Abstract
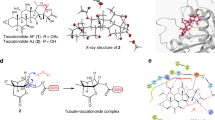
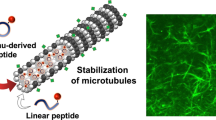
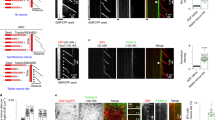
Microtubules are polymeric structures formed by the self-assembly of tubulin dimers. The growth and shrinkage of these dynamic arrays have a key role during the cell-proliferation process. This makes tubulin the molecular target of many anticancer drugs currently in use or under clinical trial. Their impressive success is limited by the onset of resistant tumour cells during the treatment, so new resistance-proof molecules need to be developed. Here we use molecular dynamics and free-energy calculations to study the network of interactions that allow microtubule formation. Modelling the protein–protein interface allows us to identify the amino acids responsible for tubulin–tubulin binding and thus to design peptides, which correspond to tubulin subsequences, that interfere with microtubule formation. We show that the application of molecular modelling techniques leads to the identification of peptides that exhibit antitubulin activity both in vitro and in cultured cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jordan, M. A. & Wilson, L. Microtubules as a target for anticancer drugs. Nature Rev. Cancer 4, 253–265 (2004).
Nogales, E. A structural view of microtubule dynamics. Cell. Mol. Life Sci. 56, 133–142 (1999).
Manfredi, J. J., Parness J. & Horwitz, S. B. Taxol binds to cellular microtubules. J. Cell. Biol. 94, 688–696 (1982).
Hastie, S. B. Interactions of colchicine with tubulin. Pharmacol. Ther. 512, 377–401 (1991).
Duflos, A., Kruczynski, A. & Barret, J. M. Novel aspects of natural and modified vinca alkaloids. Curr. Med. Chem. Anticancer Agents 2, 55–70 (2002).
Hamel, E. Antimitotic natural products and their interactions with tubulin. Med. Res. Rev. 16, 207–231 (1996).
Torin Huzil, J., Chen, K., Kurgan, L. & Tuszynski, J. A. The role of β-tubulin mutations and isotype expression in acquired drug resistance. Cancer Inf. 3, 159–181 (2007).
Orr, G. A., Verdier-Pinard, P., McDaid, H. & Horwits, S. A. Mechanisms of taxol resistance related to microtubules. Oncogene 22, 7280–7295 (2003).
Wells, J. A. & McClendon, C. L. Reaching for high-hanging fruit in drug discovery at protein–protein interfaces. Nature 450, 1001–1009 (2007).
Lo Conte, L., Chothia, C. & Janin, J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
Chène, P. Drugs targeting protein–protein interactions. ChemMedChem 1, 400–411 (2006).
Arkin, M. R. & Wells, J. A. Small-molecule inhibitors of protein–protein interactions: progressing towards the dream. Nature Rev. Drug Disc. 3, 301–317 (2004).
Janin, Y. L. Peptides with anticancer use or potential. Amino Acids 25, 1–40 (2003).
Berg, T. Modulation of protein–protein interactions with small organic molecules. Angew. Chem. Int. Ed. 42, 2462–2481 (2003).
Lowman, H. B. Bacteriophage display and discovery of peptide leads for drug development. Annu. Rev. Biophys. Biomol. Struct. 26, 401–424 (1997)
Sidhu, S. S., Fairbrothe, W. J. & Deshayes, K. Exploring protein–protein interactions with phage display. ChemBioChem 4, 14–25 (2003).
Sattler, M. et al. Structure of Bcl–XL–Bak peptide complex: recognition between regulators of apoptosis. Science 275, 983–986 (1997).
Kussie, P.H et al. Structure of the MDM2 oncoprotein bound to the p53 tumor suppressor transactivation domain. Science 274, 948–953 (1996).
Villacañas O. & Rubio-Martinez, J. Reducing CDK4/6-p16INK4a interface: computational alanine scanning of a peptide bound to CDK6 protein. Proteins 63, 797–810 (2006).
Zhong, H. & Carlson H. A. Computational studies and peptidomimetic design for the human p53–MDM2 complex. Proteins 58, 222–234 (2005).
Massova, I. & Kollman, P. A. Combined molecular mechanical and continuum solvent approach (MM-PBSA/GBSA) to predict ligand binding. Persp. Drug Disc. Des. 18, 113–135 (2000).
Massova, I. & Kollman, P. A. Computational alanine scanning to probe protein–protein interactions: a novel approach to evaluate binding free energies. J. Am. Chem. Soc. 121, 8133–8143 (1999).
Moreira, I. S., Fernandes, P. A. & Ramos, M. J. Unravelling the importance of protein–protein interactions: application of a computational alanine scanning mutagenesis to the study of the IgG1 streptococcal protein G (C2 fragment) complex. J. Phys. Chem. B 110, 10962–10969 (2006).
Moreira, I. S., Fernandes, P. A. & Ramos, M. J. Unravelling hot spots: a comprehensive computational mutagenesis study. Theor. Chem. Acc. 117, 99–113 (2007).
Nogales, E. A structural view of microtubule dynamics. Cell. Mol. Life Sci. 56, 133–142 (1999).
Wordeman, L. & Mitchison, T. J. in Microtubules (eds Hyams, J. S. & Lloyd, C. W.) 287–302 (Wiley-Liss, 1994).
McIntosh, J. R. in Microtubules (eds Hyams, J. S. & Lloyd, C. W.) 413–434 (Wiley-Liss, 1994).
Waterman-Storer, C. & Salmon, E. D. Microtubule dynamics: treadmilling comes around again. Curr. Biol. 7, 369–372 (1997).
Nogales, E., Whittaker, M., Milligan, R. A. & Downing, K. H. High-resolution model of the microtubule. Cell 96, 79–88 (1999).
Löwe, J., Li, H., Downing, K. H. & Nogales, E. Refined structure of αβ-tubulin at 3.5 Å resolution. J. Mol. Biol. 313, 1045–1057 (2001).
Richards, K. L. et al. Structure–function relationship in yeast tubulins. Mol. Biol. Cell 11, 1887–1903 (2000).
Reijo, R. A., Cooper, E. M., Beagle, G. J. & Huffaker, T. C. Systematic mutational analysis of the yeast beta-tubulin gene. Mol. Biol. Cell 5, 29–43 (1994).
Anders, K. R. & Botstein, D. Dominant-lethal α-tubulin mutants defective in microtubule depolymerization in yeast. Mol. Biol. Cell 12, 3973–3986 (2001).
Humphrey, W., Dalke, A. & Schulten, K. VMD – Visual Molecular Dynamics. J. Mol. Graphics 14, 33–38 (1996).
Johnson, K. A. & Borisy, G. G. Kinetic analysis of microtubule self-assembly in vitro. J. Mol. Biol. 117, 1–31 (1977).
Nogales, E., Wolf, S. G. & Downing, K. H. Structure of the alpha beta tubulin dimer by electron crystallography. Nature 391, 199–203 (1998).
Case, D. A., et al. AMBER 9 (University of California, San Francisco, 2006).
Duan, Y., et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
Mita, A. & Sept, D. Localization of the antimitotic peptide and depsipeptide binding site on β tubulin. Biochemistry 43, 13955–13962 (2004).
Zoete, V. & Michielin, O. Comparison between computational alanine scanning and per-residue binding free energy decomposition for protein–protein association using MM-GBSA: application to the TCR-p-MHC complex. Proteins 67, 1026–1047 (2007).
Acknowledgements
S. Rendine is acknowledged for useful discussions.
Author information
Authors and Affiliations
Contributions
S.P. and M.S. conceived and designed the research project; G.Sa. G.C., D.C., P.F., G.Sp. and P.M. performed the experiments; S.P., G.Sa., G.C. and M.S. analysed the data and co-wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Supplementary information
Supplementary information
Supplementary information (PDF 843 kb)
Rights and permissions
About this article
Cite this article
Pieraccini, S., Saladino, G., Cappelletti, G. et al. In silico design of tubulin-targeted antimitotic peptides. Nature Chem 1, 642–648 (2009). https://doi.org/10.1038/nchem.401
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.401
This article is cited by
-
Microtubule Destabilization Paves the Way to Parkinson’s Disease
Molecular Neurobiology (2017)
-
Discovery of new druggable sites in the anti-cholesterol target HMG-CoA reductase by computational alanine scanning mutagenesis
Journal of Molecular Modeling (2014)
-
Making drug design second nature
Nature Chemistry (2009)