C couplings in the structure determination of magnetically oriented macromolecules in solution. Nature Struct. Biol. 4, 732–738 (1997).
Pelton, J.G., Torchia, D.A., Meadow, N.D. & Roseman, S. Tautomeric states of the active-site histidines of phosphorylated and unphosphorylated IIIGlc, a signal-transducing protein from Escherichia coli, using two-dimensional heteronuclear NMR techniques. Protein Sci. 2, 543–558 (1993).
Clubb, R.T. et al. The wing of the enhancer-binding domain of Mu phage transposase is flexible and is essential for efficient transposition. Proc. Natl. Acad. Sci. USA 93, 1146–1150 (1996).
Billeter, M., Guntert, P., Luginbuhl, P. & Wuthrich, K. Hydration and DNA recognition by homeodomains. Cell 85, 1057–1065 (1996).
Gajiwala, K.S. & Burley, S.K. Winged helix proteins. Curr. Opin. Struct. Biol. 10, 110–116 (2000).
Clark, K.L., Hurley, E.D., Lai, E. & Burley, S.K. Co-crystal structure of the HNF-3/ fork head DNA-recognition motif resembles histone H5. Nature 364, 412–419 (1993).
Kodandapani, R. et al. A new pattern for helix-turn-helix recognition revealed by the PU.1 ETS-domain-DNA complex. Nature 380, 456–460 (1996).
Littlefield, O. & Nelson, H.C. A new use for the 'wing' of the 'winged' helix-turn-helix motif in the HSF-DNA cocrystal. Nature Struct. Biol. 6, 464–470 (1999).
Gajiwala, K.S. et al. Structure of the winged-helix protein hRFX1 reveals a new mode of DNA binding. Nature 403, 916–921 (2000).
Jin, C., Marsden, I., Chen, X. & Liao, X. Dynamic DNA contacts observed in the NMR structure of winged helix protein-DNA complex. J. Mol. Biol. 289, 683–690 (1999).
Louis, J.M., Martin, R.G., Clore, G.M. & Gronenborn, A.M. Preparation of uniformly isotope-labeled DNA oligonucleotides for NMR spectroscopy. J. Biol. Chem. 273, 2374–2378 (1998).
Grzesiek, S. & Bax, A. Improved 3D triple-resonance NMR techniques applied to a 31-kDa protein. J. Magn. Reson. 96, 432–440 (1992).
Wittekind, M. & Mueller, L. HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha-carbon and beta-carbon resonances in proteins. J. Magn. Reson. 101, 201–205 (1993).
Grzesiek, S. & Bax, A. Correlating backbone amide and side chain resonances in larger proteins by multiple relayed triple resonance NMR. J. Am. Chem. Soc. 114, 6291–6293 (1992).
Grzesiek, S., Anglister, J. & Bax, A. Correlation of backbone amide and aliphatic side-chain resonances in C-13/N-15-enriched proteins by isotropic mixing of C-13 magnetization. J. Magn. Reson. B 101, 114–119 (1993).
Bax, A., Clore, G.M. & Gronenborn, A.M. 1H-1H correlation via isotropic mixing of 13C magnetization, a new three-dimensional approach for assigning 1H and 13C spectra of 13C-enriched proteins. J. Magn. Reson. 88, 425–431 (1990).
Ikura, I., Kay, L.E. & Bax, A. Improved three-dimensional 1H-13C-1H correlation spectroscopy of a 13C-labeled protein using constant-time evolution. J. Biomol. NMR 1, 299–304 (1991).
Marion, D. et al. Overcoming the overlap problem in the assignment of 1H NMR spectra of larger proteins by use of three-dimensional heteronuclear 1H-15N Hartmann-Hahn-multiple quantum coherence and nuclear Overhauser-multiple quantum coherence spectroscopy: application to interleukin 1 beta. Biochemistry 28, 6150–6156 (1989).
Vuister, G.W. & Bax, A. Quantitative J correlation ‐ a new approach for measuring homonuclear 3-bond J(H(N)H(alpha) coupling constants in N-15-enriched proteins. J. Am. Chem. Soc. 115, 7772–7777 (1993).
Archer, S.J., Ikura, M., Torchia, D.A. & Bax, A. An alternative 3D-NMR technique for correlating backbone N-15 with side chain H-beta-resonances in larger proteins. J. Magn. Reson. 95, 636–641 (1991).
Clore, G.M., Bax, A. & Gronenborn, G.M. Stereospecific assignment of beta-methylene protons in larger proteins using 3D 15N-separated Hartman-Hahn and 13-separated rotating frame Overhauser spectroscopy. J. Biomol. NMR 1, 13–22 (1991).
Fesik, S.W. & Zuiderweg, E.R.P. Heteronuclear three-dimensional NMR spectroscopy. A strategy for the simplification of homonuclear two-dimensional NMR spectra. J. Magn. Reson. 78, 588–593 (1988).
Delaglio, F. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Garrett, D.S., Powers, R., Gronenborn, A.M. & Clore, G.M. A common sense approach to peak picking in two-, three-, and four-dimensional spectra using automated computer analysis of contour diagrams. J. Magn. Reson. 95, 214–220 (1991).
Brünger, A.T. X-PLOR (version 3.1): a system for X-ray crystallography and NMR (Yale University Press, New Haven, Connecticut; 1993).
Wojciak, J.M., Connolly, K.M. & Clubb, R.T. NMR structure of the Tn916 integrase-DNA complex. Nature Struct. Biol. 6, 366–373 (1999).
Nilges, M., Clore, G.M. & Gronenborn, A.M. Determination of three-dimensional structures of proteins from interproton distance data by hybrid distance geometry-dynamic simulated annealing. FEBS Lett. 229, 129–136 (1988).
Lavery, R. & Sklenar, H. The definition of generalized helicoidal parameters and of axis curvature for irregular nucleic acids. J. Biomol. Struct. Dyn. 6, 63–91 (1988).
Koradi, R., Billeter, M. & Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
Acknowledgements
We thank G. Daughdrill and the staff at the Pacific Northwest National Laboratory for assistance in operating their 800 MHz NMR spectrometer. We also thank K. Connolly for preparation of the Mu repressor expression plasmid and U. Ilangovan for useful discussions. This work was supported by grants from the US Department of Energy and National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wojciak, J., Iwahara, J. & Clubb, R. The Mu repressor–DNA complex contains an immobilized 'wing' within the minor groove. Nat Struct Mol Biol 8, 84–90 (2001). https://doi.org/10.1038/83103
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/83103