Abstract
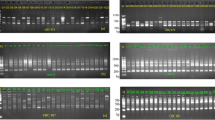
Genomic regions differentiating Quercus petraea and Quercus robur were detected by screening 2800 PCR amplification products using random primers on 22 trees of each species sampled in 11 natural populations. Only two per cent of the amplified fragments exhibited significant frequency differences between the two species and none of them was specific to a species. The nucleotide divergence between the two species estimated with RAPD data was 0.5 per cent in the overall genome and increased to 3.3 per cent in the discriminant regions. Twenty-three informative fragments were cloned and partially sequenced. New primers were derived from these sequences to obtain Sequence Characterized Amplified Region (SCAR) fragments. Southern blot experiments indicated that the SCARs were generally in low copy number in the genome. A search for similarity between SCAR sequences and sequences contained in data banks revealed that three of them corresponded to known DNA sequences.
Similar content being viewed by others
Article PDF
References
Altschul, S F, Gish, W, Miller, W, Myers, E W, and Lipman, D J. 1990. Basic local alignment search tool. J Mol Biol, 215, 403–410.
Bacilieri, R, Ducousso, A, Petit, R J, and Kremer, A. 1996. Mating system and asymmetric hybridization flow in a mixed stand of European oaks. Evolution, 50, 900–908.
Bodénès, C, Laigret, F, and Kremer, A. 1996. Inheritance and molecular variations of PCR-SSCP fragments in Pedunculate oak (Quercus robur L.). Theor Appl Genet, (in press).
Chalmers, K J, Waugh, R, Sprent, J I, Simons, A J, and Powell, W. 1992. Detection of genetic variation between and within populations of Gliricidia sepmium and G. maculata using RAPD markers. Heredity, 69, 465–472.
Chong, D K X, Yang, R E, and Yeh, F C. 1994. Nucleotide divergence between populations of trembling aspen (Populus tremuloides) estimated with RAPDs. Curr Genet, 26, 374–376.
Clark, A G, and Lanigan, C M S. 1993. Prospects for estimating nucleotide divergence with RAPDs. Mol Biol Evol, 10, 1096–1111.
Crawford, D J. 1985. Electrophoretic data and plant speciation. Syst Bot, 10, 405–416.
Denhart, D T. 1966. A membrane filter technique for the detection of complementary DNA. Biochem Biophys Res Commun, 23, 641–646.
Dupouey, J L, and Badeau, V. 1993. Morphological variability of oaks (Quercus robur L., Quercus petraea (Matt.) Liebl., Quercus pubescens Willd.) in northeastern France: preliminary results. Sci For, 50, 35s–40s.
Favre, J M, and Brown, S. 1996. A flow cytometric evaluation of the nuclear DNA content and GC per cent in genomes of European oak species. Ann Sci For, 53, 915–917.
Flavell, R B, Benett, M D, Smith, J B, and Smith, D B. 1974. Genome size and the proportion of repeated nucleotide sequence DNA in plants. Biochem Genet, 12, 257–269.
Grattapaglia, D, and Sederoff, R. 1994. Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics, 137, 1121–1137.
Harada, K, Kinoshita, A, Shukor, N A A, Tachida, H, and Yamayki, T. 1995. Genetic variation estimated in three Shorea species by the RAPD analysis. Jap J Genet, 69, 713–718.
Kendall, M G, and Stuart, A. 1977. The Advanced Theory of Statistics, Vol. 1, 4th edn. Griffin, London.
Klein-Lankhorst, R M, Vermunt, A, Weide, R, Lihar-Ska, T, and Zabel, P. 1991. Isolation of molecular markers for tomato (L. esculentum) using random amplified polymorphic DNA (RAPD). Theor Appl Genet, 83, 108–114.
Kleinschmit, J R G, Bacilieri, R, Kremer, A, and Roloff, A. 1995. Comparison of morphological traits of pedunculate oak (Q. robur L.) and sessile oak (Q. petraea (Matt.) Liebl.). Silvae Genet, 44, 256–269.
Kremer, A, and Petit, R J. 1993. Gene diversity in natural populations of oak species. Ann Sci For, 50, 186s–202s.
Krieber, M, and Rose, M R. 1986. Molecular aspects of the species barrier. Ann Rev Ecol Syst, 17, 465–485.
Lebart, L, Morineau, A, and Tabart, N. 1984. Multivariate Descriptive Stastitical Methods: Correspondance Analysis and Related Techniques for Large Matrices. J. Wiley and Sons, New York.
Lisitsyn, N, Lisitsys, N, and Wigler, M. 1993. Cloning the differences between two complex genomes. Science, 259, 946–951.
Moreau, F, Kleinschmit, J, and Kremer, A. 1994. Molecular differentiation between Q. petraea and Q. robur assessed by random amplified DNA fragments. Forest Genet, 1, 51–64.
Müller-Starck, G, Herzog, S, and Hattemer, H H. 1993. Intra and inter population genetic variation in juvenile populations of Quercus robur L. and Quercus petraea Liebl. Ann Sci For, 50, 233s–244s.
Nei, M. 1987. Molecular Evolutionary Genetics. Columbia University Press, New York.
Nei, M, and Li, W H. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA, 76, 5269–5273.
Nei, M, and Miller, J C. 1990. A simple method for estimating average number of nucleotide substitutions within and between populations from restriction data. Genetics, 125, 873–879.
N'Goran, J A, Laurent, V, Risterucci, A M, and Lanaud, C. 1994. Comparative genetic diversity studies of Theobroma cacao L. using RFLP and RAPD markers. Heredity, 73, 589–597.
Paran, I, and Michelmore, R W. 1993. Development of reliable PCR-based markers linked to downy mildew resistance genes in lettuce. Theor Appl Genet, 85, 985–993.
Peltier, D, Farcy, E, Dulieu, H, and Berville, A. 1994. Origin, distribution and mapping of RAPD markers from wild Petunia species in Petunia hybrida Hort lines. Theor Appl Genet, 88, 637–645.
Petit, R J, Wagner, D B, and Kremer, A. 1993. Ribosomal DNA gene and chloroplast DNA polymorphisms in a mixed stand of Quercus robur and Quercus petraea. Ann Sci For, 50, 41s–47s.
Plomion, C, Barhman, N, Durel, C E, and O'Malley, D M. 1995. Genomic mapping in Pinus pinaster (maritime pine) using RAPD and protein markers. Heredity, 74, 661–668.
Reed, T C, and Mann, D A. 1985. Rapid transfer of DNA from agarose gels to nylon membranes. Nucl Acids Res, 13, 7207–7221.
Reiter, R S, Williams, J G K, Feldmann, K A, Rafal-Ski, J A, Tingey, S V, and Scolnik, P A. 1992. Global and local genome mapping in Arabidopsis thaliana by using recombinant inbred lines and random amplified polymorphic DNAs. Proc Natl Acad Sci USA, 89, 1477–1481.
Rose, M R, and Doolittle, W F. 1983. Molecular biological mechanisms of speciation. Science, 200, 157–162.
Rychlik, W, Spencer, W J, and Rhoads, R E. 1990. Optimization of the annealing temperature for DNA amplification in vitro. Nucl Acids Res, 18, 6409–6412.
Saghai-Maroof, M A, Soliman, K M, Jorgensen, R A, and Allard, R W. 1984. Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA, 81, 8014–8018.
Sheldon, F H. 1995. Advances in the theory and practice of DNA hybridization as a systematic method. In: Schierwater, B., Streit, B., Wagner, G. P. and DeSalle, R. (eds) Molecular Ecology and Evolution: Approaches and Applications, pp. 285–297. Birkhäuser Verlag, Basel.
Southern, E M. 1976. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol, 98, 503–517.
Steinhoff, S. 1993. Results of species hybridization with Quercus robur L. and Quercus petraea (Matt.) Liebl. Ann Sci For, 50, 137s–143s.
Thormann, C E, Ferreira, M E, Camargo, L E A, Tivang, J G, and Osborn, T C. 1994. Comparison of RFLP and RAPD markers to estimating genetic relationships within and among cruciferous species. Theor Appl Genet, 88, 973–980.
Wilbur, W J, and Lipman, D J. 1983. Rapid similarity searches of nucleic acid and protein data banks. Proc Natl Acad Sci USA, 80, 726–730.
Wilkie, S E, Isaac, P G, and Slater, R. 1993. Random amplified polymorphic DNA (RAPD) markers for genetic analysis in Allium. Theor Appl Genet, 86, 497–504.
Williams, J G K, Kubelik, A R, Livak, K J, Rafalski, J A, and Tingey, S V. 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl Acids Res, 18, 6531–6535.
Williams, J G K, Hanafey, M K, Rafalski, J A, and Tingey, S V. 1993. Genetic analysis using random amplified polymorphic DNA markers. Methods Enzymol, 318, 705–740.
Xu, H, Wilson, D J, Arulsekar, S, and Bakalinsky, A T. 1995. Sequence specific polymerase chain-reaction markers derived from randomly amplified polymorphic DNA markers for fingerprinting grape (Vitis) root-stocks. J Amer Soc Hort Sci, 120, 714–720.
Zanetto, A, Roussel, G, and Kremer, A. 1994. Geographic variation of inter-specific differentiation between Quercus robur L. and Quercus petraea (Matt.) Liebl. Forest Genet, 1, 111–123.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bodénès, C., Joandet, S., Laigret, F. et al. Detection of genomic regions differentiating two closely related oak species Quercus petraea (Matt.) Liebl. and Quercus robur L.. Heredity 78, 433–444 (1997). https://doi.org/10.1038/hdy.1997.67
Received:
Issue Date:
DOI: https://doi.org/10.1038/hdy.1997.67
Keywords
This article is cited by
-
Effect of hybridization on the morphological differentiation of the red oaks Quercus acutifolia and Quercus grahamii (Fagaceae)
Plant Systematics and Evolution (2021)
-
Species-specific alleles at a β-tubulin gene show significant associations with leaf morphological variation within Quercus petraea and Q. robur populations
Tree Genetics & Genomes (2016)
-
Genotypic analysis and population structure of Lebanon oak (Quercus libani G. Olivier) with molecular markers
Tree Genetics & Genomes (2015)
-
Development and Characterization of Genomic and Gene-Based Microsatellite Markers in North American Red Oak Species
Plant Molecular Biology Reporter (2013)
-
Nucleotide sequence, structural organization and length heterogeneity of ribosomal DNA intergenic spacer in Quercus petraea (Matt.) Liebl. and Q. robur L.
Molecular Genetics and Genomics (2009)