Abstract
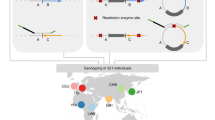
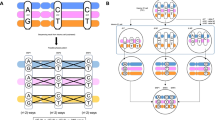
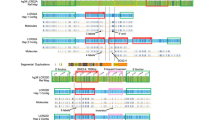
Inversions are an important form of structural variation, but they are difficult to characterize, as their breakpoints often fall within inverted repeats. We have developed a method called 'haplotype fusion' in which an inversion breakpoint is genotyped by performing fusion PCR on single molecules of human genomic DNA. Fusing single-copy sequences bracketing an inversion breakpoint generates orientation-specific PCR products, exemplified by a genotyping assay for the int22 hemophilia A inversion on Xq28. Furthermore, we demonstrated that inversion events with breakpoints embedded within long (>100 kb) inverted repeats can be genotyped by haplotype-fusion PCR followed by bead-based single-molecule haplotyping on repeat-specific markers bracketing the inversion breakpoint. We illustrate this method by genotyping a Yp paracentric inversion sponsored by >300-kb-long inverted repeats. The generality of our methods to survey for, and genotype chromosomal inversions should help our understanding of the contribution of inversions to genomic variation, inherited diseases and cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hogervorst, F.B. et al. Large genomic deletions and duplications in the BRCA1 gene identified by a novel quantitative method. Cancer Res. 63, 1449–1453 (2003).
Stankiewicz, P. & Lupski, J.R. Genome architecture, rearrangements and genomic disorders. Trends Genet. 18, 74–82 (2002).
Stefansson, H. et al. A common inversion under selection in Europeans. Nat. Genet. 37, 129–137 (2005).
Iafrate, A.J. et al. Detection of large-scale variation in the human genome. Nat. Genet. 36, 949–951 (2004).
Sebat, J. et al. Large-scale copy number polymorphism in the human genome. Science 305, 525–528 (2004).
Sharp, A.J. et al. Segmental duplications and copy-number variation in the human genome. Am. J. Hum. Genet. 77, 78–88 (2005).
Armour, J.A., Barton, D.E., Cockburn, D.J. & Taylor, G.R. The detection of large deletions or duplications in genomic DNA. Hum. Mutat. 20, 325–337 (2002).
Tuzun, E. et al. Fine-scale structural variation of the human genome. Nat. Genet. 37, 727–732 (2005).
Bunge, S. et al. Homologous nonallelic recombinations between the iduronate-sulfatase gene and pseudogene cause various intragenic deletions and inversions in patients with mucopolysaccharidosis type II. Eur. J. Hum. Genet. 6, 492–500 (1998).
Rossiter, J.P. et al. Factor VIII gene inversions causing severe hemophilia A originate almost exclusively in male germ cells. Hum. Mol. Genet. 3, 1035–1039 (1994).
Osborne, L.R. et al. A 1.5 million-base pair inversion polymorphism in families with Williams-Beuren syndrome. Nat. Genet. 29, 321–325 (2001).
Liu, Q., Nozari, G. & Sommer, S.S. Single-tube polymerase chain reaction for rapid diagnosis of the inversion hotspot of mutation in hemophilia A. Blood 92, 1458–1459 (1998).
Small, K., Iber, J. & Warren, S.T. Emerin deletion reveals a common X-chromosome inversion mediated by inverted repeats. Nat. Genet. 16, 96–99 (1997).
Antonarakis, S.E. et al. Factor VIII gene inversions in severe hemophilia A: results of an international consortium study. Blood 86, 2206–2212 (1995).
Jobling, M.A. et al. A selective difference between human Y-chromosomal DNA haplotypes. Curr. Biol. 8, 1391–1394 (1998).
Wetmur, J.G. et al. Molecular haplotyping by linking emulsion PCR: analysis of paraoxonase 1 haplotypes and phenotypes. Nucleic Acids Res. 33, 2615–2619 (2005).
Yon, J. & Fried, M. Precise gene fusion by PCR. Nucleic Acids Res. 17, 4895 (1989).
Dressman, D. et al. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc. Natl. Acad. Sci. USA 100, 8817–8822 (2003).
Ghadessy, F.J., Ong, J.L. & Holliger, P. Directed evolution of polymerase function by compartmentalized self-replication. Proc. Natl. Acad. Sci. USA 98, 4552–4557 (2001).
Bagnall, R.D., Giannelli, F. & Green, P.M. Polymorphism and hemophilia A causing inversions in distal Xq28: a complex picture. J. Thromb. Haemost. 3, 2598–2599 (2005).
Ross, M.T. et al. The DNA sequence of the human X chromosome. Nature 434, 325–337 (2005).
Vogt, P.H. et al. Report of the third international workshop on Y chromosome mapping 1997. Cytogenet. Cell Genet. 79, 2–16 (1997).
Skaletsky, H. et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 423, 825–837 (2003).
Williams, G. Mapping Studies of the Centromeric Region of the Human Y Chromosome. D. Phil. Thesis. University of Oxford, UK (1998).
Cooper, K.F., Fisher, R.B. & Tyler-Smith, C. Structure of the sequences adjacent to the centromeric alphoid satellite DNA array on the human Y-chromosome. J. Mol. Biol. 230, 787–799 (1993).
Mitra, R.D. et al. Digital genotyping and haplotyping with polymerase colonies. Proc. Natl. Acad. Sci. USA 100, 5926–5931 (2003).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
She, X. et al. Shotgun sequence assembly and recent segmental duplications within the human genome. Nature 431, 927–930 (2004).
Lupski, J.R. Hotspots of homologous recombination in the human genome: not all homologous sequences are equal. Genome Biol. 5, 242 (2004).
Feuk, L. et al. Discovery of human inversion polymorphisms by comparative analysis of human and chimpanzee DNA sequence assemblies. PLoS Genet. 1, e56 (2005).
Broman, K.W. et al. Common long human inversion polymorphism on chromosome 8p′. In Science and Statistics: A Festschrift for Terry Speed (ed, Goldstein, D.R.) 237–245 (IMS Lecture Notes - Monograph Series, Beechwood, Ohio, 2003).
Shaffer, L.G. & Lupski, J.R. Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu. Rev. Genet. 34, 297–329 (2000).
Cohen, D., Chumakov, I. & Weissenbach, J. A first-generation physical map of the human genome. Nature 366, 698–701 (1993).
Brownstein, B.H. et al. Isolation of single-copy human genes from a library of yeast artificial chromosome clones. Science 244, 1348–1351 (1989).
Monaco, A.P. et al. Isolation of the human sex determining region from a Y-enriched yeast artificial chromosome library. Genomics 11, 1049–1053 (1991).
Acknowledgements
This work was funded by the Wellcome Trust. We thank M. Jobling for the kind gift of DNA, M. Ross for insights into Xq28, J. McCafferty and W. Howat for help with imaging, J. Collins and A. Coffey for guidance on pulsed-field gel elecrophoresis, and P. Holliger and Z. Oliynyk for advice on preparation of emulsions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Blinded genotyping of ten males of known int22h inversion status. (PDF 158 kb)
Supplementary Fig. 2
Pulsed Field Gel Electrophoresis of genomic DNA before and after emulsion preparation. (PDF 227 kb)
Rights and permissions
About this article
Cite this article
Turner, D., Shendure, J., Porreca, G. et al. Assaying chromosomal inversions by single-molecule haplotyping. Nat Methods 3, 439–445 (2006). https://doi.org/10.1038/nmeth881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth881
This article is cited by
-
Copy number variation arising from gene conversion on the human Y chromosome
Human Genetics (2018)
-
High-throughput haplotype determination over long distances by haplotype fusion PCR and ligation haplotyping
Nature Protocols (2009)
-
The population genetics of structural variation
Nature Genetics (2007)
-
Revealing the hidden structure of our genome
Nature Methods (2006)