Abstract
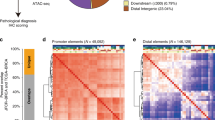
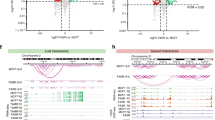
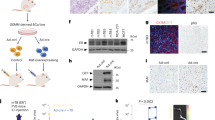
Oestrogen receptor-α (ER) is the defining and driving transcription factor in the majority of breast cancers and its target genes dictate cell growth and endocrine response, yet genomic understanding of ER function has been restricted to model systems1,2,3. Here we map genome-wide ER-binding events, by chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq), in primary breast cancers from patients with different clinical outcomes and in distant ER-positive metastases. We find that drug-resistant cancers still recruit ER to the chromatin, but that ER binding is a dynamic process, with the acquisition of unique ER-binding regions in tumours from patients that are likely to relapse. The acquired ER regulatory regions associated with poor clinical outcome observed in primary tumours reveal gene signatures that predict clinical outcome in ER-positive disease exclusively. We find that the differential ER-binding programme observed in tumours from patients with poor outcome is not due to the selection of a rare subpopulation of cells, but is due to the FOXA1-mediated reprogramming of ER binding on a rapid timescale. The parallel redistribution of ER and FOXA1 binding events in drug-resistant cellular contexts is supported by histological co-expression of ER and FOXA1 in metastatic samples. By establishing transcription-factor mapping in primary tumour material, we show that there is plasticity in ER-binding capacity, with distinct combinations of cis-regulatory elements linked with the different clinical outcomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Carroll, J. S. et al. Genome-wide analysis of estrogen receptor binding sites. Nature Genet. 38, 1289–1297 (2006)
Lin, C. Y. et al. Whole-genome cartography of estrogen receptor α binding sites. PLoS Genet. 3, e87 (2007)
Welboren, W. J. et al. ChIP-Seq of ERα and RNA polymerase II defines genes differentially responding to ligands. EMBO J. 28, 1418–1428 (2009)
Carroll, J. S. et al. Chromosome-wide mapping of estrogen receptor binding reveals long-range regulation requiring the forkhead protein FoxA1. Cell 122, 33–43 (2005)
Lupien, M. et al. FoxA1 translates epigenetic signatures into enhancer-driven lineage-specific transcription. Cell 132, 958–970 (2008)
Hurtado, A., Holmes, K. A., Ross-Innes, C. S., Schmidt, D. & Carroll, J. S. FOXA1 is a key determinant of estrogen receptor function and endocrine response. Nature Genet. 43, 27–33 (2011)
EBCTCG. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet 365, 1687–1717 (2005)
Kun, Y. et al. Classifying the estrogen receptor status of breast cancers by expression profiles reveals a poor prognosis subpopulation exhibiting high expression of the ERBB2 receptor. Hum. Mol. Genet. 12, 3245–3258 (2003)
Arpino, G. et al. Estrogen receptor-positive, progesterone receptor-negative breast cancer: association with growth factor receptor expression and tamoxifen resistance. J. Natl Cancer Inst. 97, 1254–1261 (2005)
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009)
Loi, S. et al. Definition of clinically distinct molecular subtypes in estrogen receptor-positive breast carcinomas through genomic grade. J. Clin. Oncol. 25, 1239–1246 (2007)
Knowlden, J. M. et al. Elevated levels of epidermal growth factor receptor/c-erbB2 heterodimers mediate an autocrine growth regulatory pathway in tamoxifen-resistant MCF-7 cells. Endocrinology 144, 1032–1044 (2003)
Lupien, M. et al. Growth factor stimulation induces a distinct ERα cistrome underlying breast cancer endocrine resistance. Genes Dev. 24, 2219–2227 (2010)
Nagashima, T. et al. Quantitative transcriptional control of ErbB receptor signaling undergoes graded to biphasic response for cell differentiation. J. Biol. Chem. 282, 4045–4056 (2007)
Schmidt, D. et al. ChIP-seq: using high-throughput sequencing to discover protein–DNA interactions. Methods 48, 240–248 (2009)
Gomm, J. J. et al. Isolation of pure populations of epithelial and myoepithelial cells from the normal human mammary gland using immunomagnetic separation with Dynabeads. Anal. Biochem. 226, 91–99 (1995)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Stark, R. & Brown, G. D. DiffBind: differential binding analysis of ChIP-seq peak data. Bioconductorhttp://bioconductor.org/packages/release/bioc/html/DiffBind.html.
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Pavesi, G. et al. MoD Tools: regulatory motif discovery in nucleotide sequences from co-regulated or homologous genes. Nucleic Acids Res. 34, W566–W570 (2006)
Wang, Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365, 671–679 (2005)
van de Vijver, M. J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002)
Sotiriou, C. et al. Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J. Natl. Cancer Inst. 98, 262–272 (2006)
Chin, K. et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10, 529–541 (2006)
Schmidt, M. et al. The humoral immune system has a key prognostic impact in node-negative breast cancer. Cancer Res. 68, 5405–5413 (2008)
Buffa, F. M. et al. microRNA associated progression pathways and potential therapeutic targets identified by integrated mRNA and microRNA expression profiling in breast cancer. Cancer Res. 71, 5635 (2011)
Naderi, A. et al. A gene-expression signature to predict survival in breast cancer across independent data sets. Oncogene 26, 1507–1516 (2007)
Hoadley, K. A. et al. EGFR associated expression profiles vary with breast tumor subtype. BMC Genomics 8, 258 (2007)
Bair, E. & Tibshirani, R. Semi-supervised methods to predict patient survival from gene expression data. PLoS Biol. 2, e108 (2004)
Acknowledgements
The authors would like to thank D. Schmidt for assistance with figures, J. Hadfield for Illumina sequencing, S. MacArthur, O. Rueda, S. Vowler, R. Russell and M. Wilson for technical and bioinformatics help. We thank J. Stingl and his laboratory for help with the normal mammary gland work. We would like to acknowledge the support of The University of Cambridge, Cancer Research UK and Hutchison Whampoa Limited. The authors would like to thank Imperial College Healthcare NHS Trust, Human Biomaterials Resource Centre (Tissue Bank). Tumour samples from Cambridge were obtained with support from NIHR Biomedical Research Centre and the Experimental Cancer Medicine Centre. C.S.R.-I. is supported by a Commonwealth Scholarship. O.G. is part funded by a grant awarded by the Ministry of Education of the Czech Republic (Project “Oncology” MSM 0021620808) and is also a recipient of a Translational Research Fellowship from the European Society of Medical Oncology. C.P. is funded by Cancer Research UK. J.S.C. is supported by an ERC starting grant and an EMBO Young investigator award.
Author information
Authors and Affiliations
Contributions
C.S.R.-I., R.S., C.C. and J.S.C. designed all experiments. Experimental work was conducted by C.S.R.-I. with help from K.A.H. Computational analysis was conducted by R.S. and A.E.T., with help from M.J.D. and G.D.B. All clinical samples, clinical information and help with sample processing was provided by C.C., C.P., S.-F.C., S.A., A.R.G., I.O.E. and O.G. Histological analysis was conducted by H.R.A. The manuscript was written by C.S.R.-I., R.S., C.C. and J.S.C. with assistance from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-16 with legends and additional references. (PDF 3818 kb)
Rights and permissions
About this article
Cite this article
Ross-Innes, C., Stark, R., Teschendorff, A. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012). https://doi.org/10.1038/nature10730
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10730
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.