Abstract
Background.
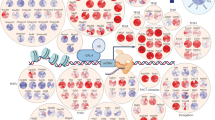
Many transcriptional regulatory proteins have been identified and classified into several families on the basis of sequence similarity (Fukami, 2003), these families usually regulate important biological processes, such as development and responses to external stimuli. Transcription factors (TFs) regulate spatial and temporal gene expression by binding to DNA and either activating or repressing the action of RNA polymerases (Latchman, 2004); in addition to TFs, other Transcriptional regulators (TRs) participate in transcriptional modulation by e.g., chromatin remodeling. With the availability of genome sequences for several organisms and computational strategies for gene functional annotation, the entire set of Transcription factors (TFs) and Transcription regulators (TRs) can be identified, described, and compared between species and lineages. The diversity among Stramenopiles is striking; they range from large multicellular seaweeds to tiny unicellular species, they are present in freshwater, marine and terrestrial habitats and embrace many ecologically important algal (e.g. diatoms, brown algae, chrysophytes), and heterotrophic (e.g., Oomycetes) groups.Methods.In order to find TF and TR genes in the deduced proteomes of Stramenopile we followed the approach developed in Perez et al. 2010. Briefly, it exploits the presence of protein domains and their combinations, in the form of boolean rules, that are specific for different families of TFs and TRs. Such rules are represented as a bipartite graph in which a set of nodes represent gene families and the other set of nodes, protein domains. Edges are drawn between nodes of different types indicating whether a protein domain is required in the gene family or it is forbidden.
Results.
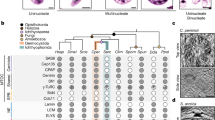
We applied an enlarged set of rules to the deduced proteomes of 8 different Stramenopiles, identifying more than 400 different regulatory genes in each species belonging to up to 59 different gene families.
Conclusions.
The identification of this class of regulatory genes will constitute and important resource that could be exploited in gene functional characterization and evolutionary analyses. All TFs and TRs families found will be publicly released via a web database.
References.
Pérez-Rodríguez P, Riaño-Pachón DM, Corrêa LG, Rensing SA, Kersten B, Mueller-Roeber B. 2010. Nucleic Acids Res. 38(Database issue):D822-7
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Buitrago, F., Restrepo, S. & Riaño-Pachón, D. Identification and Evolution of Transcription Factors in Stramenopiles. Nat Prec (2011). https://doi.org/10.1038/npre.2011.6400.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.6400.1