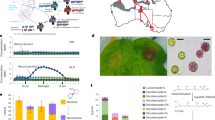
Abstract
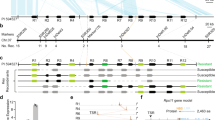
A single lineage of Nicotiana benthamiana is widely used as a model plant1 and has been instrumental in making revolutionary discoveries about RNA interference (RNAi), viral defence and vaccine production. It is peerless in its susceptibility to viruses and its amenability in transiently expressing transgenes2,3. These unparalleled characteristics have been associated both positively and negatively with a disruptive insertion in the RNA-dependent RNA polymerase 1 gene, Rdr14–6. For a plant so routinely used in research, the origin, diversity and evolution of the species, and the basis of its unusual abilities, have been relatively unexplored. Here, by comparison with wild accessions from across the spectrum of the species’ natural distribution, we show that the laboratory strain of N. benthamiana is an extremophile originating from a population that has retained a mutation in Rdr1 for ∼0.8 Myr and thereby traded its defence capacity for early vigour and survival in the extreme habitat of central Australia. Reconstituting Rdr1 activity in this isolate provided protection. Silencing the functional allele in a wild strain rendered it hypersusceptible and was associated with a doubling of seed size and enhanced early growth rate. These findings open the way to a deeper understanding of the delicate balance between protection and vigour.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Goodin, M. M., Zaitlin, D., Naidu, R. A. & Lommel, S. A. Nicotiana benthamiana: its history and future as a model for plant-pathogen interactions. Mol. Plant Microbe Interact. 21, 1015–1026 (2008).
Clemente, T. in Nicotiana (Nicotiana tobaccum, Nicotiana benthamiana) (ed Wang, K. ) Vol. 343, 143–154 (Humana, 2006).
Horsch, R. B. et al. A simple and general method of transferring genes into plants. Science 227, 1229–1231 (1985).
Yang, S. J., Carter, S. A., Cole, A. B., Cheng, N. H. & Nelson, R. S. A natural variant of a host RNA-dependent RNA polymerase is associated with increased susceptibility to viruses by Nicotiana benthamiana. Proc. Natl Acad. Sci. USA 101, 6297–6302 (2004).
Ying, X. B. et al. RNA-dependent RNA polymerase 1 from Nicotiana tabacum suppresses RNA silencing and enhances viral infection in Nicotiana benthamiana. Plant Cell 22, 1358–1372 (2010).
Akhtar, S., Briddon, R. & Mansoor, S. Reactions of Nicotiana species to inoculation with monopartite and bipartite begomoviruses. Virol. J. 8, 475 (2011).
Burbidge, N. T. The Australian species of Nicotiana L. (Solanaceae). Aust. J. Bot. 8, 342–378 (1960).
King, A. M. Q., Adams, M. J., Carstens, E. B. & Lefkowitz, E. J. (eds). Virus Taxonomy – Ninth Report of the International Committee on Taxonomy of Viruses. (Elsevier/Academic Press, 2011).
Garcia-Ruiz, H. et al. Arabidopsis RNA-dependent RNA polymerases and dicer-like proteins in antiviral defense and small interfering RNA biogenesis during Turnip Mosaic Virus infection. Plant cell 22, 481–496 (2010).
Nakasugi, K. et al. De novo transcriptome sequence assembly and analysis of RNA silencing genes of Nicotiana benthamiana. PLoS ONE 8, e59534 (2013).
Nakasugi, K., Crowhurst, R., Bally, J. & Waterhouse, P. Combining transcriptome assemblies from multiple de novo assemblers in the allo-tetraploid plant Nicotiana benthamiana. PLoS ONE 9, e91776 (2014).
Naim, F. et al. Advanced engineering of lipid metabolism in Nicotiana benthamiana using a draft genome and the V2 viral silencing-suppressor protein. PLoS ONE 7, e52717 (2012).
Duarte, J. M. et al. Identification of shared single copy nuclear genes in Arabidopsis, Populus, Vitis and Oryza and their phylogenetic utility across various taxonomic levels. BMC Evol. Biol. 10, 61 (2010).
Ladiges, P. Y., Marks, C. E. & Nelson, G. Biogeography of Nicotiana Section Suaveolentes (Solanaceae) reveals geographic tracks in arid Australia. J. Biogeog. 38, 2066–2077 (2011).
Byrne, M. et al. Birth of a biome: insights into the assembly and maintenance of the Australian arid zone biota. Mol. Ecol. 17, 4398–4417 (2008).
Mauricio, R. Costs of resistance to natural enemies in field populations of the annual plant Arabidopsis thaliana. Am. Nat. 151, 20–28 (1998).
Todesco, M. et al. Natural allelic variation underlying a major fitness trade-off in Arabidopsis thaliana. Nature 465, 632–636 (2010).
Zavala, J. A., Patankar, A. G., Gase, K. & Baldwin, I. T. Constitutive and inducible trypsin proteinase inhibitor production incurs large fitness costs in Nicotiana attenuata. Proc. Natl Acad. Sci. USA 101, 1607–1612 (2004).
Ruiz, M. T., Voinnet, O. & Baulcombe, D. C. Initiation and maintenance of virus-induced gene silencing. Plant Cell 10, 937–946 (1998).
Marks, C. E., Newbigin, E. & Ladiges, P. Y. Comparative morphology and phylogeny of Nicotiana section Suaveolentes (Solanaceae) in Australia and the South Pacific. Aust. Syst. Bot. 24, 61–86 (2011).
Fusaro, A. F. et al. The Enamovirus P0 protein is a silencing suppressor which inhibits local and systemic RNA silencing through AGO1 degradation. Virology 426, 178–187 (2012).
Eamens, A. L. & Waterhouse, P. M. Vectors and methods for hairpin RNA and artificial microRNA-mediated gene silencing in plants. Methods Mol. Biol. 701, 179–197 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Ronquist, F. & Huelsenbeck, J. P. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Drummond, A. J., Ho, S. Y. W., Phillips, M. J. & Rambaut, A. “Relaxed phylogenetics and dating with confidence”. PLoS Biol. 4, e88 (2006).
Acknowledgements
We are grateful to D. Albrecht, T. Willing, K. Courtney, T. Bean, J. Randles, E. Newbiggin, P. Ladiges and E. Dodds for their help, resource provision and discussions, and K. Lee for excellent technical assistance.
Author information
Authors and Affiliations
Contributions
J.B. and P.W. designed the experiments; J.B., F.J., M.W., C.P. and F.N. performed the experiments; J.B., K.N., H.J., P.W., S.H., R.C., C.W., R.H. and J.D. analysed data; J.B. and P.W. wrote the paper. All the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Bally, J., Nakasugi, K., Jia, F. et al. The extremophile Nicotiana benthamiana has traded viral defence for early vigour. Nature Plants 1, 15165 (2015). https://doi.org/10.1038/nplants.2015.165
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2015.165
This article is cited by
-
Nicotiana benthamiana as a model plant host for Fusarium verticillioides to investigate RNA interference, cross-kingdom RNA exchange, and competitive endogenous RNA (ceRNA) network
Molecular Biology Reports (2023)
-
A multi-omic Nicotiana benthamiana resource for fundamental research and biotechnology
Nature Plants (2023)
-
Into a dilemma of plants: the antagonism between chemical defenses and growth
Plant Molecular Biology (2022)
-
Carbon nanotube biocompatibility in plants is determined by their surface chemistry
Journal of Nanobiotechnology (2021)
-
Metabolic effects of agro-infiltration on N. benthamiana accessions
Transgenic Research (2021)