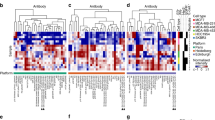
Abstract
Patient-tailored medicine can be defined as the selection of specific therapeutics to treat disease in a particular individual based on genetic, genomic or proteomic information. While individualized treatments have been used in medicine for years, advances in cancer treatment have now generated a need to more precisely define and identify those patients who will derive the most benefit from new-targeted agents. Cellular signaling pathways are a protein-based network, and the intended drug effect is to disrupt aberrant protein phosphorylation-based enzymatic activity and epigenetic phenomena. Pharmacoproteomics, or the tailoring of therapy based on proteomic knowledge, will begin to take a central role in this process. A new type of protein array platform, the reverse-phase protein microarray, shows potential for providing detailed information about the state of the cellular 'circuitry' from small samples such as patient biopsy specimens. Measurements of hundreds of specific phosphorylated proteins that span large classes of important signaling pathways can be obtained at once from only a few thousand cells. Clinical implementation of these new proteomic tools to aid the clinical, medical and surgical oncologist in making decisions about patient care will now require thoughtful communication between practicing clinicians and research scientists.
Key Points
-
An ever-increasing number of molecularly targeted agents for treating cancer are entering clinical use, generating a new need to better define and identify those patients who will most benefit from them
-
Proteomic technologies may eventually hold a dominant position in tailored medicine as we transition from pharmacogenomics to pharmacoproteomics, since gene transcript profiling cannot be used effectively to elucidate or monitor protein–protein interaction networks and signal transduction pathways
-
Protein microarrays that examine post-translational modifications, such as phosphorylation, in a global, high-throughput manner, can be used to profile the working state of cellular signaling pathways in human tissue
-
The reverse-phase protein microarray (RPA) format is uniquely suited to signal transduction profiling of small samples (e.g. biopsy specimens) such that rational selection and monitoring of patients for targeted medicine is now technically possible
-
Clinical implementation of RPAs for profiling patient tissue specimens before, during and after therapy could provide important diagnostic and prognostic information, and could help therapeutic decision making and monitoring of response or resistance
-
Establishment of RPAs or any stratification tool as routine clinical practice will require standardization of tissue collection and processing, and the cooperation and collaboration of surgeons, oncologists, pathologists and scientists
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jain KK (2002) Personalized medicine. Curr Opin Mol Ther 4: 548–558
Baak JPA et al. (2003) Genomics and proteomics in cancer. Eur J Cancer 39: 1199–1215
Segal E et al. (2005) From signatures to models: understanding cancer using microarrays. Nat Genet 37 (Suppl) S38–S45
Brennan DJ et al. (2005) Application of DNA microarray technology in determining breast cancer prognosis and therapeutic response. Expert Opin Biol Ther 5: 1069–1083
Nishizuka S et al. (2003) Proteomic profiling of the NCI-60 cancer cell lines using new high-density reverse-phase lysate microarrays. Proc Natl Acad Sci USA 100: 14229–14234
Celis JE and Gromov P (2003) Proteomics in translational cancer research: toward an integrated approach. Cancer Cell 3: 9–15
Hunter T (2000) Signaling—2000 and beyond. Cell 100: 113–127
Gorg A et al. (2004) Current two-dimensional electrophoresis technology for proteomics. Proteomics 4: 3665–3685
Gygi SP et al. (1999) Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol 17: 994–999
Krutchinsky AN et al. (2001) Automatic identification of proteins with a MALDI-quadrupole ion trap mass spectrometer. Anal Chem 73: 5066–5077
Washburn MP et al. (2001) Large scale analysis of the yeast proteome by multidimensional protein identification technology. Nat Biotechnol 19: 242–247
Zhou G et al. (2002) 2D differential in-gel electrophoresis for the identification of esophageal scans cell cancer-specific protein markers. Mol Cell Proteomics 1: 117–124
Eckel-Passow JE et al. (2005) Experimental design and analysis of antibody microarrays: applying methods from cDNA arrays. Cancer Res 65: 2985–2989
Haab BB (2005) Antibody arrays in cancer research. Mol Cell Proteomics 4: 377–383
Leuking A et al. (2005) Protein biochips: a new and versatile platform technology for molecular medicine. Drug Disc Today 10: 789–794
Liotta LA et al. (2001) Clinical proteomics: personalized molecular medicine. JAMA 286: 2211–2214
Petricoin EF et al. (2002) Clinical proteomics: translating benchside promise into bedside reality. Nat Rev Drug Discov 1: 683–695
Paweletz CP et al. (2001) Reverse phase protein microarrays which capture disease progression show activation of pro-survival pathways at the cancer invasion front. Oncogene 20: 1981–1989
Zhu H and Snyder M (2003) Protein chip technology. Curr Opin Chem Biol 7: 55–63
Cutler P (2003) Protein arrays: the current state-of-the-art. Proteomics 3: 3–18
Ge H (2000) UPA, a universal protein array system for quantitative detection of protein–protein, protein–DNA, protein–RNA and protein–ligand interactions. Nucleic Acids Res 28: e3
Liotta L and Petricoin E (2000) Molecular profiling of human cancer. Nat Rev Genet 1: 48–56
MacBeath G (2002) Protein microarrays and proteomics. Nat Genet 32 (Suppl) 526–532
Miller JC et al. (2003) Antibody microarray profiling of human prostate cancer sera: antibody screening and identification of potential biomarkers. Proteomics 3: 56–63
Liotta LA et al. (2003) Protein microarrays: meeting analytical challenges for clinical applications. Cancer Cell 3: 317–325
Angenendt P (2005) Progress in protein and antibody microarray technology. Drug Disc Today 10: 503–511
LaBaer J and Ramachandran N (2005) Protein microarrays as tools for functional proteomics. Curr Opin Chem Biol 9: 14–19
Humphery-Smith I et al. (2002) Protein arrays for assessment of target selectivity. Drug Discov World 4: 17–27
MacBeath G and Schreiber SL (2000) Printing proteins as microarrays for high-throughput function determination. Science 289: 1760–1763
Petach H and Gold L (2002) Dimensionality is the issue: use of photoaptamers in protein microarrays. Curr Opin Biotechnol 13: 309–314
Weng S et al. (2002) Generating addressable protein microarrays with PROfusion covalent mRNA-protein fusion technology. Proteomics 2: 48–57
Hultschig C et al. (2006) Recent advances of protein microarrays. Curr Opin Chem Biol 10: 4–10
Robinson WH (2006) Antigen arrays for antibody profiling. Curr Opin Chem Biol 10: 67–72
Sreekumar A et al. (2001) Profiling of cancer cells using protein microarrays: discovery of novel radiation-regulated proteins. Cancer Res 61: 7585–7593
Sanchez-Carbayo M et al. (2006) Profiling bladder cancer using targeted antibody arrays. Am J Pathol 168: 93–103
Grubb RL et al. (2003) Signal pathway profiling of prostate cancer using reverse phase protein microarrays. Proteomics 3: 2142–2146
Petricoin III EF et al. (2005) Mapping molecular networks using proteomics: a vision for patient-tailored combination therapy. J Clin Oncol 23: 3614–3621
Wulfkuhle JD et al. (2003) Signal pathway profiling of ovarian cancer from human tissue specimens using reverse-phase protein microarrays. Proteomics 3: 2085–2090
Gulmann C et al. (2005) Proteomic analysis of apoptotic pathways reveals prognostic factors in follicular lymphoma. Clin Cancer Res 11: 5847–5855
Sheehan KM et al. (2005) Use of reverse-phase protein microarrays and reference standard development for molecular network analysis of metastatic ovarian carcinoma. Mol Cell Proteomics 4: 346–355
Emmert-Buck MR et al. (1996) Laser capture microdissection. Science 274: 998–1001
Zha H et al. (2004) Similarities of prosurvival signals in Bcl-2-positive and Bcl-2-negative follicular lymhomas identified by reverse phase protein microarray. Lab Invest 84: 235–244
Giltrane JM and Rimm DL (2005) Technology insight: identification of biomarkers with tissue microarray technology. Nat Clin Pract Oncol 1: 104–111
Espina V et al. (2003) Protein microarrays: molecular profiling technologies for clinical specimens. Proteomics 3: 2091–2100
Wulfkuhle J et al. (2004) Genomic and proteomic technologies for individualisation and improvement of cancer treatment. Eur J Cancer 40: 2623–2632
Espina V et al. (2004) Use of proteomic analysis to monitor responses to biological therapies. Expert Opin Biol Ther 4: 83–93
Posadas EM et al. (2004) Proteomics and ovarian cancer: implications for diagnosis and treatment: a critical review of the recent literature. Curr Opin Oncol 16: 478–484
Krause DS and Van Etten RA (2005) Tyrosine kinases as targets for cancer therapy. N Engl J Med 353: 172–187
Awada A et al. (2004) New anticancer agents and therapeutic strategies in development for solid cancers: a clinical perspective. Expert Rev Anticancer Ther 4: 53–60
Harari PM and Huang S (2004) Combining EGFR inhibitors with radiation or chemotherapy: will preclinical studies predict clinical results? Int J Radiat Oncol Biol Phys 58: 976–983
Raben D et al. (2002) ZD1839, a selective epidermal growth factor receptor tyrosine kinase inhibitor, alone and in combination with radiation and chemotherapy as a new therapeutic strategy in non-small cell lung cancer. Semin Oncol 29: 37–46
Araujo RP et al. (2005) A mathematical model of combination therapy using the EGFR signaling network. Biosystems 80: 57–69
Araujo RP et al. (2004) Network-targeted combination therapy: a new concept in cancer treatment. Drug Disc Today 1: 425–433
Arteaga CL and Baselga J (2003) Clinical trial design and end points for epidermal growth factor receptor-targeted therapies: implications for drug development and practice. Clin Cancer Res 9: 1579–1589
Gasparini G and Gion M (2000) Molecular-targeted anticancer therapy: challenges related to study design and choice of proper endpoints. Cancer J Sci Am 6: 117–131
Jain KK (2005) Personalised medicine for cancer: from drug development into clinical practice. Expert Opin Pharmacother 6: 1463–1476
von Mehren M (2003) Gastrointestinal stromal tumors: a paradigm for molecularly targeted therapy. Cancer Invest 21: 553–563
Wiestner A and Staudt LM (2003) Towards molecular diagnosis and targeted therapy of lymphoid malignancies. Semin Hematol 40: 296–307
Hynes NE and Lane HA (2005) ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer 5: 341–354
Emens LA (2005) Trastuzumab: targeted therapy for the management of HER-2/neu-overexpressing metastatic breast cancer. Am J Therapeutics 12: 243–253
Dowsett M (2003) Correlation between immunohistochemistry (HercepTest) and fluorescence in situ hybridization (FISH) for HER2 in 426 breast carcinomas from 37 centres. J Pathol 199: 418–423
Vogel DL et al. (2002) Efficacy and safety of trastuzumab as a single agent in first-line treatment of HER2-overexpressing metastatic breast cancer. J Clin Oncol 20: 719–726
DiGiovanna MP et al. (2005) Relationship of epidermal growth factor receptor expression to ErbB-2 signaling activity and prognosis in breast cancer patients. J Clin Oncol 23: 1152–1160
Thor AD et al. (2000) Activation (tyrosine phosphorylation) of ErbB-2 (HER-2/neu): a study of incidence and correlation with outcome in breast cancer. J Clin Oncol 18: 3230–3239
Tripathy D (2005) Targeted therapies in breast cancer. Breast J 11 (Suppl 1): S30–S35
Engelman JA and Janne PA (2005) Factors predicting response to EGFR tyrosine kinase inhibitors. Semin Resp Crit Care Med 26: 314–322
Giaccone G (2005) Epidermal growth factor receptor inhibitors in the treatment of non-small-cell lung cancer. J Clin Oncol 23: 3235–3242
Baselga J et al. (2002) Phase I safety, pharmacokinetic, and pharmacodynamic trial of ZD1839, a selective, oral epidermal growth factor receptor tyrosine kinase inhibitor, in patients with five selected solid tumor types. J Clin Oncol 20: 4292–4302
Herbst RS et al. (2002) Selective oral epidermal growth factor receptor tyrosine kinase inhibitor ZD1839 is generally well-tolerated and has activity in non-small-cell lung cancer and other solid tumors: results of a phase I trial. J Clin Oncol 20: 3815–3825
Fukuoka M et al. (2003) Multi-institutional randomized phase II trial of gefitinib for previously treated patients with advanced non-small-cell lung cancer. J Clin Oncol 21: 2237–2246
Kris MG et al. (2003) Efficacy of gefitinib, an inhibitor of the epidermal growth factor receptor tyrosine kinase, in symptomatic patients with non-small cell lung cancer: a randomized trial. JAMA 290: 2149–2158
Bailey R et al. (2003) Gefitinib, (“Iressa”, ZD1839) monotherapy for pretreated advance non-small cell lung cancer in IDEAL 1 and 2: tumor response is not clinically relevantly predictable from tumor EGFR membrane staining alone. Lung Cancer 41: 71
Giaccone G et al. (2004) Gefitinib in combination with gemcitabine and cisplatin in advanced non-small-cell lung cancer: a phase III trial—INTACT 1. J Clin Oncol 22: 777–784
Lynch TJ et al. (2004) Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 350: 2129–2139
Paez JG et al. (2004) EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 304: 1497–1500
Cappuzzo F et al. (2004) Akt phosphorylation and gefitinib efficacy in patients with advance non-small-cell lung cancer. J Natl Cancer Inst 96: 1133–1141
Narayanan S (2000) The preanalytic phase: an important component of laboratory medicine. Am J Clin Pathol 113: 429–452
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Wulfkuhle, J., Edmiston, K., Liotta, L. et al. Technology Insight: pharmacoproteomics for cancer—promises of patient-tailored medicine using protein microarrays. Nat Rev Clin Oncol 3, 256–268 (2006). https://doi.org/10.1038/ncponc0485
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ncponc0485
This article is cited by
-
Histopathological markers of treatment response and recurrence risk in ovarian cancers and borderline tumors
Der Pathologe (2017)
-
Capillary nano-immunoassays: advancing quantitative proteomics analysis, biomarker assessment, and molecular diagnostics
Journal of Translational Medicine (2015)
-
Role of Bruton’s tyrosine kinase inhibitors in HIV-1-infected cells
Journal of NeuroVirology (2015)
-
HER-2 assessment in formalin-fixed paraffin-embedded breast cancer tissue by well-based reverse phase protein array
Clinical Proteomics (2014)
-
Activation of the PI3K/AKT pathway correlates with prognosis in stage II colon cancer
British Journal of Cancer (2014)