Abstract
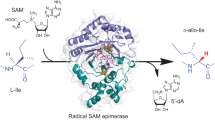
Ribosomally synthesized peptides are built out of L-amino acids, whereas D-amino acids are generally the hallmark of non-ribosomal synthetic processes. Here we show that the model bacterium Bacillus subtilis is able to produce a novel type of ribosomally synthesized and post-translationally modified peptide that contains D-amino acids, and which we propose to call epipeptides. We demonstrate that a two [4Fe–4S]-cluster radical S-adenosyl-L-methionine (SAM) enzyme converts L-amino acids into their D-counterparts by catalysing Cα-hydrogen-atom abstraction and using a critical cysteine residue as the hydrogen-atom donor. Unexpectedly, these D-amino acid residues proved to be essential for the activity of a peptide that induces the expression of LiaRS, a major component of the bacterial cell envelope stress-response system. Present in B. subtilis and in several members of the human microbiome, these epipeptides and radical SAM epimerases broaden the landscape of peptidyl structures accessible to living organisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jordan, S., Junker, A., Helmann, J. D. & Mascher, T. Regulation of LiaRS-dependent gene expression in Bacillus subtilis: identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system. J. Bacteriol. 188, 5153–5166 (2006).
Jordan, S., Hutchings, M. I. & Mascher, T. Cell envelope stress response in Gram-positive bacteria. FEMS Microbiol. Rev. 32, 107–146 (2008).
Mascher, T., Margulis, N. G., Wang, T., Ye, R. W. & Helmann, J. D. Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon. Mol. Microbiol. 50, 1591–1604 (2003).
Mascher, T., Zimmer, S. L., Smith, T. A. & Helmann, J. D. Antibiotic-inducible promoter regulated by the cell envelope stress-sensing two-component system LiaRS of Bacillus subtilis. Antimicrob. Agents Chemother. 48, 2888–2896 (2004).
Butcher, B. G., Lin, Y. P. & Helmann, J. D. The yydFGHIJ operon of Bacillus subtilis encodes a peptide that induces the LiaRS two-component system. J. Bacteriol. 189, 8616–8625 (2007).
Benjdia, A. & Berteau, O. Sulfatases and radical SAM enzymes: emerging themes in glycosaminoglycan metabolism and the human microbiota. Biochem. Soc. Trans. 44, 109–115 (2016).
Vey, J. L. & Drennan, C. L. Structural insights into radical generation by the radical SAM superfamily. Chem. Rev. 111, 2487–2506 (2011).
Broderick, J. B., Duffus, B. R., Duschene, K. S. & Shepard, E. M. Radical S-adenosylmethionine enzymes. Chem. Rev. 114, 4229–4317 (2014).
Walsby, C. J. et al. Electron-nuclear double resonance spectroscopic evidence that S-adenosylmethionine binds in contact with the catalytically active [4Fe–4S]+ cluster of pyruvate formate-lyase activating enzyme. J. Am. Chem. Soc. 124, 3143–3151 (2002).
Nicolet, Y., Amara, P., Mouesca, J. M. & Fontecilla-Camps, J. C. Unexpected electron transfer mechanism upon AdoMet cleavage in radical SAM proteins. Proc. Natl Acad. Sci. USA 106, 14867–14871 (2009).
Frey, P. A., Hegeman, A. D. & Ruzicka, F. J. The radical SAM superfamily. Crit. Rev. Biochem. Mol. Biol. 43, 63–88 (2008).
Pierre, S. et al. Thiostrepton tryptophan methyltransferase expands the chemistry of radical SAM enzymes. Nat. Chem. Biol. 8, 957–959 (2012).
Wang, S. C. & Frey, P. A. Binding energy in the one-electron reductive cleavage of S-adenosylmethionine in lysine 2,3-aminomutase, a radical SAM enzyme. Biochemistry 46, 12889–12895 (2007).
Horitani, M. et al. Why nature uses radical SAM enzymes so widely: electron nuclear double resonance studies of lysine 2,3-aminomutase show the 5′-dAdo* ‘free radical’ is never free. J. Am. Chem. Soc. 137, 7111–7121 (2015).
Wang, S. C. & Frey, P. A. S-adenosylmethionine as an oxidant: the radical SAM superfamily. Trends Biochem. Sci. 32, 101–110 (2007).
Chandor, A. et al. Dinucleotide spore photoproduct, a minimal substrate of the DNA repair spore photoproduct lyase enzyme from Bacillus subtilis. J. Biol. Chem. 281, 26922–26931 (2006).
Benjdia, A. DNA photolyases and SP lyase: structure and mechanism of light-dependent and independent DNA lyases. Curr. Opin. Struct. Biol. 22, 711–720 (2012).
Benjdia, A., Heil, K., Barends, T. R. M., Carell, T. & Schlichting, I. Structural insights into recognition and repair of UV-DNA damage by spore photoproduct lyase, a radical SAM enzyme. Nucleic Acids Res 40, 9308–9318 (2012).
Sun, X. et al. The free radical of the anaerobic ribonucleotide reductase from Escherichia coli is at glycine 681. J. Biol. Chem. 271, 6827–6831 (1996).
Benjdia, A., Deho, G., Rabot, S. & Berteau, O. First evidences for a third sulfatase maturation system in prokaryotes from E. coli aslB and ydeM deletion mutants. FEBS Lett. 581, 1009–1014 (2007).
Benjdia, A. et al. Anaerobic sulfatase-maturating enzymes: radical SAM enzymes able to catalyze in vitro sulfatase post-translational modification. J. Am. Chem. Soc. 129, 3462–3463 (2007).
Benjdia, A., Leprince, J., Sandstrom, C., Vaudry, H. & Berteau, O. Mechanistic investigations of anaerobic sulfatase-maturating enzyme: direct Cβ H-atom abstraction catalyzed by a radical AdoMet enzyme. J. Am. Chem. Soc. 131, 8348–8349 (2009).
Benjdia, A. et al. Anaerobic sulfatase-maturating enzymes—first dual substrate radical S-adenosylmethionine enzymes. J. Biol. Chem. 283, 17815–17826 (2008).
Arragain, S . et al. Post-translational modification of ribosomal proteins: structural and functional characterization of RimO from Thermotoga maritima, a radical-SAM methylthiotransferase. J. Biol. Chem. 258, 5792–5801 (2009).
Lee, K. H. et al. Characterization of RimO, a new member of the methylthiotransferase subclass of the radical SAM superfamily. Biochemistry 48, 10162–10174 (2009).
Flühe, L. K. et al. The radical SAM enzyme AlbA catalyzes thioether bond formation in subtilosin A. Nat. Chem. Biol. 8, 350–357 (2012).
Benjdia, A. et al. Thioether bond formation by SPASM domain radical SAM enzymes: Cα H-atom abstraction in subtilosin A biosynthesis. Chem. Commun. 52, 6249–6252 (2016).
Arnison, P. G. et al. Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160 (2013).
Freeman, M. F. et al. Metagenome mining reveals polytheonamides as posttranslationally modified ribosomal peptides. Science 338, 387–390 (2012).
Huo, L., Rachid, S., Stadler, M., Wenzel, S. C. & Muller, R. Synthetic biotechnology to study and engineer ribosomal bottromycin biosynthesis. Chem. Biol. 19, 1278–1287 (2012).
Duin, E. C. et al. [2Fe–2S] to [4Fe–4S] cluster conversion in Escherichia coli biotin synthase. Biochemistry 36, 11811–11820 (1997).
Bansal, P. S. et al. Substrate specificity of platypus venom L-to-D-peptide isomerase. J. Biol. Chem. 283, 8969–8975 (2008).
Morinaka, B. I. et al. Radical S-adenosyl methionine epimerases: regioselective introduction of diverse D-amino acid patterns into peptide natural products. Angew. Chem. Int. Ed. 53, 8503–8507 (2014).
Burkhart, B. J., Hudson, G. A., Dunbar, K. L. & Mitchell, D. A. A prevalent peptide-binding domain guides ribosomal natural product biosynthesis. Nat. Chem. Biol. 11, 564–570 (2015).
Benjdia, A. et al. Anaerobic sulfatase-maturating enzyme—a mechanistic link with glycyl radical-activating enzymes? FEBS J. 277, 1906–1920 (2010).
Grell, T. A., Goldman, P. J. & Drennan, C. L. SPASM and twitch domains in S-adenosylmethionine (SAM) radical enzymes. J. Biol. Chem. 290, 3964–3971 (2015).
Kudo, F., Hoshi, S., Kawashima, T., Kamachi, T. & Eguchi, T. Characterization of a radical S-adenosyl-L-methionine epimerase, NeoN, in the last step of neomycin B biosynthesis. J. Am. Chem. Soc. 136, 13909–13915 (2014).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8, 275–282 (1992).
Zharkikh, A. & Li, W. H. Estimation of confidence in phylogeny: the complete-and-partial bootstrap technique. Mol. Phylogenet. Evol. 4, 44–63 (1995).
Gerlt, J. A. et al. Enzyme function initiative-enzyme similarity tool (EFI-EST): a web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 1854, 1019–1037 (2015).
Benjdia, A., Heil, K., Winkler, A., Carell, T. & Schlichting, I. Rescuing DNA repair activity by rewiring the H-atom transfer pathway in the radical SAM enzyme, spore photoproduct lyase. Chem. Commun. 50, 14201–14204 (2014).
Chandor-Proust, A. et al. DNA repair and free radicals, new insights into the mechanism of spore photoproduct lyase revealed by single amino acid substitution. J. Biol. Chem. 283, 36361–8 (2008).
Wagner, A. F., Frey, M., Neugebauer, F. A., Schafer, W. & Knappe, J. The free radical in pyruvate formate-lyase is located on glycine-734. Proc. Natl Acad. Sci. USA 89, 996–1000 (1992).
Reichard, P. & Ehrenberg, A. Ribonucleotide reductase—a radical enzyme. Science 221, 514–519 (1983).
Moore, B. N. & Julian, R. R. Dissociation energies of X–H bonds in amino acids. Phys. Chem. Chem. Phys. 14, 3148–3154 (2012).
Stubbe, J., Nocera, D. G., Yee, C. S. & Chang, M. C. Radical initiation in the class I ribonucleotide reductase: long-range proton-coupled electron transfer? Chem. Rev. 103, 2167–2201 (2003).
Stubbe, J. & van der Donk, W. A. Ribonucleotide reductases: radical enzymes with suicidal tendencies. Chem. Biol. 2, 793–801 (1995).
Staples, C. R. et al. The function and properties of the iron–sulfur center in spinach ferredoxin: thioredoxin reductase: a new biological role for iron–sulfur clusters. Biochemistry 35, 11425–11434 (1996).
Bennati, M., Weiden, N., Dinse, K. P. & Hedderich, R. 57Fe ENDOR spectroscopy on the iron–sulfur cluster involved in substrate reduction of heterodisulfide reductase. J. Am. Chem. Soc. 126, 8378–8379 (2004).
Duin, E. C., Madadi-Kahkesh, S., Hedderich, R., Clay, M. D. & Johnson, M. K. Heterodisulfide reductase from Methanothermobacter marburgensis contains an active-site [4Fe–4S] cluster that is directly involved in mediating heterodisulfide reduction. FEBS Lett. 512, 263–268 (2002).
Dai, S., Schwendtmayer, C., Schurmann, P., Ramaswamy, S. & Eklund, H. Redox signaling in chloroplasts: cleavage of disulfides by an iron-sulfur cluster. Science 287, 655–658 (2000).
Walters, E. M. et al. Spectroscopic characterization of site-specific [Fe4S4] cluster chemistry in ferredoxin:thioredoxin reductase: implications for the catalytic mechanism. J. Am. Chem. Soc. 127, 9612–9624 (2005).
Nicolas, P. et al. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science 335, 1103–1106 (2012).
Craig, R., Cortens, J. C., Fenyo, D. & Beavis, R. C. Using annotated peptide mass spectrum libraries for protein identification. J. Proteome Res. 5, 1843–1849 (2006).
Radeck, J. et al. Anatomy of the bacitracin resistance network in Bacillus subtilis. Mol. Microbiol. 100, 607–620 (2016).
Wolf, D., Dominguez-Cuevas, P., Daniel, R. A. & Mascher, T. Cell envelope stress response in cell wall-deficient L-forms of Bacillus subtilis. Antimicrob. Agents Chemother. 56, 5907–5915 (2012).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459 (2015).
Ellermeier, C. D., Hobbs, E. C., Gonzalez-Pastor, J. E. & Losick, R. A three-protein signaling pathway governing immunity to a bacterial cannibalism toxin. Cell 124, 549–559 (2006).
Wilson, M. C. et al. An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506, 58–62 (2014).
Cotter, P. D., Ross, R. P. & Hill, C. Bacteriocins—a viable alternative to antibiotics? Nat. Rev. Microbiol. 11, 95–105 (2013).
Acknowledgements
This work was supported by grants from European Research Council (Consolidator Grant 617053 to O.B.). High-resolution MS analyses were performed on the INRA PAPPSO proteomics platform.
Author information
Authors and Affiliations
Contributions
A.B. and O.B. conceived and designed the experiments; A.B., A.G., P.R., J.L. and O.B. performed the experiments; A.B., A.G, J.L. and O.B. analysed the data; A.B. and O.B. co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 3263 kb)
Rights and permissions
About this article
Cite this article
Benjdia, A., Guillot, A., Ruffié, P. et al. Post-translational modification of ribosomally synthesized peptides by a radical SAM epimerase in Bacillus subtilis. Nature Chem 9, 698–707 (2017). https://doi.org/10.1038/nchem.2714
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2714
This article is cited by
-
Structural and mechanistic basis for RiPP epimerization by a radical SAM enzyme
Nature Chemical Biology (2024)
-
Interspecies interaction reduces selection for antibiotic resistance in Escherichia coli
Communications Biology (2023)
-
Crystallographic snapshots of a B12-dependent radical SAM methyltransferase
Nature (2022)
-
Bicyclostreptins are radical SAM enzyme-modified peptides with unique cyclization motifs
Nature Chemical Biology (2022)
-
Biosynthesis-guided discovery reveals enteropeptins as alternative sactipeptides containing N-methylornithine
Nature Chemistry (2022)