Abstract
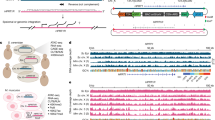
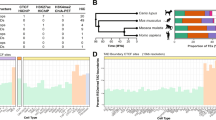
Genome-wide DNA rearrangements occur in many eukaryotes but are most exaggerated in ciliates, making them ideal model systems for epigenetic phenomena. During development of the somatic macronucleus, Oxytricha trifallax destroys 95% of its germ line, severely fragmenting its chromosomes, and then unscrambles hundreds of thousands of remaining fragments by permutation or inversion. Here we demonstrate that DNA or RNA templates can orchestrate these genome rearrangements in Oxytricha, supporting an epigenetic model for sequence-dependent comparison between germline and somatic genomes. A complete RNA cache of the maternal somatic genome may be available at a specific stage during development to provide a template for correct and precise DNA rearrangement. We show the existence of maternal RNA templates that could guide DNA assembly, and that disruption of specific RNA molecules disables rearrangement of the corresponding gene. Injection of artificial templates reprogrammes the DNA rearrangement pathway, suggesting that RNA molecules guide genome rearrangement.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
King, M. L., Messitt, T. J. & Mowry, K. L. Putting RNAs in the right place at the right time: RNA localization in the frog oocyte. Biol. Cell 97, 19–33 (2005)
Tadros, W. & Lipshitz, H. D. Setting the stage for development: mRNA translation and stability during oocyte maturation and egg activation in Drosophila . Dev. Dyn. 232, 593–608 (2005)
Tang, F. et al. Maternal microRNAs are essential for mouse zygotic development. Genes Dev. 21, 644–648 (2007)
Rassoulzadegan, M. et al. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006)
Herbert, A. & Rich, A. RNA processing and the evolution of eukaryotes. Nature Genet. 21, 265–269 (1999)
Lolle, S. J., Victor, J. L., Young, J. M. & Pruitt, R. E. Genome-wide non-mendelian inheritance of extra-genomic information in Arabidopsis . Nature 434, 505–509 (2005)
Peng, P., Chan, S. W., Shah, G. A. & Jacobsen, S. E. Plant genetics: increased outcrossing in hothead mutants. Nature 443, E8 (2006)
Lolle, S. J., Pruitt, R. E., Victor, J. L. & Young, J. M. Lolle et al. reply. Nature 443, E8–E9 (2006)
Storici, F., Bebenek, K., Kunkel, T. A., Gordenin, D. A. & Resnick, M. A. RNA-templated DNA repair. Nature 447, 338–341 (2007)
Blum, B., Bakalara, N. & Simpson, L. A model for RNA editing in kinetoplastid mitochondria: “guide” RNA molecules transcribed from maxicircle DNA provide the edited information. Cell 60, 189–198 (1990)
Maxwell, E. S. & Fournier, M. J. The small nucleolar RNAs. Annu. Rev. Biochem. 64, 897–934 (1995)
Prescott, D. M. The DNA of ciliated protozoa. Microbiol. Rev. 58, 233–267 (1994)
Chalker, D. L. & Yao, M. C. Non-mendelian, heritable blocks to DNA rearrangement are induced by loading the somatic nucleus of Tetrahymena thermophila with germ line-limited DNA. Mol. Cell. Biol. 16, 3658–3667 (1996)
Duharcourt, S., Keller, A. M. & Meyer, E. Homology-dependent maternal inhibition of developmental excision of internal eliminated sequences in Paramecium tetraurelia . Mol. Cell. Biol. 18, 7075–7085 (1998)
Mochizuki, K., Fine, N. A., Fujisawa, T. & Gorovsky, M. A. Analysis of a piwi-related gene implicates small RNAs in genome rearrangement in Tetrahymena . Cell 110, 689–699 (2002)
Prescott, D. M. & DuBois, M. L. Internal eliminated segments (IESs) of Oxytrichidae. J. Eukaryot. Microbiol. 43, 432–441 (1996)
Mayer, K. M. & Forney, J. D. A mutation in the flanking 5′-TA-3′ dinucleotide prevents excision of an internal eliminated sequence from the Paramecium tetraurelia genome. Genetics 151, 597–604 (1999)
Landweber, L. F., Kuo, T. C. & Curtis, E. A. Evolution and assembly of an extremely scrambled gene. Proc. Natl Acad. Sci. USA 97, 3298–3303 (2000)
Prescott, D. M., Ehrenfeucht, A. & Rozenberg, G. Template-guided recombination for IES elimination and unscrambling of genes in stichotrichous ciliates. J. Theor. Biol. 222, 323–330 (2003)
Angeleska, A., Jonoska, N., Saito, M. & Landweber, L. F. RNA-guided DNA assembly. J. Theor. Biol. 248, 706–720 (2007)
Prescott, J. D., DuBois, M. L. & Prescott, D. M. Evolution of the scrambled germline gene encoding α-telomere binding protein in three hypotrichous ciliates. Chromosoma 107, 293–303 (1998)
Hoffman, D. C. & Prescott, D. M. Evolution of internal eliminated segments and scrambling in the micronucleargene encoding DNA polymerase α in two Oxytricha species Oxytricha novo is extremely scrambled. Nucleic Acids Res. 25, 1883–1889 (1997)
Möllenbeck, M. et al. The pathway to detangle a scrambled gene. PLoS Biol. (submitted)
Horvath, M. P., Schweiker, V. L., Bevilacqua, J. M., Ruggles, J. A. & Schultz, S. C. Crystal structure of the Oxytricha nova telomere end binding protein complexed with single strand DNA. Cell 95, 963–974 (1998)
Yao, M. C., Fuller, P. & Xi, X. Programmed DNA deletion as an RNA-guided system of genome defense. Science 300, 1581–1584 (2003)
Paques, F. & Haber, J. E. Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae . Microbiol. Mol. Biol. Rev. 63, 349–404 (1999)
Derr, L. K. & Strathern, J. N. A role for reverse transcripts in gene conversion. Nature 361, 170–173 (1993)
Moore, J. K. & Haber, J. E. Capture of retrotransposon DNA at the sites of chromosomal double-strand breaks. Nature 383, 644–646 (1996)
Nevo-Caspi, Y. & Kupiec, M. cDNA-mediated Ty recombination can take place in the absence of plus-strand cDNA synthesis, but not in the absence of the integrase protein. Curr. Genet. 32, 32–40 (1997)
Teng, S. C., Kim, B. & Gabriel, A. Retrotransposon reverse-transcriptase-mediated repair of chromosomal breaks. Nature 383, 641–644 (1996)
Chang, W.-J. et al. Intron evolution and information processing in the DNA polymerase alpha gene in spirotrichous ciliates: a hypothesis for interconversion between DNA and RNA deletion. Biol. Direct 2, 6 (2007)
Galvani, A. & Sperling, L. RNA interference by feeding in Paramecium . Trends Genet. 18, 11–12 (2002)
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004)
Paschka, A. G. et al. The use of RNAi to analyze gene function in spirotrichous ciliates. Eur. J. Protistol. 39, 449–454 (2003)
Williams, K., Doak, T. G. & Herrick, G. Developmental precise excision of Oxytricha trifallax telomere-bearing elements and formation of circles closed by a copy of the flanking target duplication. EMBO J. 12, 4593–4601 (1993)
Chang, W.-J., Stover, N. A., Addis, V. M. & Landweber, L. F. A micronuclear locus containing three protein-coding genes remains linked during macronuclear development in the spirotrichous ciliate Holosticha . Protist 155, 245–255 (2004)
Laughlin, T. J., Henry, J. M., Phares, E. F., Long, M. V. & Olins, D. E. Methods for the large-scale cultivation of an Oxytricha (Ciliophora: Hypotrichida). J. Eukaryot. Microbiol. 30, 63–64 (1983)
Acknowledgements
This work was supported by awards from the NSF and NIH to L.F.L. and the SEAS senior thesis research fund to V.V. We thank J. Wang for technical assistance and all members of the laboratory for discussion.
Author Contributions M.N., V.V., Y.Z., T.G.D. and L.F.L. designed experiments; M.N., V.V., Y.Z. and K.S. performed the experiments; T.G.D. provided cells; M.N., V.V., Y.Z. and L.F.L. analysed the data; and M.N., V.V., Y.Z. and L.F.L. wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
TITLE
The file contains Supplementary Figures 1-5 with Legends, Primer sequences and experimental notes; sequences used for RNAi and a guide to Supplementary FASTA Data files with additional sequence discussion. (PDF 2282 kb)
Supplementary Data
This folder contains Supplementary Data FASTA files. (ZIP 10 kb)
Rights and permissions
About this article
Cite this article
Nowacki, M., Vijayan, V., Zhou, Y. et al. RNA-mediated epigenetic programming of a genome-rearrangement pathway. Nature 451, 153–158 (2008). https://doi.org/10.1038/nature06452
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06452
This article is cited by
-
Application of RNA interference and protein localization to investigate housekeeping and developmentally regulated genes in the emerging model protozoan Paramecium caudatum
Communications Biology (2024)
-
Many Paths to Anticipatory Behavior: Anticipatory Model Acquisition Across Phylogenetic and Ontogenetic Timescales
Biological Theory (2023)
-
Timing and characteristics of nuclear events during conjugation and genomic exclusion in Paramecium multimicronucleatum
Marine Life Science & Technology (2022)
-
RNA: a double-edged sword in genome maintenance
Nature Reviews Genetics (2020)
-
Precise gene replacement in plants through CRISPR/Cas genome editing technology: current status and future perspectives
aBIOTECH (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.