Abstract
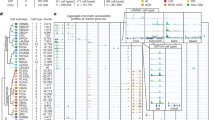
This report contains a gene expression summary of the inferior colliculus (IC), derived from the Allen Brain Atlas (ABA) in-situ hybridization (ISH) mouse data set. The structure’s location and morphological characteristics in the mouse brain are described using the Nissl data found in the Allen Reference Atlas. Using an established algorithm, the expression values of the IC were compared to the values of the macro/parent-structure, in this case the midbrain, for the purpose of extracting regionally specific gene expression data. The highest ranking ratios were then manually curated and verified. The 50 Select Genes were compiled for expression characterization. The experimental data for each gene may be accessed via the links provided; complementary sagittal data may also be accessed using the ABA. Correlation between gene expression in the IC and the rest of the brain, across all genes in the coronal dataset (~4300 genes), were derived computationally and are presented below. A gene ontology table (derived from DAVID Bioinformatics Resources 2007) is also included, highlighting possible functions of these 50 Select Genes.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Consortia
Rights and permissions
About this article
Cite this article
Allen Institute for Brain Science., Solan, B., Ng, L. et al. Inferior Colliculus . Nat Prec (2008). https://doi.org/10.1038/npre.2008.2033.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2008.2033.1