Abstract
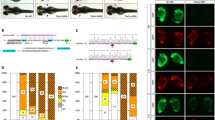
We describe the use of zinc-finger nucleases (ZFNs) for somatic and germline disruption of genes in zebrafish (Danio rerio), in which targeted mutagenesis was previously intractable. ZFNs induce a targeted double-strand break in the genome that is repaired to generate small insertions and deletions. We designed ZFNs targeting the zebrafish golden and no tail/Brachyury (ntl) genes and developed a budding yeast–based assay to identify the most active ZFNs for use in vivo. Injection of ZFN-encoding mRNA into one-cell embryos yielded a high percentage of animals carrying distinct mutations at the ZFN-specified position and exhibiting expected loss-of-function phenotypes. Over half the ZFN mRNA-injected founder animals transmitted disrupted ntl alleles at frequencies averaging 20%. The frequency and precision of gene-disruption events observed suggest that this approach should be applicable to any loci in zebrafish or in other organisms that allow mRNA delivery into the fertilized egg.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lieschke, G.J. & Currie, P.D. Animal models of human disease: zebrafish swim into view. Nat. Rev. Genet. 8, 353–367 (2007).
Nasevicius, A. & Ekker, S.C. Effective targeted gene 'knockdown' in zebrafish. Nat. Genet. 26, 216–220 (2000).
Stemple, D.L. TILLING–a high-throughput harvest for functional genomics. Nat. Rev. Genet. 5, 145–150 (2004).
Thomas, K.R., Folger, K.R. & Capecchi, M.R. High frequency targeting of genes to specific sites in the mammalian genome. Cell 44, 419–428 (1986).
Sedivy, J.M. & Joyner, A.L. Gene Targeting. (Oxford University Press, Oxford, 1992).
Kim, Y.G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 93, 1156–1160 (1996).
Smith, J., Berg, J. & Chandrasegaran, S. A detailed study of the substrate specificity of a chimeric restriction enzyme. Nucleic Acids Res. 27, 674–681 (1999).
Valerie, K. & Povirk, L.F. Regulation and mechanisms of mammalian double-strand break repair. Oncogene 22, 5792–5812 (2003).
Bibikova, M., Golic, M., Golic, K.G. & Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161, 1169–1175 (2002).
Morton, J., Davis, M.W., Jorgensen, E.M. & Carroll, D. Induction and repair of zinc-finger nuclease-targeted double-strand breaks in Caenorhabditis elegans somatic cells. Proc. Natl. Acad. Sci. USA 103, 16370–16375 (2006).
Santiago, Y. et al. Targeted gene knockout in mammalian cells using engineered zinc-finger nucleases. Proc. Natl. Acad. Sci. USA 105, 5809–5814 (2008).
Lamason, R.L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005).
Streisinger, G., Coale, F., Taggart, C., Walker, C. & Grunwald, D.J. Clonal origins of cells in the pigmented retina of the zebrafish eye. Dev. Biol. 131, 60–69 (1989).
Pabo, C.O., Peisach, E. & Grant, R.A. Design and selection of novel Cys2His2 zinc finger proteins. Annu. Rev. Biochem. 70, 313–340 (2001).
Haber, J.E. In vivo biochemistry: physical monitoring of recombination induced by site-specific endonucleases. Bioessays 17, 609–620 (1995).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Halpern, M.E., Ho, R.K., Walker, C. & Kimmel, C.B. Induction of muscle pioneers and floor plate is distinguished by the zebrafish no tail mutation. Cell 75, 99–111 (1993).
Urnov, F.D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005).
Miller, J., McLachlan, A.D. & Klug, A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 4, 1609–1614 (1985).
Pavletich, N.P. & Pabo, C.O. Zinc finger-DNA recognition: crystal structure of a Zi268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Choo, Y., Sanchez-Garcia, I. & Klug, A. In vivo repression by a site-specific DNA-binding protein designed against an oncogenic sequence. Nature 372, 642–645 (1994).
Zhang, L. et al. Synthetic zinc finger transcription factor action at an endogenous chromosomal site. J. Biol. Chem. 275, 33850–33860 (2000).
Voth, W.P., Richards, J.D., Shaw, J.M. & Stillman, D.J. Yeast vectors for integration at the HO locus. Nucleic Acids Res. 29, E59 (2001).
Mumberg, D., Muller, R. & Funk, M. Regulatable promoters of Saccharomyces cerevisiae: a comparison of transcriptional activity and their use for heterologous expression. Nucleic Acids Res. 22, 5767–5768 (1994).
Haber, J.E. Uses and abuses of HO endonuclease. Methods Enzymol. 350, 141–164 (2002).
Chen, Q. et al. A yeast two-hybrid technology-based system for the discovery of PPARgamma agonist and antagonist. Anal. Biochem. 335, 253–259 (2004).
Kimmel, C.B., Ballard, W.W., Kimmel, S.R., Ullmann, B. & Schilling, T.F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Schulte-Merker, S., van Eeden, F.J., Halpern, M.E., Kimmel, C.B. & Nüsslein-Volhard, C. no tail (ntl) is the zebrafish homologue of the mouse T (Brachyury) gene. Development 120, 1009–1015 (1994).
Jowett, T. Analysis of protein and gene expression. Methods Cell Biol. 59, 63–85 (1999).
Acknowledgements
We thank Jasper Rine, Michael Holmes, Jacques Côté and Amine Nourani for helpful suggestions regarding yeast-based screening of ZFNs, Randy Morse and Amine Nourani for providing reagents, Keith Cheng for gol cDNA, Keith Cheng and Herwig Baier for gol stocks, and Jennifer St. Hilaire, Emily Janus and Kimberly Blum for excellent zebrafish care and technical support. We thank the anonymous referees for helpful suggestions on the manuscript. The ntlb487 allele was isolated in Eugene, Oregon, in a National Institutes of Health (NIH)-funded screen (HD22486); S.L.A. gratefully acknowledges Charles Kimmel and our many colleagues at the University of Oregon who participated in mutagenesis screening. This work was supported by grants to S.L.A. from the NIH (1-R01-GM061952) and March of Dimes Birth Defects Foundation (1FY05-118). J.M.M. is supported by a National Science Foundation Predoctoral Fellowship.
Author information
Authors and Affiliations
Contributions
F.D.U. and S.L.A. conceived the project; Y.D., J.M.M., J.C.M., L.Z., E.J.R., F.D.U. and S.L.A. designed experiments; Y.D., J.M.M., F.F., C.N., G.E.K., R.A., T.D.H. and S.L.A. performed experiments; Y.D., J.M.M., P.D.G., F.D.U. and S.L.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
All of the authors, except J.M.M. and S.L.A., are full-time employees of Sangamo BioSciences, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Tables 1–4, Figures 1–11 (PDF 1677 kb)
Rights and permissions
About this article
Cite this article
Doyon, Y., McCammon, J., Miller, J. et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat Biotechnol 26, 702–708 (2008). https://doi.org/10.1038/nbt1409
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1409
This article is cited by
-
Effects of editing DFR genes on flowers, leaves, and roots of tobacco
BMC Plant Biology (2023)
-
Large-scale F0 CRISPR screens in vivo using MIC-Drop
Nature Protocols (2023)
-
Genome centric engineering using ZFNs, TALENs and CRISPR-Cas9 systems for trait improvement and disease control in Animals
Veterinary Research Communications (2023)
-
The Zebrafish model in dermatology: an update for clinicians
Discover Oncology (2022)