Key Points
-
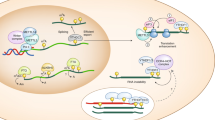
Multiple molecular events take place during the life of a B cell, first when the B-cell receptor (BCR) is assembled in the bone marrow during the generation of the pre-immune repertoire and later during an immune response in secondary lymphoid organs through the process of affinity maturation (known as somatic hypermutation, SHM) and isotype switching. Specific DNA polymerases are involved at each of these molecular steps.
-
Mammals have 15 DNA polymerases (including terminal deoxynucleotidyl transferase). Four are involved in semi-conservative replication and 11 are involved in different tasks, such as the removal of damaged bases (Polβ), the rejoining of DNA ends during non-homologous end joining (NHEJ) (Polλ and Polμ) and the bypass of DNA lesions that block the replication fork (Y family polymerases and Polζ).
-
The assembly of the immunoglobulin receptor involves the joining of immunoglobulin coding regions that are distantly located in the genome. These coding ends are subject to several enzymatic activities that include nucleotide trimming and template-dependent and -independent DNA synthesis. A large diversity is thus created at the sites of the exon junctions, by varying the length of the complementarity-determining region 3 (CDR3). Polλ and Polμ participate in these processes, being involved during the recombination of the heavy- and light-chain loci, respectively.
-
During SHM, mostly point mutations are introduced in the variable heavy- and light-chain genes of the BCR presented by B cells engaged in an immune response. The mechanism is initiated by the deamination of cytidines into uracils through the action of activation-induced cytidine deaminase (AID). Thereafter, these U-G mismatches are recognized by two different repair pathways, mediated by either uracil glycosylase (UNG) or the mismatch recognition complex, MSH2–MSH6. Two DNA polymerases have been shown unequivocally to be recruited during these error-prone repair processes: Polη, which generates all mutations at A/T bases during the normal physiological SHM process, and Rev1, which contributes some of the G/C mutation transversion mutations.
-
A model for SHM is proposed that attempts to include all the data collected from various mouse and human experimental settings. In this model, the two repair pathways involved in SHM appear to function outside of their normal roles in the base-excision repair or mismatch repair processes, and to compete rather than collaborate once the U-G mismatches have been generated by AID.
Abstract
To cope with an unpredictable variety of potential pathogenic insults, the immune system must generate an enormous diversity of recognition structures, and it does so by making stepwise modifications at key genetic loci in each lymphoid cell. These modifications proceed through the action of lymphoid-specific proteins acting together with the general DNA-repair machinery of the cell. Strikingly, these general mechanisms are usually diverted from their normal functions, being used in rather atypical ways in order to privilege diversity over accuracy. In this Review, we focus on the contribution of a set of DNA polymerases discovered in the past decade to these unique DNA transactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tonegawa, S. Somatic generation of antibody diversity. Nature 302, 575–581 (1983).
Rajewsky, K. Clonal selection and learning in the antibody system. Nature 381, 751–758 (1996).
Weill, J. C. & Reynaud, C. A. Rearrangement/hypermutation/gene conversion: when, where and why? Immunol. Today 17, 92–97 (1996).
Goodman, M. F. & Tippin, B. The expanding polymerase universe. Nature Rev. Mol. Cell Biol. 1, 101–109 (2000).
Burgers, P. M. et al. Eukaryotic DNA polymerases: proposal for a revised nomenclature. J. Biol. Chem. 276, 43487–43490 (2001).
Hubscher, U., Maga, G. & Spadari, S. Eukaryotic DNA polymerases. Annu. Rev. Biochem. 71, 133–163 (2002).
Friedberg, E. C., Wagner, R. & Radman, M. Specialized DNA polymerases, cellular survival, and the genesis of mutations. Science 296, 1627–1630 (2002).
Gellert, M. V(D)J recombination: RAG proteins, repair factors, and regulation. Annu. Rev. Biochem. 71, 101–132 (2002).
Schatz, D. G. & Spanopoulou, E. Biochemistry of V(D)J recombination. Curr. Top. Microbiol. Immunol. 290, 49–85 (2005).
Revy, P., Malivert, L. & de Villartay, J. P. Cernunnos-XLF, a recently identified non-homologous end-joining factor required for the development of the immune system. Curr. Opin. Allergy Clin. Immunol. 6, 416–420 (2006).
Mahajan, K. N., Nick McElhinny, S. A., Mitchell, B. S. & Ramsden, D. A. Association of DNA polymerase μ (polμ) with Ku and ligase IV: role for polμ in end-joining double-strand break repair. Mol. Cell Biol. 22, 5194–5202 (2002).
Ma, Y. et al. A biochemically defined system for mammalian nonhomologous DNA end joining. Mol. Cell 16, 701–713 (2004). This paper provides a detailed biochemical analysis of the relative contribution of TdT, Polλ and Polμ to DNA end processing during NHEJ in vitro.
Sobol, R. W. & Wilson, S. H. Mammalian DNA β-polymerase in base excision repair of alkylation damage. Prog. Nucleic Acid Res. Mol. Biol. 68, 57–74 (2001).
Wilson, T. E. & Lieber, M. R. Efficient processing of DNA ends during yeast nonhomologous end joining. Evidence for a DNA polymerase β (Pol4)-dependent pathway. J. Biol. Chem. 274, 23599–23609 (1999).
Aoufouchi, S. et al. Two novel human and mouse DNA polymerases of the polX family. Nucleic Acids Res. 28, 3684–3693 (2000).
Dominguez, O. et al. DNA polymerase μ (Polμ), homologous to TdT, could act as a DNA mutator in eukaryotic cells. EMBO J. 19, 1731–1742 (2000).
Garcia-Diaz, M. et al. DNA polymerase λ (Polλ), a novel eukaryotic DNA polymerase with a potential role in meiosis. J. Mol. Biol. 301, 851–867 (2000).
Lee, J. W. et al. Implication of DNA polymerase λ in alignment-based gap filling for nonhomologous DNA end joining in human nuclear extracts. J. Biol. Chem. 279, 805–811 (2004).
Daley, J. M., Laan, R. L., Suresh, A. & Wilson, T. E. DNA joint dependence of pol X family polymerase action in nonhomologous end joining. J. Biol. Chem. 280, 29030–29037 (2005).
Nick McElhinny, S. A. et al. A gradient of template dependence defines distinct biological roles for family X polymerases in nonhomologous end joining. Mol. Cell 19, 357–366 (2005).
Bertocci, B., De Smet, A., Weill, J. C. & Reynaud, C. A. Nonoverlapping functions of DNA polymerases μ, λ, and terminal deoxynucleotidyltransferase during immunoglobulin V(D)J recombination in vivo. Immunity 25, 31–41 (2006). This paper describes the independent and specific contribution of Polλ and Polμ to heavy-chain and light-chain V(D)J junctions, respectively.
Thai, T. H., Purugganan, M. M., Roth, D. B. & Kearney, J. F. Distinct and opposite diversifying activities of terminal transferase splice variants. Nature Immunol. 3, 457–462 (2002).
Repasky, J. A., Corbett, E., Boboila, C. & Schatz, D. G. Mutational analysis of terminal deoxynucleotidyltransferase-mediated N-nucleotide addition in V(D)J recombination. J. Immunol. 172, 5478–5488 (2004).
Doyen, N., Boule, J. B., Rougeon, F. & Papanicolaou, C. Evidence that the long murine terminal deoxynucleotidyltransferase isoform plays no role in the control of V(D)J junctional diversity. J. Immunol. 172, 6764–6767 (2004).
Ma, Y., Pannicke, U., Schwarz, K. & Lieber, M. R. Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination. Cell 108, 781–794 (2002).
Moshous, D. et al. Artemis, a novel DNA double-strand break repair/V(D)J recombination protein, is mutated in human severe combined immune deficiency. Cell 105, 177–186 (2001).
Yang, Y. G., Lindahl, T. & Barnes, D. E. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 131, 873–886 (2007).
Bertocci, B., De Smet, A., Berek, C., Weill, J. C. & Reynaud, C. A. Immunoglobulin κ light chain gene rearrangement is impaired in mice deficient for DNA polymerase μ. Immunity 19, 203–211 (2003).
Tomlinson, I. M., Cox, J. P., Gherardi, E., Lesk, A. M. & Chothia, C. The structural repertoire of the human Vκ domain. EMBO J. 14, 4628–4638 (1995).
Lederberg, J. Genes and antibodies. Science 129, 1649–1653 (1959).
Burnett, F. The clonal selection theory of acquired immunity (The University Press, Cambridge, 1959).
Weigert, M. G., Cesari, I. M., Yonkovich, S. J. & Cohn, M. Variability in the λ light chain sequences of mouse antibody. Nature 228, 1045–1047 (1970).
Bothwell, A. L. et al. Heavy chain variable region contribution to the NPb family of antibodies: somatic mutation evident in a γ2a variable region. Cell 24, 625–637 (1981).
Gearhart, P. J., Johnson, N. D., Douglas, R. & Hood, L. IgG antibodies to phosphorylcholine exhibit more diversity than their IgM counterparts. Nature 291, 29–34 (1981).
Selsing, E. & Storb, U. Somatic mutation of immunoglobulin light-chain variable-region genes. Cell 25, 47–58 (1981).
Griffiths, G. M., Berek, C., Kaartinen, M. & Milstein, C. Somatic mutation and the maturation of immune response to 2-phenyl oxazolone. Nature 312, 271–275 (1984).
Brenner, S. & Milstein, C. Origin of antibody variation. Nature 211, 242–243 (1966).
Muramatsu, M. et al. Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. J. Biol. Chem. 274, 18470–18476 (1999).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Revy, P. et al. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the hyper-IgM syndrome (HIGM2). Cell 102, 565–575 (2000).
Di Noia, J. M. & Neuberger, M. S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007).
Reynaud, C. A. et al. Mismatch repair and immunoglobulin gene hypermutation: did we learn something? Immunol. Today 20, 522–527 (1999).
Martin, A. & Scharff, M. D. AID and mismatch repair in antibody diversification. Nature Rev. Immunol. 2, 605–614 (2002).
Martomo, S. A. & Gearhart, P. J. Somatic hypermutation: subverted DNA repair. Curr. Opin. Immunol. 18, 243–248 (2006).
Esposito, G. et al. Mice reconstituted with DNA polymerase β-deficient fetal liver cells are able to mount a T cell-dependent immune response and mutate their Ig genes normally. Proc. Natl Acad. Sci. USA 97, 1166–1171 (2000).
Prakash, S., Johnson, R. E. & Prakash, L. Eukaryotic translesion synthesis DNA polymerases: specificity of structure and function. Annu. Rev. Biochem. 74, 317–353 (2005).
Goodman, M. F. Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu. Rev. Biochem. 71, 17–50 (2002).
Friedberg, E. C. Suffering in silence: the tolerance of DNA damage. Nature Rev. Mol. Cell Biol. 6, 943–953 (2005).
Zhang, Y. et al. Lesion bypass activities of human DNA polymerase μ. J. Biol. Chem. 277, 44582–44587 (2002).
Maga, G. et al. 8-oxo-guanine bypass by human DNA polymerases in the presence of auxiliary proteins. Nature 447, 606–608 (2007).
Seki, M. et al. High-efficiency bypass of DNA damage by human DNA polymerase Q. EMBO J. 23, 4484–4494 (2004).
Masutani, C. et al. The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature 399, 700–704 (1999).
Johnson, R. E., Kondratick, C. M., Prakash, S. & Prakash, L. hRAD30 mutations in the variant form of xeroderma pigmentosum. Science 285, 263–265 (1999).
Zeng, X. et al. DNA polymerase η is an A-T mutator in somatic hypermutation of immunoglobulin variable genes. Nature Immunol. 2, 537–541 (2001). The first demonstration of the contribution of a TLS polymerase to SHM, through the analysis of patients with the XP-V syndrome, a genetic disease caused by a deficiency in Polη activity.
Faili, A. et al. DNA polymerase η is involved in hypermutation occurring during immunoglobulin class switch recombination. J. Exp. Med. 199, 265–270 (2004).
Delbos, F. et al. Contribution of DNA polymerase η to immunoglobulin gene hypermutation in the mouse. J. Exp. Med. 201, 1191–1196 (2005).
Martomo, S. A. et al. Different mutation signatures in DNA polymerase η- and MSH6-deficient mice suggest separate roles in antibody diversification. Proc. Natl Acad. Sci. USA 102, 8656–8661 (2005).
Martomo, S. A. et al. Normal hypermutation in antibody genes from congenic mice defective for DNA polymerase ι. DNA Repair (Amst) 5, 392–398 (2006).
Matsuda, T. et al. Error rate and specificity of human and murine DNA polymerase ɛ. J. Mol. Biol. 312, 335–346 (2001).
Pavlov, Y. I. et al. Correlation of somatic hypermutation specificity and A-T base pair substitution errors by DNA polymerase ɛ during copying of a mouse immunoglobulin κ light chain transgene. Proc. Natl Acad. Sci. USA 99, 9954–9959 (2002).
Ohashi, E. et al. Fidelity and processivity of DNA synthesis by DNA polymerase κ, the product of the human DINB1 gene. J. Biol. Chem. 275, 39678–39684 (2000).
Zhang, Y. et al. Error-free and error-prone lesion bypass by human DNA polymerase κ in vitro. Nucleic Acids Res. 28, 4138–4146 (2000).
Schenten, D. et al. DNA polymerase κ deficiency does not affect somatic hypermutation in mice. Eur. J. Immunol. 32, 3152–3160 (2002).
Shimizu, T., Shinkai, Y., Ogi, T., Ohmori, H. & Azuma, T. The absence of DNA polymerase κ does not affect somatic hypermutation of the mouse immunoglobulin heavy chain gene. Immunol. Lett. 86, 265–270 (2003).
Delbos, F., Aoufouchi, S., Faili, A., Weill, J. C. & Reynaud, C. A. DNA polymerase η is the sole contributor of A/T modifications during immunoglobulin gene hypermutation in the mouse. J. Exp. Med. 204, 17–23 (2007). A paper establishing Polη as the sole contributor of A/T mutations during the normal SHM process in mice, and proposing that the two UNG and MSH2–MSH6 pathways that process uracils are competitive rather than alternative.
Nelson, J. R., Lawrence, C. W. & Hinkle, D. C. Deoxycytidyl transferase activity of yeast REV1 protein. Nature 382, 729–731 (1996).
Zhang, Y. et al. Response of human REV1 to different DNA damage: preferential dCMP insertion opposite the lesion. Nucleic Acids Res. 30, 1630–1638 (2002).
Guo, C. et al. Mouse Rev1 protein interacts with multiple DNA polymerases involved in translesion DNA synthesis. EMBO J. 22, 6621–6630 (2003).
Jansen, J. G. et al. The BRCT domain of mammalian Rev1 is involved in regulating DNA translesion synthesis. Nucleic Acids Res. 33, 356–365 (2005).
Jansen, J. G. et al. Strand-biased defect in C/G. transversions in hypermutating immunoglobulin genes in Rev1-deficient mice. J. Exp. Med. 203, 319–323 (2006). This paper demonstrates the contribution of Rev1 to G/C transversion mutations, with an unexpected strand bias.
Johnson, R. E., Washington, M. T., Haracska, L., Prakash, S. & Prakash, L. Eukaryotic polymerases ι and ζ act sequentially to bypass DNA lesions. Nature 406, 1015–1019 (2000).
Esposito, G. et al. Disruption of the Rev3l-encoded catalytic subunit of polymerase ζ in mice results in early embryonic lethality. Curr. Biol. 10, 1221–1224 (2000).
Bemark, M., Khamlichi, A. A., Davies, S. L. & Neuberger, M. S. Disruption of mouse polymerase ζ (Rev3) leads to embryonic lethality and impairs blastocyst development in vitro. Curr. Biol. 10, 1213–1216 (2000).
Wittschieben, J. et al. Disruption of the developmentally regulated Rev3l gene causes embryonic lethality. Curr. Biol. 10, 1217–1220 (2000).
Zhong, X. et al. The fidelity of DNA synthesis by yeast DNA polymerase ζ alone and with accessory proteins. Nucleic Acids Res. 34, 4731–4742 (2006).
Diaz, M., Verkoczy, L. K., Flajnik, M. F. & Klinman, N. R. Decreased frequency of somatic hypermutation and impaired affinity maturation but intact germinal center formation in mice expressing antisense RNA to DNA polymerase ζ. J. Immunol. 167, 327–335 (2001).
Zan, H. et al. The translesion DNA polymerase ζ plays a major role in Ig and bcl-6 somatic hypermutation. Immunity 14, 643–653 (2001).
Tissier, A., McDonald, J. P., Frank, E. G. & Woodgate, R. polι, a remarkably error-prone human DNA polymerase. Genes Dev. 14, 1642–1650 (2000).
Frank, E. G. & Woodgate, R. Increased catalytic activity and altered fidelity of human DNA polymerase ι in the presence of manganese. J. Biol. Chem. 282, 24689–24696 (2007).
Sale, J. E. & Neuberger, M. S. TdT-accessible breaks are scattered over the immunoglobulin V domain in a constitutively hypermutating B cell line. Immunity 9, 859–869 (1998).
Denepoux, S. et al. Induction of somatic mutation in a human B cell line in vitro. Immunity 6, 35–46 (1997).
Zan, H. et al. Induction of Ig somatic hypermutation and class switching in a human monoclonal IgM+ IgD+ B cell line in vitro: definition of the requirements and modalities of hypermutation. J. Immunol. 162, 3437–3447 (1999).
Faili, A. et al. AID-dependent somatic hypermutation occurs as a DNA single-strand event in the BL2 cell line. Nature Immunol. 3, 815–821 (2002).
Xiao, Z. et al. Known components of the immunoglobulin A:T mutational machinery are intact in Burkitt lymphoma cell lines with G:C bias. Mol. Immunol. 44, 2659–2666 (2007).
Faili, A. et al. Induction of somatic hypermutation in immunoglobulin genes is dependent on DNA polymerase ι. Nature 419, 944–947 (2002).
McDonald, J. P. et al. 129-derived strains of mice are deficient in DNA polymerase ι and have normal immunoglobulin hypermutation. J. Exp. Med. 198, 635–643 (2003).
Poltoratsky, V., Prasad, R., Horton, J. K. & Wilson, S. H. Down-regulation of DNA polymerase β accompanies somatic hypermutation in human BL2 cell lines. DNA Repair (Amst) 6, 244–253 (2007).
Zan, H. et al. The translesion DNA polymerase θ plays a dominant role in immunoglobulin gene somatic hypermutation. EMBO J. 24, 3757–3769 (2005).
Masuda, K. et al. Absence of DNA polymerase θ results in decreased somatic hypermutation frequency and altered mutation patterns in Ig genes. DNA Repair (Amst) 5, 1384–1391 (2006).
Masuda, K. et al. DNA polymerases η and θ function in the same genetic pathway to generate mutations at A/T during somatic hypermutation of Ig genes. J. Biol. Chem. 282, 17387–17394 (2007).
Masuda, K. et al. DNA polymerase θ contributes to the generation of C/G mutations during somatic hypermutation of Ig genes. Proc. Natl Acad. Sci. USA 102, 13986–13991 (2005).
Kannouche, P. L., Wing, J. & Lehmann, A. R. Interaction of human DNA polymerase η with monoubiquitinated PCNA: a possible mechanism for the polymerase switch in response to DNA damage. Mol. Cell 14, 491–500 (2004).
Warbrick, E. PCNA binding through a conserved motif. Bioessays 20, 195–199 (1998).
Bienko, M. et al. Ubiquitin-binding domains in Y-family polymerases regulate translesion synthesis. Science 310, 1821–1824 (2005).
Langerak, P., Nygren, A. O., Krijger, P. H., van den Berk, P. C. & Jacobs, H. A/T mutagenesis in hypermutated immunoglobulin genes strongly depends on PCNAK164 modification. J. Exp. Med. 204, 1989–1998 (2007). This reference provides a demonstration of the major role of PCNA ubiquitylation in the A/T mutation pathway.
Moldovan, G. L., Pfander, B. & Jentsch, S. PCNA, the maestro of the replication fork. Cell 129, 665–679 (2007).
Rada, C., Di Noia, J. M. & Neuberger, M. S. Mismatch recognition and uracil excision provide complementary paths to both Ig switching and the A/T-focused phase of somatic mutation. Mol. Cell 16, 163–171 (2004). A landmark paper establishing that uracils produced by AID-mediated cytidine deamination are processed by only two pathways, UNG and MSH2–MSH6. The mutations observed in the MSH2 and UNG double-deficient background therefore represent the footprint of AID deamination at the immunoglobulin locus.
Rada, C. et al. Immunoglobulin isotype switching is inhibited and somatic hypermutation perturbed in UNG-deficient mice. Curr. Biol. 12, 1748–1755 (2002).
Wiesendanger, M., Kneitz, B., Edelmann, W. & Scharff, M. D. Somatic hypermutation in MutS homologue (MSH)3-, MSH6-, and MSH3/MSH6-deficient mice reveals a role for the MSH2–MSH6 heterodimer in modulating the base substitution pattern. J. Exp. Med. 191, 579–584 (2000).
Martomo, S. A., Yang, W. W. & Gearhart, P. J. A role for Msh6 but not Msh3 in somatic hypermutation and class switch recombination. J. Exp. Med. 200, 61–68 (2004).
Shen, H. M., Tanaka, A., Bozek, G., Nicolae, D. & Storb, U. Somatic hypermutation and class switch recombination in Msh6−/−Ung−/− double-knockout mice. J. Immunol. 177, 5386–5392 (2006).
Bardwell, P. D. et al. Altered somatic hypermutation and reduced class-switch recombination in exonuclease 1-mutant mice. Nature Immunol. 5, 224–229 (2004).
Phung, Q. H., Winter, D. B., Alrefai, R. & Gearhart, P. J. Hypermutation in Ig V genes from mice deficient in the MLH1 mismatch repair protein. J. Immunol. 162, 3121–3124 (1999).
Unniraman, S. & Schatz, D. G. Strand-biased spreading of mutations during somatic hypermutation. Science 317, 1227–1230 (2007).
Avkin, S., Adar, S., Blander, G. & Livneh, Z. Quantitative measurement of translesion replication in human cells: evidence for bypass of abasic sites by a replicative DNA polymerase. Proc. Natl Acad. Sci. USA 99, 3764–3769 (2002).
Zeng, X., Negrete, G. A., Kasmer, C., Yang, W. W. & Gearhart, P. J. Absence of DNA polymerase η reveals targeting of C mutations on the nontranscribed strand in immunoglobulin switch regions. J. Exp. Med. 199, 917–924 (2004).
Wu, X. & Stavnezer, J. DNA polymerase β is able to repair breaks in switch regions and plays an inhibitory role during immunoglobulin class switch recombination. J. Exp. Med. 204, 1677–1689 (2007).
Buerstedde, J. M. et al. Light chain gene conversion continues at high rate in an ALV-induced cell line. EMBO J. 9, 921–927 (1990).
Kim, S., Humphries, E. H., Tjoelker, L., Carlson, L. & Thompson, C. B. Ongoing diversification of the rearranged immunoglobulin light-chain gene in a bursal lymphoma cell line. Mol. Cell Biol. 10, 3224–3231 (1990).
Harris, R. S., Sale, J. E., Petersen-Mahrt, S. K. & Neuberger, M. S. AID is essential for immunoglobulin V gene conversion in a cultured B cell line. Curr. Biol. 12, 435–438 (2002).
Arakawa, H., Hauschild, J. & Buerstedde, J. M. Requirement of the activation-induced deaminase (AID) gene for immunoglobulin gene conversion. Science 295, 1301–1306 (2002).
Arakawa, H., Saribasak, H. & Buerstedde, J. M. Activation-induced cytidine deaminase initiates immunoglobulin gene conversion and hypermutation by a common intermediate. PLoS Biol. 2, E179 (2004).
Kawamoto, T. et al. Dual roles for DNA polymerase η in homologous DNA recombination and translesion DNA synthesis. Mol. Cell 20, 793–799 (2005).
Reynaud, C. A., Aoufouchi, S., Faili, A. & Weill, J. C. What role for AID: mutator, or assembler of the immunoglobulin mutasome? Nature Immunol. 4, 631–638 (2003).
McIlwraith, M. J. et al. Human DNA polymerase η promotes DNA synthesis from strand invasion intermediates of homologous recombination. Mol. Cell 20, 783–792 (2005).
Taddei, F. et al. Role of mutator alleles in adaptive evolution. Nature 387, 700–702 (1997).
Yeiser, B., Pepper, E. D., Goodman, M. F. & Finkel, S. E. SOS-induced DNA polymerases enhance long-term survival and evolutionary fitness. Proc. Natl Acad. Sci. USA 99, 8737–8741 (2002).
Saribasak, H. et al. Uracil DNA glycosylase disruption blocks Ig gene conversion and induces transition mutations. J. Immunol. 176, 365–371 (2006).
Simpson, L. J. & Sale, J. E. Rev1 is essential for DNA damage tolerance and non-templated immunoglobulin gene mutation in a vertebrate cell line. EMBO J. 22, 1654–1664 (2003).
Arakawa, H. et al. A role for PCNA ubiquitination in immunoglobulin hypermutation. PLoS Biol. 4, e366 (2006).
Acknowledgements
We thank our colleagues at INSERM U783 for contributing to this Review by many discussions and for sharing unpublished data. We apologize to those whose work could not be cited because of space limitations.
Author information
Authors and Affiliations
Related links
Glossary
- B-cell receptor
-
(BCR). A term used to refer to the immunoglobulin molecule present at the B-cell surface, irrespective of its heavy chain isotype.
- V(D)J recombination
-
Somatic rearrangement of variable (V), diversity (D) and joining (J) regions of the genes that encode antigen receptors, leading to repertoire diversity of both T-cell and B-cell receptors.
- Somatic hypermutation
-
(SHM). A process by which antigen-activated B cells in germinal centres acquire point mutations targeted to the variable regions of rearranged immunoglobulin gene segments. The B cells are subsequently selected for those expressing the 'best' mutations on the basis of the ability of the surface immunoglobulin to bind antigen. This process occurs in activated B cells in all jawed vertebrates, but it also occurs in immature B cells in sheep to generate the pre-immune repertoire.
- Class-switch recombination
-
(CSR). Also known as class or isotype switching, this term refers to a region-specific recombination process that occurs in antigen-activated B cells. Recombination takes place between 'switch region' DNA sequences located upstream from each immunoglobulin heavy-chain constant region genes and results in a change from the IgM to one of the IgG, IgA or IgE isotypes. This imparts flexibility to the humoral immune response and allows it to exploit the different capacities of the immunoglobulins to activate the appropriate downstream effector mechanisms.
- Gene conversion
-
A non-reciprocal homologous recombination event in which the donor gene(s) remains unmodified and an acceptor gene acquires the recombined segment. In chickens, variable (V) pseudogenes are donors that modify the functional, rearranged V gene in follicles of the bursa of Fabricius to generate a diverse pre-immune repertoire.
- Base-excision repair
-
A DNA-repair pathway that removes single non-canonical bases from the DNA, such as deaminated or oxidized bases, and replaces them with an appropriate base templated on the opposite DNA strand. Repair is initiated by a DNA glycosylase that is specialized for a particular type of damage, and nucleotide replacement is performed by DNA polymerase β.
- Non-homologous end joining
-
(NHEJ). A process that joins broken DNA ends independently of extended base homology. Components of this pathway include the proteins Ku70, Ku80, Artemis, X-ray repair cross-complementing protein 4 (XRCC4), DNA ligase IV, Cernunnos (also known as XLF) and the catalytic subunit of DNA-dependent protein kinase (DNA-PKcs).
- Replication fork
-
The site in double-stranded DNA at which the template strands are separated, allowing a newly formed copy of the DNA to be synthesized, with the fork moving in the direction of leading strand synthesis.
- Translesion DNA synthesis
-
(TLS). A process that allows the bypass of DNA damage that otherwise blocks the progression of the replication fork, through the replacement of the replicative polymerase with specialized polymerases (known as TLS polymerases) that can copy non-instructive DNA lesions. TLS polymerases have low fidelity on undamaged templates. Most TLS polymerases belong to the Y family polymerases.
- Recombination-activating genes
-
(RAG1 and RAG2). RAG1 and RAG2 are involved in creating the synapsis and the double-strand DNA breaks required for the assembly of the dispersed gene segments that encode the complete protein chains of B-cell and T-cell receptors.
- Synapsis
-
Non-covalent juxtaposition of two non-adjacent stretches of DNA.
- Recombination signal sequences
-
(RSSs). Conserved elements that constitute recognition sites for the V(D)J recombinase proteins that are encoded by recombination-activating gene 1 (RAG1) and RAG2. They consist of a palindromic heptamer that is immediately adjacent to the coding gene segments — V (variable), D (diversity) or J (joining) — and is separated by a 12- or 23-base-pair spacer from a conserved nonamer sequence.
- Complementarity-determining regions
-
(CDRs). The three hypervariable antigen-receptor regions (in both chains of the T-cell and B-cell receptors) that interact with the antigen. The third CDR (CDR3) is partly encoded by the germline variable (V), diversity (D) and joining (J) regions of each receptor chain. Extensive diversity is generated in CDR3 during gene rearrangement by nucleotide trimming and/or template-independent nucleotide additions by terminal deoxynucleotidyl transferase.
- Deamination
-
Removal of an amine group from a pyrimidine or purine nucleic-acid base. Deamination of cytosine and adenosine yields uracil and inosine, respectively.
- Mismatch repair
-
A repair pathway that recognizes mismatched bases that arise in DNA because of errors made by replicative DNA polymerases. These are then repaired by an excision system that removes a tract of DNA including the mismatch, and re-copies the original strand. The mismatch repair complex includes a mismatch-binding moiety (such as MSH2–MSH6), and an effector part (such as MLH1–PMS2) that triggers excision of the mismatch-containing DNA sequence.
- Abasic site
-
A common form of DNA damage in which a base is lost from a strand of DNA, spontaneously or by the action of DNA repair enzymes, such as uracil glycosylase, while leaving the phosphodiester bond intact.
- Xeroderma pigmentosum
-
(XP). A rare inherited human disorder, in which patients are sensitive to the DNA-damaging effects of sunlight. XP can be caused by disabling any of eight different genes. Seven of the genes, denoted XPA–XPG, encode components of the nucleotide excision-repair pathway. A variant form of the syndrome, XPV, corresponds to the inactivation of the gene encoding DNA polymerase η.
- Transition mutations
-
Base changes in DNA in which a pyrimidine (C or T) is replaced by another pyrimidine, or a purine (A or G) is replaced by another purine.
- Hotspot motif
-
A short DNA motif (DGYW or WRCH; where D denotes adenosine (A), guanosine (G) or thymidine (T); Y denotes cytidine (C) or T; W denotes A or T; R denotes A or G; and H denotes T, C or A) at which mutations are preferentially targeted during somatic hypermutation.
- Transversion mutations
-
Base changes in DNA in which a pyrimidine (C or T) is replaced by a purine (A or G), or a purine is replaced by a pyrimidine.
- Proliferating cell nuclear antigen
-
(PCNA). PCNA was first identified as a DNA sliding clamp for replicative polymerases, but is now known to coordinate the organization of protein partners that are involved in many processes, including DNA replication, DNA repair and cell-cycle control.
Rights and permissions
About this article
Cite this article
Weill, JC., Reynaud, CA. DNA polymerases in adaptive immunity. Nat Rev Immunol 8, 302–312 (2008). https://doi.org/10.1038/nri2281
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri2281