Abstract
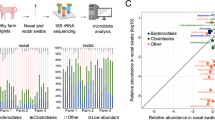
Upper respiratory tract (URT) infection caused the leading and devastating diseases in pigs. It was believed that the normal microbiome of URT plays a vital role in health and disease development. As the entry point of the URT, little knowledge of bacterial microbiome in porcine nasal was known. A cultivation-independent approach directly to 16s ribosomal RNA genes enabled us to reveal the nasal bacterial community, structure and diversity. Here, we found that an unprecedented 207 phylotypes were characterized from 933 qualified clones, indicating the variable, species richness but particularly dominant bacterial microbiome. The dominant species were from genus Comamonas and Acinetobacter, which constitute core normal bacterial microbiome in porcine nasal. Moreover, a set of swine specific pathogens and zoonotic agents were detected in the swine nasal microbiome. Collectively, we provided a snapshot of our current knowledge of the community structure of porcine nasal bacterial ecosystem in a healthy family that will further enhance our view to understand URT infection and public health.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yue, M., Bei, W. & Chen, H. Nasal Bacterial Microbiome: Probing a Healthy Porcine Family. Nat Prec (2011). https://doi.org/10.1038/npre.2011.6699.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.6699.1