Abstract
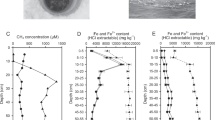
Aquatic habitats beneath ice masses contain active microbial ecosystems capable of cycling important greenhouse gases, such as methane (CH4). A large methane reservoir is thought to exist beneath the West Antarctic Ice Sheet, but its quantity, source and ultimate fate are poorly understood. For instance, O2 supplied by basal melting should result in conditions favourable for aerobic methane oxidation. Here we use measurements of methane concentrations and stable isotope compositions along with genomic analyses to assess the sources and cycling of methane in Subglacial Lake Whillans (SLW) in West Antarctica. We show that sub-ice-sheet methane is produced through the biological reduction of CO2 using H2. This methane pool is subsequently consumed by aerobic, bacterial methane oxidation at the SLW sediment–water interface. Bacterial oxidation consumes >99% of the methane and represents a significant methane sink, and source of biomass carbon and metabolic energy to the surficial SLW sediments. We conclude that aerobic methanotrophy may mitigate the release of methane to the atmosphere upon subglacial water drainage to ice sheet margins and during periods of deglaciation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kirschke, S. et al. Three decades of global methane sources and sinks. Nat. Geosci. 6, 813–823 (2013).
Thauer, R. K., Kaster, A-K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–591 (2008).
Conrad, R. The global methane cycle: recent advances in understanding the microbial processes involved. Environ. Microbiol. Rep. 1, 285–292 (2009).
Wadham, J. L. et al. Potential methane reservoirs beneath Antarctica. Nature 488, 633–637 (2012).
Dieser, M. et al. Molecular and biogeochemical evidence for methane cycling beneath the western margin of the Greenland Ice Sheet. ISME J. 8, 2305–2316 (2014).
Wadham, J. L. et al. The potential role of the Antarctic Ice Sheet in global biogeochemical cycles. Earth Environ. Sci. Trans. R. Soc. Edinburgh 104, 55–67 (2013).
Christner, B. C. et al. A microbial ecosystem beneath the West Antarctic Ice Sheet. Nature 512, 310–313 (2014).
Skidmore, M. in Antarctic Subglacial Environments, Geophysical Monograph Series (eds Siegert, M. J., Kennicutt II, M. C. & Bindschandler, R. A.) 61–81 (American Geophysical Union, 2011).
Fisher, A. T., Mankoff, K. D., Tulaczyk, S. M. & Tyler, S. W. High geothermal heat flux measured below the West Antarctic Ice Sheet. Sci. Rep. 1, e1500093 (2015).
Whiticar, M. J. Carbon and hydrogen isotope systematics of bacterial formation and oxidation of methane. Chem. Geol. 161, 291–314 (1999).
Coleman, D. D., Liu, C.-L. & Riley, K. M. Microbial methane in the shallow Paleozoic sediments and glacial deposits of Illinois, USA. Chem. Geol. 71, 23–40 (1988).
Conrad, R. in Advances in Agronomy (ed. Sparks, S.) 1–63 (Elsevier, 2007).
Schoell, M. Multiple origins of methane in the Earth. Chem. Geol. 71, 1–10 (1988).
Michaud, A. B. et al. Solute sources and geochemical processes in Subglacial Lake Whillans, West Antarctica. Geology 44, 347–350 (2016).
Telling, J. et al. Rock comminution as a source of hydrogen for subglacial ecosystems. Nat. Geosci. 8, 851–855 (2015).
Lin, L.-H., Slater, G. F., Sherwood Lollar, B., Lacrampe-Couloume, G. & Onstott, T. C. The yield and isotopic composition of radiolytic H2, a potential energy source for the deep subsurface biosphere. Geochim. Cosmochim. Acta 69, 893–903 (2005).
Matheus Carnevali, P. B. et al. Methane sources in arctic thermokarst lake sediments on the North Slope of Alaska. Geobiology 13, 181–197 (2015).
Lever, M. A. et al. Evidence for microbial carbon and sulfur cycling in deeply buried ridge flank basalt. Science 339, 1305–1308 (2013).
Blazewicz, S. J., Barnard, R. L., Daly, R. A. & Firestone, M. K. Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J. 7, 2061–2068 (2013).
Jones, S. E. & Lennon, J. T. Dormancy contributes to the maintenance of microbial diversity. Proc. Natl Acad. Sci. USA 107, 5881–5886 (2010).
Achberger, A. M. et al. Microbial community structure of Subglacial Lake Whillans, West Antarctica. Front. Microbiol. 7, 1457 (2016).
Grant, N. J. & Whiticar, M. J. Stable carbon isotopic evidence for methane oxidation in plumes above Hydrate Ridge, Cascadia Oregon Margin. Global Biogeochem. Cycles 16, 1124 (2002).
Coleman, D. D., Risatti, J. B. & Schoell, M. Fractionation of carbon and hydrogen isotopes by methane-oxidizing bacteria. Geochim. Cosmochim. Acta 45, 1033–1037 (1981).
McDonald, I. R., Bodrossy, L., Chen, Y. & Murrell, J. C. Molecular ecology techniques for the study of aerobic methanotrophs. Appl. Environ. Microbiol. 74, 1305–1315 (2008).
Knief, C. Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front. Microbiol. 6, 1346 (2015).
Ho, A. et al. Conceptualizing functional traits and ecological characteristics of methane-oxidizing bacteria as life strategies. Environ. Microbiol. Rep. 5, 335–345 (2013).
Martineau, C., Whyte, L. G. & Greer, C. W. Stable isotope probing analysis of the diversity and activity of methanotrophic bacteria in soils from the Canadian high Arctic. Appl. Environ. Microbiol. 76, 5773–5784 (2010).
Graef, C., Hestnes, A. G., Svenning, M. M. & Frenzel, P. The active methanotrophic community in a wetland from the High Arctic. Environ. Microbiol. Rep. 3, 466–472 (2011).
He, R. et al. Shifts in identity and activity of methanotrophs in Arctic lake sediments in response to temperature changes. Appl. Environ. Microbiol. 78, 4715–4723 (2012).
Shock, E. L. et al. Quantifying inorganic sources of geochemical energy in hydrothermal ecosystems, Yellowstone National Park, USA. Geochim. Cosmochim. Acta 74, 4005–4043 (2010).
Amend, J. P. & Shock, E. L. Energetics of overall metabolic reactions of thermophilic and hyperthermophilic Archaea and Bacteria. FEMS Microbiol. Rev. 25, 175–243 (2001).
Vick-Majors, T. J. et al. Physiological ecology of microorganisms in Subglacial Lake Whillans. Front. Microbiol. 7, 1705 (2016).
Trimmer, M. et al. Riverbed methanotrophy sustained by high carbon conversion efficiency. ISME J. 9, 2304–2314 (2015).
Iversen, N. & Blackburn, T. H. Seasonal rates of methane oxidation in anoxic marine sediments. Appl. Environ. Microbiol. 41, 1295–1300 (1981).
Marlow, J. J. et al. Carbonate-hosted methanotrophy represents an unrecognized methane sink in the deep sea. Nat. Commun. 5, 5094 (2014).
Bastviken, D., Ejlertsson, J., Sundh, I. & Tranvik, L. Methane as a source of carbon and energy for lake pelagic food webs. Ecology 84, 969–981 (2003).
Lough, A. C. et al. Seismic detection of an active subglacial magmatic complex in Marie Byrd Land, Antarctica. Nat. Geosci. 6, 1031–1035 (2013).
Beem, L. H., Jezek, K. C. & Van Der Veen, C. J. Basal melt rates beneath Whillans Ice Stream, West Antarctica. J. Glaciol. 56, 647–654 (2010).
Yvon-Durocher, G. et al. Methane fluxes show consistent temperature dependence across microbial to ecosystem scales. Nature 507, 488–491 (2014).
Priscu, J. C. et al. A microbiologically clean strategy for access to the Whillans Ice Stream subglacial environment. Antarct. Sci. 11, 1–11 (2013).
Tulaczyk, S. et al. WISSARD at Subglacial Lake Whillans, West Antarctica: scientific operations and initial observations. Ann. Glaciol. 55, 51–58 (2014).
National Resource Council Exploration of Antarctic Subglacial Aquatic Environments (The National Academies Press, 2007).
Riedinger, N. et al. Methane at the sediment–water transition in Black Sea sediments. Chem. Geol. 274, 29–37 (2010).
Wiesenburg, D. A. & Guinasso, N. L. Jr Equilibrium solubilities of methane, carbon monoxide, and hydrogen in water and sea water. J. Chem. Eng. Data 24, 356–360 (1979).
Avnimelech, Y., Ritvo, G., Meijer, L. E. & Kochba, M. Water content, organic carbon and dry bulk density in flooded sediments. Aquacult. Eng. 25, 25–33 (2001).
Yarnes, C. 13C and 2H measurement of methane from ecological and geological sources by gas chromatography/combustion/pyrolysis isotope-ratio mass spectrometry. Rapid Commun. Mass Spectrom. 27, 1036–1044 (2013).
Vick-Majors, T. J. Biogeochemical Processes in Antarctic Aquatic Environments: Linkages and Limitations (Montana State University, 2016).
Solorzano, L. Determination of ammonia in natural waters by the phenolhypochlorite method. Limnol. Oceanogr. 14, 799–801 (1969).
Lever, M. A. et al. A modular method for the extraction of DNA and RNA, and the separation of DNA pools from diverse environmental sample types. Front. Microbiol. 6, 476 (2015).
Paegel, B. M., Emrich, C. A., Wedemayer, G. J., Scherer, J. R. & Mathies, R. A. High throughput DNA sequencing with a microfabricated 96-lane capillary array electrophoresis bioprocessor. Proc. Natl Acad. Sci. USA 99, 574–579 (2002).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Stumm, W. & Morgan, J. J. Aquatic Chemistry: Chemical Equilibria and Rates in Natural Waters (Wiley-Interscience, 1996).
Parkhurst, D. L. & Appelo, C. A. J. US Geological Survey Techniques and Methods 497 (US Geological Survey, 2013).
Boetius, A. & Wenzhöfer, F. Seafloor oxygen consumption fuelled by methane from cold seeps. Nat. Geosci. 6, 725–734 (2013).
LaRowe, D. E. & Amend, J. P. in Microbial Life of the Deep Biosphere (eds Kallmeyer, J. & Wagner, D.) 279–302 (Walter de Gruyter, 2014).
Osburn, M. R. et al. Chemolithotrophy in the continental deep subsurface: Sanford Underground Research Facility (SURF), USA. Front. Microbiol. 5, 610 (2014).
Shen, L. & Chen, Z. Critical review of the impact of tortuosity on diffusion. Chem. Eng. Sci. 62, 3748–3755 (2007).
Broecker, W. S. & Peng, T.-H. Gas exchange rates between air and seal. Tellus 26, 21–35 (1974).
Shelley, F., Abdullahi, F., Grey, J. & Trimmer, M. Microbial methane cycling in the bed of a chalk river: oxidation has the potential to match methanogenesis enhanced by warming. Freshw. Biol. 60, 150–160 (2015).
Whalen, S. C., Reeburgh, W. S. & Sandbeck, K. A. Rapid methane oxidation in a landfill cover soil. Appl. Environ. Microbiol. 56, 3405–3411 (1990).
Urmann, K., Lazzaro, A., Gandolfi, I., Schroth, M. H. & Zeyer, J. Response of methanotrophic activity and community structure to temperature changes in a diffusive CH/O counter gradient in an unsaturated porous medium. FEMS Microbiol. Ecol. 69, 202–212 (2009).
Siegfried, M. R., Fricker, H. A., Carter, S. P. & Tulaczyk, S. Episodic ice velocity fluctuations triggered by a subglacial flood in West Antarctica. Geophys. Res. Lett. 43, 2640–2648 (2016).
Acknowledgements
This study was funded by National Science Foundation – Division of Polar Programs grants (0838933, 1346250, 1439774 to J.C.P.; 0838941 to B.C.C.) awarded as part of the Whillans Ice Stream Subglacial Access Research Drilling (WISSARD) project. We thank the WISSARD Science Team (see http://wissard.org for the full list of team members) for their assistance in expedition planning and with collecting and processing samples. Partial support was provided by graduate fellowships from the NSF-IGERT Program (0654336), Montana Space Grant Consortium and NSF-Center for Dark Energy Biosphere Investigations (A.B.M.); a dissertation grant from the American Association of University Women (T.J.V.-M.); a NSF-Graduate Research Fellowship (A.M.A.); and a Sêr Cymru National Research Network for Low Carbon, Energy and the Environment Grant from the Welsh Government and Higher Education Funding Council for Wales (A.C.M.). We thank R. Scherer and R. Powell for sediment cores. B.B. Jørgensen, M. A. Lever and S. Nielsen provided support and assistance with DNA extraction and pmoA/mcrA amplification. Logistics were conducted by the 139th Expeditionary Airlift Squadron of the New York Air National Guard, Kenn Borek Air, and Antarctic Support Contractor, managed by Lockheed-Martin. Hot-water drill support was provided by University of Nebraska-Lincoln and directed by F. Rack and D. Duling (chief driller). D. Blythe, J. Burnett, C. Carpenter, D. Gibson, J. Lemery, A. Melby and G. Roberts provided drill support at SLW. This is C-DEBI contribution #371.
Author information
Authors and Affiliations
Contributions
A.B.M., J.E.D., T.J.V.-M., J.C.P. and M.L.S. wrote the manuscript. A.B.M., J.E.D., M.L.S. and T.J.V.-M. conducted and analysed methane concentration and isotopic data. A.M.A., A.B.M. and B.C.C. processed, analysed and interpreted the molecular data. A.C.M. conducted thermodynamic calculations. All authors contributed to the study design, collection of samples and approved the final draft of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 545 kb)
Rights and permissions
About this article
Cite this article
Michaud, A., Dore, J., Achberger, A. et al. Microbial oxidation as a methane sink beneath the West Antarctic Ice Sheet. Nature Geosci 10, 582–586 (2017). https://doi.org/10.1038/ngeo2992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ngeo2992
This article is cited by
-
Electrochemically coupled CH4 and CO2 consumption driven by microbial processes
Nature Communications (2024)
-
CH4 emissions from runoff water of Alaskan mountain glaciers
Scientific Reports (2024)
-
Groundwater springs formed during glacial retreat are a large source of methane in the high Arctic
Nature Geoscience (2023)
-
Methylotrophic Communities Associated with a Greenland Ice Sheet Methane Release Hotspot
Microbial Ecology (2023)
-
Biogeochemical and historical drivers of microbial community composition and structure in sediments from Mercer Subglacial Lake, West Antarctica
ISME Communications (2023)