Abstract
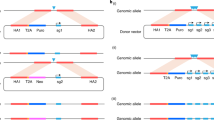
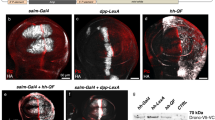
The conditional expression of hairpin constructs in Drosophila melanogaster has emerged in recent years as a method of choice in functional genomic studies. To date, upstream activating site–driven RNA interference constructs have been inserted into the genome randomly using P-element–mediated transformation, which can result in false negatives due to variable expression. To avoid this problem, we have developed a transgenic RNA interference vector based on the phiC31 site-specific integration method.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Dietzl, G. et al. Nature 448, 151–156 (2007).
Brand, A.H. & Perrimon, N. Development 118, 401–415 (1993).
Groth, A.C., Fish, M., Nusse, R. & Calos, M.P. Genetics 166, 1775–1782 (2004).
Perrimon, N. & Mathey-Prevot, B. Genetics 175, 7–16 (2007).
Bischof, J., Maeda, R.K., Hediger, M., Karch, F. & Basler, K. Proc. Natl. Acad. Sci. USA 104, 3312–3317 (2007).
Presente, A., Shaw, S., Nye, J.S. & Andres, A.J. Genesis 34, 165–169 (2002).
Gustafson, K. & Boulianne, G.L. Genome 39, 174–182 (1996).
Mondal, K. et al. J. Mol. Biol. 370, 939–950 (2007).
Fridell, Y.W. & Searles, L.L. Nucleic Acids Res. 19, 5082 (1991).
Siegal, M.L. & Hartl, D.L. Genetics 144, 715–726 (1996).
An, X., Armstrong, J.D., Kaiser, K. & O'Dell, K.M. J. Neurogenet. 14, 227–243 (2000).
Acknowledgements
We thank G. Rubin, C. Zuker, K. Moses and B. Mathey-Prevot for critical input on the project, B. Dickson (IMP, Vienna) for the gift of UAS-Dcr2 and F. Karch (University of Geneva) for nanos-integrase. M.M. is a fellow of the Jane Coffin Childs Memorial Fund. M.B. is supported by R01 GM067761 from the US National Institute of General Medical Sciences. This work was supported in part by the Janelia Farm Visitor program. N.P. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Table 1, Supplementary Methods, Supplementary Figure 1 and Supplementary Note (PDF 1804 kb)
Rights and permissions
About this article
Cite this article
Ni, JQ., Markstein, M., Binari, R. et al. Vector and parameters for targeted transgenic RNA interference in Drosophila melanogaster. Nat Methods 5, 49–51 (2008). https://doi.org/10.1038/nmeth1146
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1146
This article is cited by
-
TnaA, a trithorax group protein, modulates wingless expression in different regions of the Drosophila wing imaginal disc
Scientific Reports (2023)
-
Sestrin mediates detection of and adaptation to low-leucine diets in Drosophila
Nature (2022)
-
A LexAop > UAS > QUAS trimeric plasmid to generate inducible and interconvertible Drosophila overexpression transgenes
Scientific Reports (2022)
-
Heritable shifts in redox metabolites during mitochondrial quiescence reprogramme progeny metabolism
Nature Metabolism (2021)
-
The sugar-responsive enteroendocrine neuropeptide F regulates lipid metabolism through glucagon-like and insulin-like hormones in Drosophila melanogaster
Nature Communications (2021)