Abstract
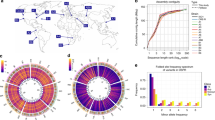
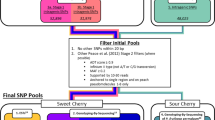
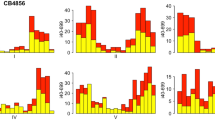
Single nucleotide polymorphisms (SNPs) are useful markers for genetic mapping experiments in model organisms. Here we report the establishment of a high-density SNP map and high-throughput genotyping assays for Drosophila melanogaster. Our map comprises 27,367 SNPs in common laboratory Drosophila stocks. These SNPs were clustered within 2,238 amplifiable markers at an average density of 1 marker every 50.3 kb, or 6.3 genes. We have also constructed a set of 62 Drosophila stocks, each of which facilitates the generation of recombinants within a defined genetic interval of 1–2 Mb. For flexible, high-throughput SNP genotyping, we used fluorescent tag-array mini-sequencing (TAMS) assays. We designed and validated TAMS assays for 293 SNPs at an average resolution of 391.3 kb, and demonstrated the utility of these tools by rapidly mapping 14 mutations that disrupt embryonic muscle patterning. These resources enable high-resolution high-throughput genetic mapping in Drosophila.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Davis, M.W. et al. Rapid single nucleotide polymorphism mapping in C. elegans. BMC Genomics 6, 118 (2005).
Davis, M.W. & Hammarlund, M. Single-nucleotide polymorphism mapping. Methods Mol. Biol. 351, 75–92 (2006).
Zipperlen, P. et al. A universal method for automated gene mapping. Genome Biol. 6, R19 (2005).
Wicks, S.R., Yeh, R.T., Gish, W.R., Waterston, R.H. & Plasterk, R.H. Rapid gene mapping in Caenorhabditis elegans using a high density polymorphism map. Nat. Genet. 28, 160–164 (2001).
Teeter, K. et al. Haplotype dimorphism in a SNP collection from Drosophila melanogaster. J. Exp. Zool. 288, 63–75 (2000).
Berger, J. et al. Genetic mapping with SNP markers in Drosophila. Nat. Genet. 29, 475–481 (2001).
Hoskins, R.A. et al. Single nucleotide polymorphism markers for genetic mapping in Drosophila melanogaster. Genome Res. 11, 1100–1113 (2001).
Martin, S.G., Dobi, K.C. & St. Johnston, D. A rapid method to map mutations in Drosophila. Genome Biol. 2, Research0036 (2001).
Stickney, H.L. et al. Rapid mapping of zebrafish mutations with SNPs and oligonucleotide microarrays. Genome Res. 12, 1929–1934 (2002).
Cho, R.J. et al. Genome-wide mapping with biallelic markers in Arabidopsis thaliana. Nat. Genet. 23, 203–207 (1999).
Torjek, O. et al. Establishment of a high-efficiency SNP-based framework marker set for Arabidopsis. Plant J. 36, 122–140 (2003).
Nairz, K., Stocker, H., Schindelholz, B. & Hafen, E. High-resolution SNP mapping by denaturing HPLC. Proc. Natl. Acad. Sci. USA 99, 10575–10580 (2002).
Underhill, P.A. et al. Detection of numerous Y chromosome biallelic polymorphisms by denaturing high-performance liquid chromatography. Genome Res. 7, 996–1005 (1997).
Orita, M., Iwahana, H., Kanazawa, H., Hayashi, K. & Sekiya, T. Detection of polymorphisms of human DNA by gel electrophoresis as single-strand conformation polymorphisms. Proc. Natl. Acad. Sci. USA 86, 2766–2770 (1989).
Macdonald, S.J., Pastinen, T., Genissel, A., Cornforth, T.W. & Long, A.D. A low-cost open-source SNP genotyping platform for association mapping applications. Genome Biol. 6, R105 (2005).
Xu, T. & Rubin, G.M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993).
Rorth, P. et al. Systematic gain-of-function genetics in Drosophila. Development 125, 1049–1057 (1998).
Lindroos, K., Sigurdsson, S., Johansson, K., Ronnblom, L. & Syvänen, A.C. Multiplex SNP genotyping in pooled DNA samples by a four-colour microarray system. Nucleic Acids Res. 30, e70 (2002).
Lovmar, L., Fredriksson, M., Liljedahl, U., Sigurdsson, S. & Syvänen, A.C. Quantitative evaluation by minisequencing and microarrays reveals accurate multiplexed SNP genotyping of whole genome amplified DNA. Nucleic Acids Res. 31, e129 (2003).
Bradley, P.S., Mangasarian, O.L. & Street, W.N. Clustering via concave minimization. in Advances in Neural Information Processing Systems (eds., Mozer, M.C., Jordan, M.I. & Petsche, T.) 368–374 (MIT Press, Cambridge, Massachusetts, 1997).
Jain, A.K. & Dubes, R.C. Algorithms for Clustering Data. (Prentice-Hall, Englewood Cliffs, New Jersey, 1988).
Schnorrer, F., Kalchhauser, I. & Dickson, B.J. The transmembrane protein kon-tiki couples to Dgrip to mediate myotube targeting in Drosophila. Dev. Cell 12, 751–766 (2007).
Ryder, E. et al. The DROSDEL collection: A set of P-element insertions for generating custom chromosomal aberrations in Drosophila melanogaster. Genetics 167, 797–813 (2004).
Parks, A.L. et al. Systematic generation of high-resolution deletion coverage of the Drosophila melanogaster genome. Nat. Genet. 36, 288–292 (2004).
Steemers, F.J. et al. Whole-genome genotyping with the single-base extension assay. Nat. Methods 3, 31–33 (2006).
Kurg, A. et al. Arrayed primer extension: solid-phase four-color DNA resequencing and mutation detection technology. Genet. Test. 4, 1–7 (2000).
Banér, J. et al. Parallel gene analysis with allele-specific padlock probes and tag microarrays. Nucleic Acids Res. 31, e103 (2003).
Fan, J.B. et al. Highly parallel SNP genotyping. Cold Spring Harb. Symp. Quant. Biol. 68, 69–78 (2003).
Hardenbol, P. et al. Multiplexed genotyping with sequence-tagged molecular inversion probes. Nat. Biotechnol. 21, 673–678 (2003).
Chen, E.H. & Olson, E.N. Antisocial, an intracellular adaptor protein, is required for myoblast fusion in Drosophila. Dev. Cell 1, 705–715 (2001).
Acknowledgements
We thank J. Berger and T. Suzuki for initial help in the creation of the SNP map, R. Figueroa for printing the tag arrays, S. Krüttner for help mapping the muscle mutants, J. Saharinen for the SNPSnapper v4.0 software (National Public Health Institute, Finland), and J. Braisted and N. Bhagabati for support in using the source code of the Multiple Experiment Viewer. This work was supported by funds from the EU Fifth Framework Programme, the Austrian Science Fund, and the Swedish Research Council for Science and Technology. Work at the IMP is supported by Boehringer Ingelheim GmbH. F.S. was supported by a long term fellowship from the Human Frontier Science Program.
Author information
Authors and Affiliations
Contributions
D.C. and M.F. generated the SNP map; D.C. developed SNPmapper; A.A., D.C. and F.S. established the TAMS assays; E.V. and Is.K. generated and M.F., E.V. and D.C. verified the 2EP stocks; F.S. generated the muscle mutants; F.S., Ir.K., A.A. and D.C. mapped the muscle mutants; B.J.D. and A.-C.S. coordinated the project; D.C., A.A., F.S., A.-C.S. and B.J.D. wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Tables 1–4, Supplementary Methods (PDF 812 kb)
Supplementary Table 5
Pre-computed 2EP TAMS assays. (XLS 562 kb)
Rights and permissions
About this article
Cite this article
Chen, D., Ahlford, A., Schnorrer, F. et al. High-resolution, high-throughput SNP mapping in Drosophila melanogaster. Nat Methods 5, 323–329 (2008). https://doi.org/10.1038/nmeth.1191
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1191
This article is cited by
-
Strategies for gene disruption in Drosophila
Cell & Bioscience (2014)
-
SNPing and chipping away at the fly genome
Nature Methods (2008)
-
Positional cloning by fast-track SNP-mapping in Drosophila melanogaster
Nature Protocols (2008)