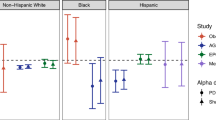
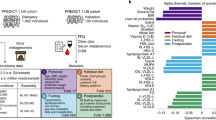
Abstract
Complex gene–environment interactions are considered important in the development of obesity1. The composition of the gut microbiota can determine the efficacy of energy harvest from food2,3,4 and changes in dietary composition have been associated with changes in the composition of gut microbial populations5,6. The capacity to explore microbiota composition was markedly improved by the development of metagenomic approaches7,8, which have already allowed production of the first human gut microbial gene catalogue9 and stratifying individuals by their gut genomic profile into different enterotypes10, but the analyses were carried out mainly in non-intervention settings. To investigate the temporal relationships between food intake, gut microbiota and metabolic and inflammatory phenotypes, we conducted diet-induced weight-loss and weight-stabilization interventions in a study sample of 38 obese and 11 overweight individuals. Here we report that individuals with reduced microbial gene richness (40%) present more pronounced dys-metabolism and low-grade inflammation, as observed concomitantly in the accompanying paper11. Dietary intervention improves low gene richness and clinical phenotypes, but seems to be less efficient for inflammation variables in individuals with lower gene richness. Low gene richness may therefore have predictive potential for the efficacy of intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mutch, D. M. & Clément, K. Unraveling the genetics of human obesity. PLoS Genet. 2, e188 (2006)
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl Acad. Sci. USA 101, 15718–15723 (2004)
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006)
Bäckhed, F., Manchester, J. K., Semenkovich, C. F. & Gordon, J. I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl Acad. Sci. USA 104, 979–984 (2007)
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006)
Duncan, S. H. et al. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl. Environ. Microbiol. 73, 1073–1078 (2007)
Riesenfeld, C. S., Schloss, P. D. & Handelsman, J. Metagenomics: genomic analysis of microbial communities. Annu. Rev. Genet. 38, 525–552 (2004)
National Research Council The New Science of Metagenomics: Revealing the Secrets of Our Microbial Planet (The National Academies Press, 2007)
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011)
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature http://dx.doi.org/10.1038/nature12506. (this issue)
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009)
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184 (2012)
Wasserman, S. & Faust, K. Social Network Analysis: Methods and Applications. (Cambridge Univ. Press, 1994)
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334, 105–108 (2011)
Ouchi, N., Parker, J. L., Lugus, J. J. & Walsh, K. Adipokines in inflammation and metabolic disease. Nature Rev. Immunol. 11, 85–97 (2011)
Shoelson, S. E., Lee, J. & Goldfine, A. B. Inflammation and insulin resistance. J. Clin. Invest. 116, 1793–1801 (2006)
Renehan, A. G., Tyson, M., Egger, M., Heller, R. F. & Zwahlen, M. Body-mass index and incidence of cancer: a systematic review and meta-analysis of prospective observational studies. Lancet 371, 569–578 (2008)
Rizkalla, S. W. et al. Differential effects of macronutrient content in 2 energy-restricted diets on cardiovascular risk factors and adipose tissue cell size in moderately obese individuals: a randomized controlled trial. Am. J. Clin. Nutr. 95, 49–63 (2012)
Bouché, C. et al. Five-week, low-glycemic index diet decreases total fat mass and improves plasma lipid profile in moderately overweight nondiabetic men. Diabetes Care 25, 822–828 (2002)
Tordjman, J. et al. Structural and inflammatory heterogeneity in subcutaneous adipose tissue: Relation with liver histopathology in morbid obesity. J. Hepatol. 56, 1152–1158 (2012)
Disse, E. et al. A lipid-parameter-based index for estimating insulin sensitivity and identifying insulin resistance in a healthy population. Diabetes Metab. 34, 457–463 (2008)
Antuna-Puente, B. et al. Evaluation of insulin sensitivity with a new lipid-based index in non-diabetic postmenopausal overweight and obese women before and after a weight loss intervention. Eur. J. Endocrinol. 161, 51–56 (2009)
Prat-Larquemin, L. et al. Adipose angiotensinogen secretion, blood pressure, and AGT M235T polymorphism in obese patients. Obes. Res. 12, 556–561 (2004)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B 57, 289–300 (1995)
Pons, N. et al. METEOR, a platform for quantitative metagenomic profiling of complex ecosystems. Journées Ouvertes en Biologie, Informatique et Mathématiques http://www.jobim2010.fr/sites/default/files/presentations/27Pons.pdf (2010)
Jiang, D., Huang, J. & Zhang, Y. The cross-validated AUC for MCP-logistic regression with high-dimensional data. Stat. Methods Med. Res http://dx.doi.org/10.1177/0962280211428385 (28 November 2011)
Shannon, C. E. A mathematical theory of communication. Bell Sys. Tech. J. 27, 379–423 (1995) 623–656 (1948)
Silverman, B. W. Density Estimation for Statistics and Data Analysis (Chapman and Hall, 1986)
R Development Core Team. R: A Language and Environment for Statistical Computinghttp://www.R-project.org (R Foundation for Statistical Computing, 2011)
Acknowledgements
We are grateful to O. Pedersen (Univ. Copenhagen) for helpful comments on this manuscript and to the MetaHIT consortium for providing the gene profiles of the Danish subjects used to test the ROC models in advance of publication and the DNA samples sequenced on the SOLiD platform for comparison with the Illumina platform used in the accompanying manuscript. We thank C. Baudoin, P. Ancel and V. Pelloux who contributed to the clinical investigation study; S. Fellahi and J.-P. Bastard for analyses of inflammatory markers; D. Bonnefont-Rousselot and R. Bittar for help with the analysis of plasma lipid profile. This work was supported by Agence Nationale de la Recherche (ANR MICRO-Obes, ANR, Nutra2sens, ANR-10-IAHU-05), the Metagenopolis grant ANR-11-DPBS-0001, KOT-Ceprodi (Florence Massiera), Danone Research (Damien Paineau) and the associations Fondacoeur, and Louis-Bonduelle. Additional funding came from the European Commission FP7 grant HEALTH-F4-2007-201052 and METACARDIS.
Author information
Authors and Affiliations
Consortia
Contributions
S.D.E., J.D. and K.C. designed the study; S.D.E., J.D., K.C. and P.R. managed the study; K.C. and S.R. designed the clinical research; S.R. and L.C.K. conducted the clinical research and clinical data management; A.C., S.R. and L.C.K. conducted clinical and dietary data analysis; S.G. gave dietary counselling to the patients and carried out analysis of dietary data; F.L. prepared the DNA for sequencing; S.K. managed DNA sequencing, which B.Q. and N.G. carried out; N.P. and J.-M.B. established the sequence analysis pipeline; A.C., J.-D.Z., E.P., N.P., E.L.C., M.A., J.-M.B., S.K. and S.D.E. carried out microbial data analysis; A.C., K.C., L.C.K. and S.D.E. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A list of authors and affiliations appears at the end of the paper.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-5, Supplementary Tables 1-6, 9-12, 14-15 and Supplementary Cluster Sheets. (PDF 5243 kb)
Supplementary Data
This file contains Supplementary Table 7. (XLS 30 kb)
Supplementary Data
This file contains Supplementary Table 8. (XLSX 22 kb)
Supplementary Data
This file contains Supplementary Table 13. (XLS 54 kb)
Rights and permissions
About this article
Cite this article
Cotillard, A., Kennedy, S., Kong, L. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588 (2013). https://doi.org/10.1038/nature12480
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12480
This article is cited by
-
The role of gut microbiota in human metabolism and inflammatory diseases: a focus on elderly individuals
Annals of Microbiology (2024)
-
Gut microbiome signature of metabolically healthy obese individuals according to anthropometric, metabolic and inflammatory parameters
Scientific Reports (2024)
-
Fewer culturable Lactobacillaceae species identified in faecal samples of pigs performing manipulative behaviour
Scientific Reports (2024)
-
Roxadustat alleviates the inflammatory status in patients receiving maintenance hemodialysis with erythropoiesis-stimulating agent resistance by increasing the short-chain fatty acids producing gut bacteria
European Journal of Medical Research (2023)
-
Metagenomic analysis reveals the relationship between intestinal protozoan parasites and the intestinal microecological balance in calves
Parasites & Vectors (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.