Abstract
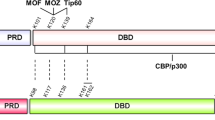
p53 is a tumour suppressor that regulates the cellular response to genotoxic stresses. p53 is a short-lived protein and its activity is regulated mostly by stabilization via different post-translational modifications. Here we report a novel mechanism of p53 regulation through lysine methylation by Set9 methyltransferase. Set9 specifically methylates p53 at one residue within the carboxyl-terminus regulatory region. Methylated p53 is restricted to the nucleus and the modification positively affects its stability. Set9 regulates the expression of p53 target genes in a manner dependent on the p53-methylation site. The crystal structure of a ternary complex of Set9 with a p53 peptide and the cofactor product S-adenosyl-l-homocysteine (AdoHcy) provides the molecular basis for recognition of p53 by this lysine methyltransferase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001)
Wang, H. et al. Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol. Cell 8, 1207–1217 (2001)
Nishioka, K. et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev. 16, 479–489 (2002)
Appella, E. & Anderson, C. W. Post-translational modifications and activation of p53 by genotoxic stresses. Eur. J. Biochem. 268, 2764–2772 (2001)
Shieh, S. Y., Ikeda, M., Taya, Y. & Prives, C. DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell 91, 325–334 (1997)
Momand, J., Wu, H. H. & Dasgupta, G. MDM2—master regulator of the p53 tumor suppressor protein. Gene 242, 15–29 (2000)
Brooks, C. L. & Gu, W. Ubiquitination, phosphorylation and acetylation: the molecular basis for p53 regulation. Curr. Opin. Cell Biol. 15, 164–171 (2003)
Li, M., Luo, J., Brooks, C. L. & Gu, W. Acetylation of p53 inhibits its ubiquitination by Mdm2. J. Biol. Chem. 277, 50607–50611 (2002)
Luo, J. et al. Acetylation of p53 augments its site-specific DNA binding both in vitro and in vivo. Proc. Natl Acad. Sci. USA 101, 2259–2264 (2004)
Gu, W. & Roeder, R. G. Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 90, 595–606 (1997)
Barlev, N. A. et al. Acetylation of p53 activates transcription through recruitment of coactivators/histone acetyltransferases. Mol. Cell 8, 1243–1254 (2001)
Gu, W., Luo, J., Brooks, C. L., Nikolaev, A. Y. & Li, M. Dynamics of the p53 acetylation pathway. Nov. Found. Symp. 259, 197–205 (2004)
Gostissa, M. et al. Activation of p53 by conjugation to the ubiquitin-like protein SUMO-1. Embo J. 18, 6462–6471 (1999)
Kwek, S. S., Derry, J., Tyner, A. L., Shen, Z. & Gudkov, A. V. Functional analysis and intracellular localization of p53 modified by SUMO-1. Oncogene 20, 2587–2599 (2001)
Rodriguez, M. S. et al. SUMO-1 modification activates the transcriptional response of p53. Embo J. 18, 6455–6461 (1999)
Kouskouti, A., Scheer, E., Staub, A., Tora, L. & Talianidis, I. Gene-specific modulation of TAF10 function by SET9-mediated methylation. Mol. Cell 14, 175–182 (2004)
Xiao, B. et al. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature 421, 652–656 (2003)
Grand, R. J. et al. The high levels of p53 present in adenovirus early region 1-transformed human cells do not cause up-regulation of MDM2 expression. Virology 210, 323–334 (1995)
Tachibana, M., Sugimoto, K., Fukushima, T. & Shinkai, Y. SET domain-containing protein, G9a, is a novel lysine-preferring mammalian histone methyltransferase with hyperactivity and specific selectivity to lysines 9 and 27 of histone H3. J. Biol. Chem. 276, 25309–25317 (2001)
Otwinowski, Z. & Minor, W. in Data Collection and Processing (eds Sawyer, L., Isaacs, N. & Bailey, S.) 552–562 (SERC Daresbury Laboratory, Warrington, 1993)
CCP4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Jones, T. A., Zhou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Acknowledgements
We thank L. Vales for comments on the manuscript. We also thank S. L. Berger and members of the Reinberg laboratory, especially A. Kuzmichev and K. Sarma for discussions. We also thank E. White for discussions and P. Chumakov for the gift of the lentivirus LSL–GFP vector, and A. Ivanov for generation of the siRNA-Set9 U2OS cell line. We thank S. Martin for assistance with binding experiments. The GRASP centre at Tufts is acknowledged for providing excellent technical support. This work was supported by a grant from the NIH and the Howard Hughes Medical Institute (D.R.). N.A.B. acknowledges support from the Charlton Award. S.J.G. acknowledges support from the Medical Research Council and from the Association for International Cancer Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Set 9 mono-methylates p53. p53 10 mer peptide was methylated by Set9 (52-366aa). (PDF 174 kb)
Supplementary Figure S2
Overlay of cartoon representations of Set9 ternary structures with p53 substrate. (PDF 200 kb)
Supplementary Figure S3
Set 9 regulates expression of BAX and MDM2 genes. (PDF 36 kb)
Supplementary Figure S4
Set9 mediated activation of p21 gene expression requires p53. H1299 cells were stably transfected with Flag-Set9 wt or control vector. (PDF 42 kb)
Supplementary Figure S5
Set9 facilitates apoptosis in p53-dependent manner. (PDF 16 kb)
Supplementary Table S1
Data and refinement statistics for the ternary complex of SET9 with p53(369-374). (PDF 53 kb)
Rights and permissions
About this article
Cite this article
Chuikov, S., Kurash, J., Wilson, J. et al. Regulation of p53 activity through lysine methylation. Nature 432, 353–360 (2004). https://doi.org/10.1038/nature03117
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03117
This article is cited by
-
Set7/9 aggravates ischemic brain injury via enhancing glutamine metabolism in a blocking Sirt5 manner
Cell Death & Differentiation (2024)
-
Lysine methylation promotes NFAT5 activation and determines temozolomide efficacy in glioblastoma
Nature Communications (2023)
-
Mettl13 protects against cardiac contractile dysfunction by negatively regulating C-Cbl-mediated ubiquitination of SERCA2a in ischemic heart failure
Science China Life Sciences (2023)
-
Regulation of Keap1-Nrf2 axis in temporal lobe epilepsy—hippocampal sclerosis patients may limit the seizure outcomes
Neurological Sciences (2023)
-
Unconventional protein post-translational modifications: the helmsmen in breast cancer
Cell & Bioscience (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.