Abstract
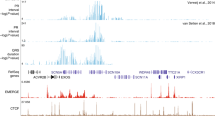
The SCL gene encodes a highly conserved bHLH transcription factor with a pivotal role in hemopoiesis and vasculogenesis. We have sequenced and analyzed 320 kb of genomic DNA composing the SCL loci from human, mouse, and chicken. Long-range sequence comparisons demonstrated multiple peaks of human/mouse homology, a subset of which corresponded precisely with known SCL enhancers. Comparisons between mammalian and chicken sequences identified some, but not all, SCL enhancers. Moreover, one peak of human/mouse homology (+23 region), which did not correspond to a known enhancer, showed significant homology to an analogous region of the chicken SCL locus. A transgenic Xenopus reporter assay was established and demonstrated that the +23 region contained a new neural enhancer. This combination of long-range comparative sequence analysis with a high-throughput transgenic bioassay provides a powerful strategy for identifying and characterizing developmentally important enhancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Popperl, H. et al. Segmental expression of Hoxb-1 is controlled by a highly conserved autoregulatory loop dependent upon exd/pbx. Cell 81, 1031 –1042 (1995).
Aparicio, S. et al. Detecting conserved regulatory elements with the model genome of the Japanese puffer fish, Fugu rubripes. Proc Natl. Acad. Sci. USA 92, 1684–1688 ( 1995).
Hardison, R.C., Oeltjen, J. & Miller, W. Long human-mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997).
Rowitch, D.H. et al. Identification of an evolutionary conserved 110 base-pair cis -acting regulatory sequence that governs wnt-1 expression in the murine neural plate. Development 125, 2735– 2746 (1998).
Nonchev, S. et al. The conserved role of Krox-20 in directing Hox gene-expression during vertebrate hindbrain segmentation. Proc Natl. Acad. Sci. USA 93 , 9339–9345 (1996).
Oeltjen, J.C. et al. Large-scale comparative sequence analysis of the human and murine Bruton's tyrosine kinase loci reveals conserved regulatory domains. Genome Res. 7, 315–329 ( 1997).
Lamerdin, J.E. et al. Genomic sequence comparison of the human and mouse XRCC1 DNA repair gene regions. Genomics 25, 547– 554 (1995).
Lamerdin, J.E., Stilwagen, S.A., Ramirez, M.H., Stubbs, L. & Carrano, A.V. Sequence analysis of the ERCC2 gene regions in human, mouse, and hamster reveals three linked genes. Genomics 34, 399–409 ( 1996).
Epp, T.A., Wang, R., Sole, M.J. & Liew, C.C. Concerted evolution of mammalian cardiac myosin heavy chain genes. J. Mol. Evol. 41, 284–292 (1995).
Hardison, R. et al. Locus control regions of mammalian beta-globin gene clusters: combining phylogenetic analyses and experimental results to gain functional insights . Gene 205, 73–94 (1997).
Begley, C.G. & Green, A.R. The SCL gene: from case report to critical hematopoietic regulator. Blood 93, 2760–70 (1999).
Porcher, C. et al. The T cell leukemia oncoprotein SCL/tal-1 is essential for development of all hematopoietic lineages. Cell 86, 47–57 (1996).
Robb, L. et al. The SCL gene product is required for the generation of all hematopoietic lineages in the adult mouse. EMBO J. 15, 4123– 4129 (1996).
Visvader, J.E., Fujiwara, Y. & Orkin, S.H. Unsuspected role for the T-cell leukemia protein SCL/tal-1 in vascular development. Genes Dev. 12, 473–479 (1998).
Gering, M., Rodaway, A.R.F., Göttgens, B., Patient, R.K. & Green, A.R. The SCL gene specifies haemangioblast development from early mesoderm. EMBO J. 17, 4029–4045 (1998).
Drake, C.J., Brandt, S.J., Trusk, T.C. & Little, C.D. TAL1/SCL is expressed in endothelial progenitor cells/angioblasts and defines a dorsal-to-ventral gradient of vasculogenesis. Dev. Biol. 192, 17–30 (1997).
Sinclair, A.M. et al. Distinct 5′ SCL enhancers direct transcription to developing brain, spinal chord and endothelium; conserved neural expression is GATA factor dependent. Dev. Biol. 209, 128– 142 (1999).
Mead, P.E., Kelley, C.M., Hahn, P.S., Piedad, O. & Zon, L.I. SCL specifies hematopoietic mesoderm in Xenopus embryos . Development 125, 2611– 2620 (1998).
Lecointe, N. et al. GATA- and SP1-binding sites are required for the full activity of the tissue-specific promoter of the tal-1 gene. Oncogene 9, 2623–2632 (1994).
Bockamp, E.-O et al. Lineage-restricted regulation of the murine SCL/TAL-1 promoter. Blood 86, 1502–1514 ( 1995).
Bockamp, E.O. et al. Distinct mechanisms direct SCL/tal-1 expression in erythroid cells and CD34 positive primitive myeloid cells. J. Biol. Chem. 272, 8781–8790 (1997).
Bockamp, E.O. et al. Transcriptional regulation of the stem cell leukemia gene by PU.1 and Elf-1. J. Biol. Chem. 273, 29032– 29042 (1998).
Göttgens, B. et al. Transcription of the SCL gene in erythroid and CD34 positive primitive myeloid cells is controlled by a complex network of lineage-restricted chromatin-dependent and chromatin-independent regulatory elements. Oncogene 15, 2419–2428 (1997).
Sanchez, M.J. et al. An SCL 3′ enhancer targets developing endothelium together with embryonic and adult haematopoietic progenitors. Development 126, 3891–3904 ( 1999).
Aplan, P.D. et al. The SCL gene is formed from a transcriptionally complex locus. Mol. Cell Biol. 10, 6426–6435 (1990).
Begley, C.G. et al. Structure of the gene encoding the murine SCL protein. Gene 138, 93–99 ( 1994).
Morgenstern, B., Frech, K., Dress, A. & Werner, T. DIALIGN: Finding local similarities by multiple sequence alignment. Bioinformatics 14, 290–294 ( 1998).
Sonnhammer, E.L. & Durbin, R. A dot-matrix program with dynamic threshold control suited for genomic DNA and protein sequence analysis. Gene 167, GC1– GC10 (1995).
Leroy-Viard, K., Vinit, M.-A, Lecointe, N., Mathieu-Mahul, D. & Romeo, P.-H Distinct DNase-I hypersensitive sites are associated with TAL-1 transcription in erythroid and T-cell lines . Blood 84, 3619–3827 (1994).
Xu, G. & Goodridge, A.G. Characterization of a polypyrimidine/polypurine tract in the promoter of the gene for chicken malic enzyme. J. Biol. Chem. 271, 16008–16019 (1996).
Levitt, N., Briggs, D., Gil, A. & Proudfoot, N.J. Definition of an efficient synthetic poly(A) site. Genes Dev. 3, 1019–1025 (1989).
Moreira, A., Wollerton, M., Monks, J. & Proudfoot, N.J. Upstream sequence elements enhance poly(A) site efficiency of the C2 complement gene and are phylogenetically conserved. EMBO J. 14, 3809–3819 (1995).
Moreira, A. et al. The upstream sequence element of the C2 complement poly(A) signal activates mRNA 3′ end formation by two distinct mechanisms. Genes Dev. 12, 2522–2534 (1998).
Kroll, K.K. & Amaya, E. Transgenic Xenopus embryos from sperm nuclear transplantations reveal FGF signaling requirements during gastrulation. Development 122, 3173– 3183 (1996).
Kroll, K.K. & Amaya, E. in Early development of Xenopus laevis . (eds Sive, H.L., Grainger, R.M. & Harland, R.M.) 393– 414, (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1998).
Kallianpur, A.R., Jordan, J.E. & Brandt, S.J. The SCL/TAL-1 gene is expressed in progenitors of both the hematopoietic and vascular systems during embryogenesis. Blood 83, 1200–1208 ( 1994).
Green, A.R., Visvader, J., Lints, T., Harvey, R. & Begley, C.G. SCL is co-expressed with GATA-1 in haemopoietic cells but is also expressed in developing brain. Oncogene 7, 653–660 (1992).
Duret, L. & Bucher, P. Searching for regulatory elements in human noncoding sequences. Curr. Opin. Struct Biol. 7, 399–406 (1997).
Wingender, E., Karas, H. & Knuppel, R. TRANSFAC database as a bridge between sequence data libraries and biological function. Pac. Symp. Biocomput. 477–485 (1997).
Kolchanov, N.A. et al. GeneExpress: a computer system for description, analysis, and recognition of regulatory sequences in eukaryotic genome. Intelligent Systems for Molecular Biology 6, 95– 104 (1998).
Long, Q. et al. GATA-1 expression pattern can be recapitulated in living transgenic zebrafish using GFP reporter gene. Development 124, 4105–4111 (1997).
Meng, A. et al. Positive and negative cis-acting elements are required for hematopoietic expression of zebrafish GATA-1. Blood 93, 500–508 (1999).
Yuh, C.H. & Davidson, E.H. Modular cis-regulatory organization of Endo16, a gut-specific gene of the sea urchin embryo. Development 122, 1069–1082 (1996).
Kirchhamer, C.V. & Davidson, E.H. Spatial and temporal information processing in the sea urchin embryo: modular and intramodular organization of the CyIIIa gene cis-regulatory system. Development 122, 333–348 (1996).
Arnone, M.I. & Davidson, E.H. The hardwiring of development: organization and function of genomic regulatory systems. Development 124, 1851–1864 ( 1997).
Eeckman, F.H. & Durbin, R. ACeDB and macace. Methods Cell Biol. 48, 583–605 ( 1995).
Göttgens, B. et al. The pufferfish SLP-1 gene, a new member of the SCL/TAL-1 family of transcription factors. Genomics 48, 52–62 (1998).
Altschul, S.F., Gish, W., Miller, W., Meyers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 ( 1990).
Quandt, K., Frech, K., Karas, H., Wingender, E. & Werner, T. MatInd and MatInspector: new fast and versatile tools for detection of consensus matches in nucleotide sequence data. Nucleic Acids Res. 23, 4878–4884 (1995).
Acknowledgements
Work in the authors' laboratories is supported by the Wellcome Trust, the Medical Research Council, the Leukaemia Research Fund, and the Kay Kendall Leukaemia Fund. We are grateful to Christine Holt for expert help in characterizing neural expression patterns, Kathy Hartley for the polyGFP3 plasmid, Rima Zoorob for a filter set of the chicken PAC library, and Jonathan Frampton for the HD57 RNA. We have enjoyed stimulating discussions with Doug Higgs, Sam Aparicio, Rob Krumlauf, and Victor Solovyev, and would also like to acknowledge the unstinting support of Jane Rogers and sequencing team 32 at the Sanger Centre.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Göttgens, B., Barton, L., Gilbert, J. et al. Analysis of vertebrate SCL loci identifies conserved enhancers . Nat Biotechnol 18, 181–186 (2000). https://doi.org/10.1038/72635
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72635
This article is cited by
-
Klf5 Mediates Odontoblastic Differentiation through Regulating Dentin-Specific Extracellular Matrix Gene Expression during Mouse Tooth Development
Scientific Reports (2017)
-
Stem Cell Leukemia: how a TALented actor can go awry on the hematopoietic stage
Leukemia (2016)
-
Aberrant TAL1 activation is mediated by an interchromosomal interaction in human T-cell acute lymphoblastic leukemia
Leukemia (2014)
-
Transcriptional regulation of haematopoietic transcription factors
Stem Cell Research & Therapy (2011)
-
Unravelling the world of cis-regulatory elements
Medical & Biological Engineering & Computing (2007)