Abstract
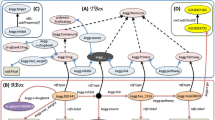
Integration of the scientific literature into a biomedical research infrastructure requires the processing of the literature, identification of the contained named entities (NEs) and concepts, and to represent the content in a standardised way.The CALBC project partners (PPs) have produced a large-scale annotated biomedical corpus with four different semantic groups through the harmonisation of annotations from automatic text mining solutions (Silver Standard Corpus, SSC). The four semantic groups were chemical entities and drugs (CHED), genes and proteins (PRGE), diseases and disorders (DISO) and species (SPE). The content of the SSC has been fully integrated into RDF Triple Store (4,568,678 triples) and has been aligned with content from the GeneAtlas (182,840 triples), UniProtKb (12,552,239 triples for human) and the lexical resource LexEBI (BioLexicon). RDF Triple Store enables querying the scientific literature and bioinformatics resources at the same time for evidence of genetic causes, such as drug targets and disease involvement.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Croset, S. The CALBC RDF Triple store: retrieval over large literature content. Nat Prec (2010). https://doi.org/10.1038/npre.2010.5383.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2010.5383.1