Abstract
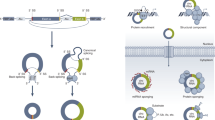
In eukaryotes, RNA processing events, including alternative splicing and RNA editing, can generate many different messages from a single gene. As a consequence, the RNA pool, which we refer to here as the 'ribotype', has a different information content from the genotype and can vary as circumstances change. The outcome of a single RNA processing event often regulates the outcome of another, giving rise to networks that affect the composition and expression of a particular ribotype. Successful ribotypes are determined by natural selection, and can be incorporated into the genome over time by reverse transcription. Eukaryotic evolution is therefore influenced by the alternate ways in which RNAs are processed and the continual interplay between RNA and DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Crick, F. Central dogma of molecular biology. Nature 227, 561–563 (1970).
Wang, Y.-C., Selvakumar, M. & Helfman, D. Alternative pre-mRNA splicing. in Eukaryotic mRNA Processing (ed. Krainer, A.R.) 242–278 (Oxford University Press, New York, 1997).
Bass, B.L. RNA editing and hypermutation by adenosine deamination. Trends Biochem. Sci. 22, 157–162 (1997).
Poole, A.M., Jeffares, D.C. & Penny, D. The path from the RNA world. J. Mol. Evol. 46, 1–17 (1998).
Ricchetti, M. & Buc, H. E. coli DNA polymerase I as a reverse transcriptase. EMBO J. 12, 387–396 (1993).
Jones, M.D. & Foulkes, N.S. Reverse transcription of mRNA by Thermus aquaticus DNA polymerase. Nucleic Acids Res. 17, 8387–8388 (1989).
Allaudeen, H.S. DNA and RNA polymerases of mammalian cells and tumor viruses. Pharmacol. Ther. 2, 447–448 (1978).
Baker, B.S. Sex in flies: the splice of life. Nature 340, 521–524 (1989).
Keyes, L.N., Cline, T.W. & Schedl, P. The primary sex determination signal of Drosophila acts at the level of transcription. Cell 68, 933–943 (1992).
Kren, B.T. & Steer, C.J. Posttranscriptional regulation of gene expression in liver regeneration: role of mRNA stability. FASEB J. 10, 559–573 (1996).
Ross, J. mRNA stability in mammalian cells. Microbiol. Rev. 59, 423–450 (1995).
Lee, R., Feinbaum, R. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 643–654 (1993).
Ha, I., Wrightman, B. & Ruvkun, G. Post-transcriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993).
Ha, I., Wrightman, B. & Ruvkun, G. A bulged lin-4/lin-14 RNA duplex is sufficient for Caenorhabditis elegans lin-14 temporal gradient formation. Genes Dev. 10, 3041–3050 (1996).
Dubnau, J. & Struhl, G. RNA recognition and translational regulation by a homeodomain protein. Nature 379, 694–699 (1996).
Rivera-Pomar, R., Niessing, D., Schmidt-Ott, U., Gehring, W.J. & Jackle, H. RNA binding and translational repression by bicoid. Nature 379, 746–749 (1996).
Bashaw, G.J. & Baker, B.S. The regulation of the Drosophila msl-2 gene reveals a function for sex-lethal in translational control. Cell 89, 789–798 (1997).
Gebauer, F., Merendino, L., Hentze, M.W. & Valcarcel, J. The Drosophila splicing regulator sex-lethal directly inhibits translation of male-specific-lethal 2 mRNA. RNA 4, 142–150 (1998).
Simpson, L. & Thiemann, O.H. mRNA editing. in Eukaryotic mRNA Processing (ed. Krainer, A.R.) 335–376 (Oxford University Press, New York, 1997).
Tollervey, D. & Kiss, T. Function and synthesis of small nucleolar RNAs. Curr. Opin. Cell Biol. 9, 337–342 (1997).
Bachellerie, J.-P. et al. Antisense snoRNAs: a family of nucleolar RNAs with long complementarities to rRNA. Trends Biochem. Sci. 20, 261–264 (1995).
Eichler, D.C. & Craig, N. Processing of eukaryotic ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol. 49, 197–239 (1994).
Sommer, B., Kohler, M., Sprengel, R. & Seeburg, P.H. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell 67, 11–19 (1991).
Hume, R.I., Dingledine, R. & Heinemann, S.F. Identification of a site in the glutamate receptor subunits that controls calcium permeability. Science 253, 1028–1031 (1991).
Verdoorn, T.A., Burnashev, N., Monyer, H., Seeburg, P.H. & Sakmann, B. Structural determinants of ion flow through recombinant glutamate receptor channels. Science 252, 1715–1718 (1991).
Higuchi, M. et al. RNA editing of AMPA receptor subunit GluR-B: a base-paired intron-exon structure determines position and efficiency. Cell 75, 1361–1370 (1993).
Kohler, M., Burnashev, N., Sakmann, B. & Seeburg, P.H. Determinants of Ca2+ permeability in both TM1 and TM2 of high affinity kainate receptor channels: diversity by RNA editing. Neuron 10, 491–500 (1993).
Lomeli, H. et al. Control of kinetic properties of AMPA receptor channels by nuclear RNA editing. Science 266, 1709–1713 (1994).
Herb, A., Higuchi, M., Sprengel, R. & Seeburg, P.H. Q/R site editing in kainate receptor GluR5 and GluR6 pre-mRNAs requires distant intronic sequences. Proc. Natl Acad. Sci. USA 93, 1875–1880 (1996).
Egebjerg, J., Kukekov, V. & Heinemann, S.F. Intron sequence directs RNA editing of the glutamate receptor GluR2 coding sequence. Proc. Natl Acad. Sci. USA 91, 10270–10274 (1994).
Burns, C.N. et al. Regulation of serotonin-2C receptor G-protein coupling by RNA editing. Nature 387, 303–308 (1997).
Ma, J., Qian, R., Rausa, F.M. III & Colley, K.J. Two naturally occurring α2,6-Sialyltransferase forms with a single amino acid change in the catalytic domain differ in their catalytic activity and proteolytic processing. J. Biol. Chem. 272, 672–679 (1997).
Patton, D.E., Silva, T. & Bezanilla, F. RNA editing generates a diverse array of transcripts encoding squid Kv2 K+ channels with altered functional properties. Neuron 19, 711–722 (1997).
Bass, B.L. RNA editing: new uses for old players in the RNA world. in The RNA World (eds Gesteland, R.F. & Atkins, J.F.) 383–418 (Cold Spring Harbor Laboratory Press, Plainview, 1993).
Herbert, A.G. RNA editing, introns and evolution. Trends Genet. 12, 6–9 (1996).
Polson, A.G., Bass, B.L. & Casey, J.L. RNA editing of hepatitis δ virus antigenome by dsRNA-adenosine deaminase. Nature 380, 454–456 (1996).
Petschek, J.P., Scheckelhoff, M.R., Mermer, M.J. & Vaughn, J.C. RNA editing and alternative splicing generate mRNA transcript diversity from the Drosophila 4f-rnp locus. Gene 204, 267–276 (1997).
Lai, M.M.C. The molecular biology of hepatitis δ virus. Annu. Rev. Biochem. 64, 259–286 (1995).
Dougherty, W.G. & Parks, T.D. Transgenes and gene suppression: telling us something new? Curr. Opin. Cell Biol. 7, 399–405 (1995).
Schiebel, W., Haas, B., Marinkovic, S., Klanner, A. & Sanger, H.L. RNA-directed RNA polymerase from tomato leaves. J. Biol. Chem. 263, 11858–11867 (1993).
Tabari, H., Grishok, A. & Mello, C.C. RNAi in C. elegans: soaking in the genome sequence. Science 282, 430–431 (1998).
Smit, A.F.A. The origin of interspersed repeats in the human genome. Curr. Opin. Genet. Dev. 6, 743–748 (1996).
Weiner, A.M., Deninger, P.L. & Efstratiadis, A. Nonviral retroposons: genes, pseudogenes and transposable elements generated by the reverse flow of genetic information. Annu. Rev. Biochem. 55, 631–661 (1986).
Vanin, E.F. Processed psuedogenes: characteristics and evolution. Annu. Rev. Genet. 19, 253–272 (1985).
Moore, J.K. & Haber, J.E. Capture of retrotransposon DNA at the sites of chromosomal double-stranded breaks. Nature 383, 644–646 (1996).
Teng, S.-C., Kim, B. & Gabriel, A. Retrotransposon reverse-transcriptase-mediated repair of chromosomal breaks. Nature 383, 641–644 (1996).
Eickbush, T.H. Telomerase and retrotransposons: which came first? Science 277, 911–912 (1997).
Taghian, D.G. & Nickoloff, J.A. Chromosomal double-strand breaks induce gene conversion at high frequency in mammalian cells. Mol. Cell. Biol. 17, 6386–6393 (1997).
Liang, F., Han, M., Romanienko, P.J. & Jasin, M. Homology-directed repair is a major double-strand break repair pathway in mammalian cells. Proc. Natl Acad. Sci. USA 95, 5172–5177 (1998).
Britten, R.J. Mobile elements inserted in the distant past have taken on important functions. Gene 205, 177–182 (1997).
Murnane, J.P. & Morales, J.F. Use of a mammalian interspersed repetitive (MIR) element in the coding and processing sequences of mammalian genes. Nucleic Acids Res. 23, 2837–2839 (1995).
Wallace, M.R. et al. A de novo Alu insertion results in neurofibromatosis type I. Nature 353, 864–866 (1991).
Miki, Y., Katagiri, M.Y., Kasumui, F., Yoshimoto, T. & Nakamura, Y. Mutation analysis in the BRCA2 gene in primary breast cancers. Nature Genet. 13, 245–247 (1996).
Kidwell, M.G. & Lisch, D. Transposable elements as sources of variation in animals and plants. Proc. Natl Acad. Sci. USA 94, 7704–7711 (1997).
Cramer, P., Pesce, C.P., Baralle, C.G. & Kornblihtt, A.R. Functional association between promoter structure and transcript alternative splicing. Proc. Natl Acad. Sci. USA 94, 11456–11460 (1997).
Branchiforte, D. & Martin, S.L. Developmental and cell type specificity of LINE-1 expression in mouse testis: implications for transposition. Mol. Cell. Biol. 14, 2584–2592 (1994).
Lewin, B. Gene Expression: Eukaryotic Genomes 728–760 (Wiley, New York, 1980).
Acknowledgements
We thank K. Lowenhaupt and A. Varshavsky for a critical reading of the manuscript. This work was supported by grants from the National Institutes of Health and the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Herbert, A., Rich, A. RNA processing and the evolution of eukaryotes. Nat Genet 21, 265–269 (1999). https://doi.org/10.1038/6780
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/6780
This article is cited by
-
The genome of the giant Nomura’s jellyfish sheds light on the early evolution of active predation
BMC Biology (2019)
-
Nature inspired genetic algorithms for hard packing problems
Annals of Operations Research (2010)
-
RNA-mediated epigenetic programming of a genome-rearrangement pathway
Nature (2008)
-
The four Rs of RNA-directed evolution
Nature Genetics (2004)
-
IG20, in contrast to DENN-SV, (MADD splice variants) suppresses tumor cell survival, and enhances their susceptibility to apoptosis and cancer drugs
Oncogene (2004)