Abstract
The aims of this study were to use computerized morphometry in order to differentiate between the degree of dysplasia and to predict progression to invasive adenocarcinoma in Barrett's esophagus (BE). Biopsies from 97 patients with BE graded by a consensus forum of expert gastrointestinal pathologists were available for morphometrical analysis. The study group included 36 biopsies negative for dysplasia (ND), none of which progressed to carcinoma; 16 indefinite for dysplasia (IND) and 21 low-grade dysplasia (LGD), of which three progressed in each group and 24 high-grade dysplasia (HGD), of which 15 progressed to invasive carcinoma. Computerized morphometry was used for measuring indices of size, shape, texture, symmetry and architectural distribution of the epithelial nuclei. Low-grade dysplasia was best differentiated from the ND group by nuclear pseudostratification (P=0.036), pleomorphism (P<0.01), and chromatin texture (margination, P<0.01) and from the HGD group by nuclear area (P<0.01), pleomorphism (P<0.01), chromatin texture (margination, P<0.01), symmetry (P<0.01), and orientation (P=0.027). These results were validated on a new set of cases (n=55) using a neural network model, resulting in an accuracy of 89% for differentiating between the ND and LGD groups and 86% for differentiating between the LGD and HGD groups. Within the HGD group, univariate significant predictors of the progression interval to carcinoma were: indices of nuclear texture (heterogeneity: P=0.0019, s.d.-OD: P=0.005) and orientation: P=0.022. Nuclear texture (heterogeneity) was the only independent predictor of progression (P=0.004, hazard=11.54) by Cox's multivariate test. This study proposes that computerized morphometry is a valid tool for determining the grade of dysplasia in BE. Moreover, histomorphometric quantification of nuclear texture is a powerful tool for predicting progression to invasive adenocarcinoma in patients with HGD.
Similar content being viewed by others
Main
Epithelial dysplasia in Barrett's esophagus (BE) is the precursor of invasive adenocarcinoma and provides the basis for endoscopic surveillance and treatment decisions.1, 2, 3, 4 A significant correlation exists between the degree of dysplasia and the time of progression to invasive adenocarcinoma.5, 6, 7, 8 However, considerable interobserver variability in the grading of dysplasia has been encountered.5, 6, 7, 9, 10, 11, 12, 13, 14 Molecular studies exploring genetic aberrations, oncogene and tumor suppressor gene expression, ploidy, apoptosis, and angiogenesis have all contributed greatly to the understanding of the biology of BE-related neoplasia.14, 15, 16, 17, 18 Despite these advances, no single molecular marker has emerged that has proven to reliably predict who will and who will not develop cancer in the setting of BE.14
The current grading scheme of dysplasia is primarily based on the subjective evaluation of nuclear characteristics such as pleomorphism, enlargement, and hyperchromasia and nuclear spatial features such as orientation and stratification.19 The utility of computerized morphometry in the grading of Barrett's dysplasia has been examined in a few studies. Baak et al20 detected a positive correlation between certain morphometrical parameters, the proliferation index and the histological grading of dysplasia. Other studies also combined quantitative morphometry with ki-67 and p53 immunohistochemical markers and found a significant decrease in the interobserver variability in grading of esophageal dysplasia.21, 22 The role of computerized morphometry in predicting the time to progression from dysplasia to invasive carcinoma has not yet been evaluated.
The aims of this study were to determine the utility of computerized morphometry as an objective method for grading dysplasia and to evaluate the role of morphometry in predicting progression to adenocarcinoma in BE patients. In this study, we concentrated on aspects of nuclear structure such as size, shape, texture, as well as novel quantitative indices of nuclear pleomorphism and symmetry. These measurements were further complemented by a more general analysis, at the architectural level, of epithelial cell organization, which included indices of orientation, pseudostratification, and nuclear crowding.
Materials and methods
Patients and Tissues
This study included tissue sections from two sources as follows:
-
a)
A set of 138 patients described in a previous multicentric retrospective study,5, 9 which focused on interobserver variability in BE-associated dysplasia. All biopsies in the original study were reviewed and graded for dysplasia by 12 expert gastrointestinal pathologists after reaching a consensus agreement upon the grade of dysplasia in all cases. Out of these original 138 cases, only 97 were technically suitable for morphometric analysis, based on the quality of hematoxylin and eosin staining. Of these 97 BE cases, 36 were negative for dysplasia (ND), 16 were diagnosed as indefinite for dysplasia (IND), 21 low-grade dysplasia (LGD), and 24 with high-grade dysplasia (HGD). These cases were used as a gold standard to develop our predictive model.
-
b)
In order to validate our model for evaluation, an additional set of biopsies from 55 BE patients (ND=21; LGD=15; HGD=19) was obtained from the archives of the Department of Pathology at Rhode Island Hospital. The diagnoses were assessed and confirmed by three of the authors (MR, JW, ES), none of who was part of the original consensus forum.
Computerized Morphometry
Hematoxylin and eosin-stained histologic images were scanned and digitized using a computerized image analysis system comprising a high-resolution digital camera (MicroPublisher 3.3 RTV, QIMAGING, Burnaby, BC, Canada) attached to a microscope (Olympus BX41, Japan) and to an IBM-compatible computer (Pentium R 4 CPU 3.2 GHz, 0.99GB RAM). Images were displayed at a pixel resolution of 1024 × 768 pixels (spatial resolution=0.11 microns per pixel). The Image Pro Plus version 5.1 software (MediaCybernetics, MD, USA) was used for image segmentation, threshold establishment, and nuclear measurements. Nuclear contours were automatically highlighted. Manual corrections were performed as needed.
For each biopsy, the entire surface epithelium was digitally scanned at a magnification of × 600. The scanned images were then reviewed by MR and ES. All areas that appeared dysplastic or suspicious for dysplasia were sampled for morphometrical analysis. In the normal-appearing biopsies, representative fields were sampled.
Overall, 19 780 nuclei from the metaplastic and/or dysplastic surface epithelium were measured by computerized morphometry. For each nucleus, multiple morphometric indices were recorded, covering a multitude of dimensions, including size (area, diameters), shape (ellipticity, contour regularity index, roundness, fractal dimension), and nuclear chromatin texture. The latter included multiple textural indices quantifying the variation in pixel density of the nuclei. Examples of such indices include chromatin margination and heterogeneity.23 Margination is derived from the second moment of inertia of an object and it represents the relative distribution of the optical density between the center and the edges of a nucleus. High margination indicates that most of the intensity is along the edges while low margination indicates that most of the intensity is near the geometric center of the nucleus.
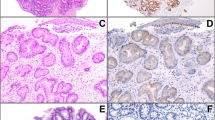
Heterogeneity is another morphometric descriptor of the nuclear chromatin texture and it is defined as the fraction of pixels that deviate more than 10% from the average optical density of the nucleus. Larger values of heterogeneity are obtained from more irregular chromatin patterns (Figure 1a). All the above indices are provided by the Image Pro Plus program for the quantitative analysis of patterns and objects in the images.
Morphometric concepts applied for quantitative evaluation of chromatin texture, nuclear symmetry, and orientation. (a) Heterogeneity: the fraction of pixels that deviate more than 10% from the average optical density of the nuclear chromatin. Two nuclei are displayed, each showing a different optical density (gray levels) distribution pattern. The left nucleus is more homogenous and therefore has a lower heterogeneity as compared to the right nucleus. (b) Symmetry: the steps applied for evaluating nuclear symmetry are shown. The segmented nuclei are oriented along their vertical axes and divided in two fragments along their horizontal axes (passing through their geometric centers of gravity). Thus, indices of size, shape, and textural symmetry are calculated as described in the Materials and methods section. (c) Orientation: degree of alignment of the nuclei along their longest axes was evaluated, as shown in the figure. Microscopic images and their frequency distributions were examined before and after applying a Fast Fourier Transform (FFT) algorithm. High-grade dysplasia (HGD) showed less-oriented nuclei resulting in rounder frequency distributions as opposed to low-grade dysplasia (LGD) or no-dysplasia (ND) cases showing more oriented nuclei providing more elliptic frequency distributions.
Using the predesigned variables, additional morphometric indices were computed in order to quantitatively assess nuclear pleomorphism and nuclear symmetry. Nuclear pleomorphism (variation in the nuclear size, shape, and texture) was determined by calculating the standard deviations of all measured variables, ensuring a multidimensional evaluation of the concept of nuclear variability. A similar approach was already used in morphometric studies measuring tumor pleomorphism in tissues other than BE.24
Novel indices of nuclear symmetry were also designed. In order to measure nuclear symmetry, the nuclei were sorted by their longest axis and a Macro algorithm obtained from the Media Cybernetics website was applied for splitting each nucleus along its shortest axis (reflectional symmetry), passing through a calculated digital center of gravity called centroid. Measurements were repeated for each one of the two resulting nuclear segments. Then, a ratio of symmetry was calculated for each variable, by dividing the smaller to the larger value of the two halves of the nucleus (Figure 1b). Lower indices are obtained from less-symmetric nuclei.
In order to enhance the observer's confidence regarding the perception of nuclear orientation, images were transformed from their space into their frequency domains, using a two-dimensional (2D) Fast Fourier Transform (FFT) algorithm. Digitized images are composed of 2D arrays of pixels, each having a light intensity value (gray level) ranging from 0 (black) through different shades of gray to 255 (white). Every row or column in this 2D array represents a signal within a range of coordinates defining a so-called space domain. Fourier Transform is an important image processing tool, which is used to decompose such 2D arrays (signals) into more basic components composed of sine and cosine functions.25 FFT is an efficient computer algorithm for calculating the basic frequencies of these sinusoidal functions. The graphical output of such transformation is called the Fourier frequency domain of the image. In this Fourier frequency domain, each point represents a particular frequency contained in the spatial domain image. The 2D Fourier frequency plots are frequently used in image analysis in purpose to analyze pattern periodicity and orientation. The frequency plots obtained from the Fourier transformation of the BE histological images were displayed. A well-oriented nuclear arrangement provided an elliptic frequency distribution. A disoriented nuclear pattern generated a round distribution of gray level frequencies. Different grades of orientation exhibited a spectrum of frequency patterns in between elliptic and round distributions. A semiquantitative score of nuclear orientation was then created, ranging from 1 (good) to 3 (poor orientation), reflecting the progressive loss of nuclear orientation (Figure 1c).
Nuclear pseudostratification was assessed by a semiquantitative score ranging from +1 where two nuclear layers were seen in up to 25% of the epithelium examined, +2 where two nuclear layers were seen in more than 25% or three or more layers were present in less than 25% of the epithelium, and +3 were three or more layers are seen in more than 25% of the epithelium. A similar semiquantitative score was applied for the nuclear crowding ranging from +1 (low crowding) to +3 (high crowding) of the surface metaplastic or dysplastic epithelium.
Statistical Analysis
For standardization of the variances within the groups, measurement values were normalized by dividing them by their corresponding variable averages.26, 27
Normality of the groups was tested using the Kolmogorov–Smirnov test. Comparison between multiple continuous groups was done using the one-way ANOVA test, followed by the Bonferroni post hoc test for detecting significant differences between individual subgroups. Reproducibility (evaluating the interoperator variability) was assessed by computing the R Pearson's coefficient of correlation between two repetitive measurements, done by two different operators (ES, AB).
In order to handle data complexity, technically related or unexplained variation, a two-layered feed forward back-propagation NNET algorithm28 was trained using the statistically predictive variables that were obtained by analyzing the first 97 cases. This NNET model was aimed to differentiate BE from LGD and LGD from HGD. For each classification task, the NNET algorithm contained an input neuron (the morphometric results), two hidden neurons (three log-sigmoid transfer functions in the first hidden neuron and one log-sigmoid function in the second hidden neuron) and an output neuron (the grade of dysplasia). In order to maintain the model ability to generalize (by avoiding overtraining), training was stopped after 100 iterations. Validation of the trained NNET model was subsequently done by testing the accuracy of its classification on a second group of images obtained from 55 new cases collected from the Rhode Island Hospital data set. Time to progression from dysplasia to invasive adenocarcinoma was evaluated using Kaplan–Meier curves. Univariate analysis was performed using the log-rank test for categorical groups, whereas the univariate Cox's regression test was used for numerical groups. Multivariate analysis for progression time from dysplasia to invasive carcinoma was performed using the Cox's proportional hazard model.
Statistical and NNET analyzes were performed using the SPSS 11 (SPSS Inc., Chicago, IL, USA) and the Matlab 6.5.1, R13 (The MathWorks, MA, USA) programs, respectively.
Results
The results are separately displayed for the two sets (n=97 and n=55, respectively). The first data set (training set: n=97) was used as a training model for differentiating between the grades of dysplasia and the results were validated using the second data set (testing set: n=55). The study group included biopsies from 68 males and 29 female patients. Ages ranged between 23–85 years (62.9±13) and the grading of dysplasia was achieved by a consensus forum of 12 expert GI pathologists.5 All variables were grouped according to their geometrical representation into major morphologic categories as follows: architecture, nuclear size, shape, pleomorphism, texture, and symmetry. As seen in Table 1, LGD was best differentiated from the ND group by nuclear pseudostratification (P=0.036), pleomorphism (P<0.01) and texture (P<0.01), and from the HGD group by nuclear texture (P<0.01) pleomorphism (P<0.01), symmetry (P<0.01), area (P<0.01), and orientation (P=0.027). No significant variables were found to distinguish between ND and IND groups.
The reproducibility of the morphometric analysis as indicated by the Pearson's coefficient of correlation (R) computed for the linear relationship between two repetitive morphometric measurements, ranged between 0.85 and 0.97 (P<0.01).
Neural Network Model (NNET)
Using the significant variables obtained by the statistical analysis (training set: n=97), two separate neural network algorithms were designed and trained (see Materials and methods for NNET configuration). For differentiating between ND and the LGD groups, the NNET classifier was stopped after 100 learning iterations in order to avoid overtraining of the training set which would reduce its ability to accurately classify the new cases (testing set). The NNET algorithm was able to correctly classify 86% of the cases (70% of ND and 95% of LGD cases were correctly identified). Similarly, in order to differentiate between the LGD and HGD groups (in the training set), the NNET algorithm adequately classified 87% of the cases (correctly identifying 76% of HGD cases and 100% of the LGD cases). For differentiating between ND and HGD, no predictive model was developed as these two categories are usually distinguished with high precision. The ability of this NNET-based morphometric classification system was validated by testing it on new cases (ie testing set) as described below.
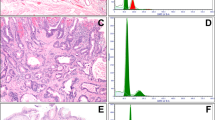
Fifty-five new cases from the archives of pathology at the Rhode Island Hospital were used as a testing group, for validation of the previous NNET algorithm. In this group, the patient age ranged between 32 and 87 years (40 male and 15 female subjects). Twenty-one cases were classified as ND, 15 LGD and 19 HGD. A schematic representation of the NNET configuration used in this model, as well as the error curves associated with the training process are shown in Figure 2. When the NNET algorithm trained on the 97 cases mentioned above was applied to the testing group (n=55), it was able to adequately differentiate ND from LGD in 89% of the cases (80% of ND and 91.7% of LGD correctly identified) and to differentiate LGD from HGD in 85.7% of the cases (71.4% of LGD and 100% of HGD cases correctly identified).
A schematic representation of the Neural Network (NNET) configuration used to predict different grades of dysplasia (a) and the error curves (b) associated with the training process for differentiating between no dysplasia (ND), low-grade dysplasia (LGD), and high-grade dysplasia (HGD), respectively.
Analysis of Progression to Invasive Adenocarcinoma
The intervals for progression to invasive adenocarcinoma were available for the first set of 97 cases. In the ND (n=36), IND (n=16), and LGD (n=21) groups, only three cases progressed to invasive adenocarcinoma. In the HGD category (n=24), 15 progressed to invasive adenocarcinoma (mean progression time of 13 months, range: 1–60 months).
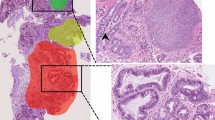
When analyzing the overall relationship between the morphometrical parameters and the time to progression to invasive carcinoma, in all groups (n=24), the best predictors of progression were: grade of dysplasia (P<0.0001), nuclear orientation (P=0.016), pseudostratification (P=0.018), and texture (heterogeneity, P=0.013). A similar analysis was not performed separately within the specific groups owing to the small number of cases which progressed, except for the HGD category. Within the HGD category, the best Cox's univariate predictors of progression time from HGD to invasive adenocarcinoma were nuclear texture (heterogeneity: P=0.0019, margination: P=0.039), variability (OD-s.d.: P=0.03, s.d.-OD: P=0.005, Het-s.d.: P=0.004), and nuclear orientation (P=0.022), as shown in Table 2. Biopsies from patients who progressed exhibited a different textural distribution (heterogeneity) and a less-oriented nuclear pattern when compared to the nonprogressive cases (see Figure 3). Textural heterogeneity was the only significant independent predictor for progression to adenocarcinoma, in the HGD group (Cox's multivariate: P=0.004, OR=11.54, shown in Table 3). Figure 3 displays a Kaplan–Meier curve of interval from diagnosis of HGD to invasive carcinoma, according to a significant cutoff point of the nuclear textural heterogeneity. A visual example of the differences in the textural heterogeneity between progressing vs nonprogressing cases are illustrated in Figure 4.
Discussion
Patients with BE diagnosed with HGD are at the highest risk for progression to invasive adenocarcinoma.5, 6, 7, 29, 30 The diagnosis of HGD may lead to complete esophageal resection, partial resection, or ablation.31, 32, 33, 34 Because the consequences of the diagnosis of HGD in BE are profound, diagnostic accuracy is critical. However, a relatively high rate of interobserver variability in the grading of dysplasia exists among pathologists.9, 10, 11, 12, 13 When asked to classify a given lesion as LGD or HGD, the rate of disagreement among pathologists ranges from 50 to 70%.12 Such interobserver variability may lead to diagnostic inaccuracies and consequently, to erroneous treatment decisions resulting in either an excessive surgical procedure (ie esophagectomy) or to inappropriate delay in the removal of an advanced premalignant lesion.
In an attempt to increase the accuracy of dysplasia grading in BE, a few studies have combined morphometry with the use of molecular markers in order to increase objectivity. Baak et al20 found excellent agreement between certain morphometrical variables such as nuclear area and the stratification index when combined with the proliferation index as compared to histological grading of dysplasia. Other studies that examined quantitative morphometry and p53 expression found a significant decrease in the interobserver variability in the grading of dysplasia.21, 22
As compared to the studies described above, the uniqueness of this study lies in the relatively large number of BE biopsies that were evaluated, and the fact that morphometrical parameters were compared to biopsies graded by a consensus of 12 expert gastrointestinal pathologists.5 In addition, novel morphometric parameters such as indices of nuclear symmetry (size, shape, and texture) were developed and analyzed. Finally, this is the only study which correlated morphometrical parameters with progression to adenocarcinoma in BE patients.
The relatively high pretest probability of Barrett's-associated dysplasia (low+high grade) seen in our study (46%) is due to a selection bias resulting from our aim to address the reproducibility of dysplasia grading rather than to detect dysplasia among the general population of BE patients. We suggest determining whether the criteria identified here will maintain their significance with routinely diagnosed cases of BE.
In this study, LGD could be differentiated from ND by nuclear pseudostratification, pleomorphism and texture and from the HGD group by nuclear texture, pleomorphism, symmetry, area, and orientation. When analyzing the relation between morphometry and time to progression to invasive carcinoma in all of the BE groups, the best independent predictors of progression were: grade of dysplasia, nuclear architecture (orientation, stratification), and chromatin texture (heterogeneity). However, within the HGD group, textural heterogeneity was the only significant independent predictor for progression. Increased textural heterogeneity was associated with an 11.54-fold increased risk for progression from HGD to invasive adenocarcinoma.
In our study, nuclear texture was found to be a powerful variable useful both for differentiating grades of dysplasia and in the prediction of progression. Texture represents the human perception of patterns. Quantification of texture is usually performed using computer vision algorithms. Investigators have used different textural parameters to evaluate a variety of patterns in biology (repetitive, fractal, stochastic, etc.). All textural parameters are designed to quantify the amount and distribution of the gray values of the pixels of an image as well as to assess the mutual relationship between close and remote pixel intensities.
Morphometry has been used to explore the diagnostic or prognostic value of nuclear texture in both cytological and histological tissue specimens. Einstein et al35 utilized fractal and lacunar methods for characterization of chromatin textures in breast cytology and concluded that fractal and lacunarity dimensions are useful tools for the quantitative characterization of chromatin appearance, and can potentially be incorporated into image analysis devices to assure the quality and reproducibility of diagnosis by breast fine-needle aspiration biopsy. Guillaud et al36 were able to predict HPV positivity in cervical cytology, using nuclear chromatin texture analysis.
A few morphometric studies have emphasized the importance of texture as an index of the severity of cellular dysplasia, in diagnostic pathology.37, 38, 39, 40, 41, 42, 43, 44 Deligdish et al40 used an autocorrelation based textural approach for quantifying epithelial nuclear dysplasia in prophylactic oophorectomies obtained from BRCA-positive patients. Weyn et al41 applied different textural descriptors (global, co-occurrence matrix and run length matrix based parameters) for assessing nuclear dysplasia in colonic, esophageal and prostatic premalignant epithelia.
Nuclear texture has been shown to correlate with the degree of nuclear chromatin organization as well as with DNA distribution.42 Changes in the textural pattern of chromatin can be associated with altered gene expression and may indicate premalignant or malignant transformation.42 Khan et al38 found significant associations between nuclear texture, DNA ploidy, and patient outcome in prostate cancer patients. Friedrich et al37 correlated nuclear texture patterns with DNA ploidy along with p53 status and proliferation rate in breast carcinomas. Bartels et al described a predictive role for chromatin textures in the prognosis of prostatic neoplasia.45, 46
It is well known that aneuploidy is associated with dysplasia and progression to carcinoma in BE.8, 15, 17 As mentioned above, there is an association between alterations in ploidy and nuclear texture.37, 38 Therefore, we propose to validate the role of nuclear texture in future studies by using computer-assisted image analysis for evaluating nuclear ploidy of Feulgen-stained BE biopsies from patients with available follow-up information.
In this study, novel morphometric indices of nuclear symmetry were also developed and found to be useful in differentiating between LGD and HGD. Symmetry may be seen as the property of an object being identical on both sides of a central dividing line. Two studies have morphometrically quantified nuclear symmetry in histopathological specimens. Ricco et al47 evaluated the role of nuclear contour symmetry for differentiating between grades of astrocytomas. Bufo et al48 showed that nuclear contour symmetry along with other morphometric variables serve to differentiate between normal, dysplastic, and malignant cells in colonic specimens. Both these studies measured the nuclear contour symmetry based on a Fourier harmonic analysis of the nuclear contours. However, symmetry may be attributed to many other aspects of the nucleus. This is the first study to evaluate the predictive role of multiple indices of shape, size, and textural symmetry in general and in esophageal pathology in particular. In our study, we found contour symmetry and textural symmetry (of margination) to significantly distinguish between HGD and LGD. Textural symmetry was also able to predict progression to carcinoma by Cox's univariate analysis.
Architectural organization of the nuclei was also quantified. The architectural indices measured in this study included the degree of nuclear pseudostratification, nuclear crowding, and nuclear orientation. In our study, nuclear orientation was found to significantly differentiate between LGD and HGD categories. For quantifying nuclear orientation, a semiquantitative score was created whereby two pathologists (MR and ES) independently scored the level of nuclear orientation after evaluating the images both in their original (space) domain and in their frequency domain (Figure 1c). A similar approach was used in other studies for the analysis of periodicity and orientation of the epithelial nuclei in urothelial carcinomas, for increasing the accuracy of the histopathological grading,23 or to quantify the orientation of dermal collagen fibers for the evaluation of different skin pathologies.49, 50
It is well established that architectural changes at the glandular level are also useful in differentiating LGD from HGD. In this study, all of the morphometrical parameters examined were limited to nuclear structure, stratification, and crowding. Computerized variables for the characterization of glandular complexity, crowding, and cribriforming are currently in the course of development and may contribute to increased sensitivity and specificity when introduced into the algorithm.
The main steps in computerized morphometry are sampling, segmentation, and measurement. Digital sampling of the morphometric areas of interest is likely to encounter a degree of interobserver variability, like the one seen in the process of microscopic grading of dysplasia, routinely performed by the pathologist. Nevertheless, multiple studies suggested the use of computerized morphometry as an adjunct tool able to decrease observer variability, especially when grading epithelial dysplasia.22, 23 Polkowski et al,51 support the use of computerized quantitative pathology for reducing diagnostic variability in the grading of dysplasia, during surveillance of patients with BE. The author also recommends limiting the use of computerized quantitative grading of BE dysplasia to expert centers. In this study, we used strict sampling criteria as described in the Materials and methods section.
Another widely documented problem involving instrumental image analysis is segmentation.52 Segmentation is the delimitation of boundaries between two compartments. In our case, it means tracing out precisely the perimeter of the nucleus to separate it from the surrounding cytoplasm in order to be able to individually measure the nuclei. Image analysis is one of the branches of electrical engineering and all its textbooks contain multiple algorithms for scene segmentation that generally are straightforward and dependable.53 In many fields including medical image analysis, automatic segmentation is successful. However, in pathology, the automatic segmentation of the nuclei is still problematic owing to the inherent complexity of the material. Although current digital technology presents powerful quantitative tools, it still lacks the human's qualitative and discriminatory skills. Therefore in pathology, automatic segmentation may not be adequate, especially when measuring highly crowded cellular regions with overlapping nuclei. In such instances, the use of a semiautomatic approach is more appropriate, where digital algorithms of segmentation are manually corrected when needed, by the operator. This method although laboriously intensive, allows the expert to enjoy the automatic tools available in the software and at the same time to exercise full control on the morphometric process.
Along with the laborious interactive (semiquantitative) approach used for the nuclei segmentation, another limiting rate of computerized morphometry is the analysis time. The complex nature of this study including microscopic examination of the slides by the pathologists, sampling of regions of interest, performing a semiautomatic segmentation of the nuclei, measuring the nuclei, database management, statistical, neural (NNET), and reproducibility analyses. However, when strictly considering the kernel of the morphometric analysis, namely the sampling, the segmentation and the measurements, when applied to one biopsy, the analysis time ranged between 15 to 30 min (depending on the size and quality of the biopsy). Despite being a relatively time-consuming analytical method, when considering the profound clinical consequences resulting from a diagnosis of dysplasia, computerized morphometry may be considered as an ancillary method to be applied in borderline cases where grading of dysplasia is controversial or when a pathologist wishes to verify his diagnosis by inputting specific fields to be analyzed according to the present criteria.
The morphometric analysis revealed nuclear pseudostratification, pleomorphism chromatin texture, symmetry, and orientation to be the best criteria for determining the grade of dysplasia. These criteria are largely similar to those used by pathologists to distinguish between the different grades of dysplasia. These findings support the qualitative approach used by the pathologist. Moreover, morphometry offers the pathologist a way to standardize grading, decrease interobserver variability, and to increase the accuracy of the grading of dysplasia in BE.
BE is a particularly fertile area for the use of noninvasive optical imaging by endoscopic techniques.53, 54, 55, 56 Noninvasive optical imaging techniques use a variety of spectral methods and light sources (near-infrared, fluorescent) in order to detect microscopic premalignant changes in the shape or texture of the cells, without the need of a tissue excision. The textural, shape, and architectural variables described in our study may also be relevant to the ongoing efforts to utilize noninvasive optical imaging of dysplasia in BE. Nuclear morphology may be the key for identification of dysplasia during screening, either as a guide for obtaining biopsy, or as a stand-alone technique.
In conclusion, our study illustrates that computerized morphometry is a valid tool for determining the degree of BE-associated dysplasia. Moreover, quantitative evaluation of the nuclear texture pattern is a powerful predictor of time to progression to invasive carcinoma in patients with BE-associated HGD. This is a preliminary study that will have to be validated both prospectively and retrospectively on additional cases where progression was documented. We also suggest that such a system may be used as an adjunct in problematic cases to validate the diagnosis and degree of dysplasia. It is clear that further developments in computer vision technology, such as increased speed of digital scanning and improvements in the field of object segmentation algorithms, will have to be established before such a system can be used routinely as a screening tool.
References
Geboes K, Van Eyken P . The diagnosis of dysplasia and malignancy in Barrett's esophagus. Histopathology 2000;37:99–107.
Goldblum JR . Barrett's esophagus and Barrett's-related dysplasia. Mod Pathol 2003;163:16–24.
Lao CD, Simmons M, Syngal S, et al. Dysplasia in Barrett esophagus. Cancer 2004;100:1622–1627.
Robert ME . Defining dysplasia in Barrett esophagus. J Clin Gastroenterol 2003;36:S19–S25.
Montgomery E, Goldblum JR, Greenson JK, et al. Dysplasia as a predictive marker for invasive carcinoma in Barrett esophagus: a follow-up study based on 138 cases from a diagnostic variability study. Hum Pathol 2001;32:379–388.
Skacel M, Petras RE, Gramlich TL, et al. The diagnosis of low-grade dysplasia in Barrett's esophagus and its implications, for disease progression. Am J Gastroenterol 2000;95:3383–3387.
Haggitt RC . Barrett's esophagus, dysplasia, and adenocarcinoma. Hum Pathol 1994;25:982–993.
Reid BJ, Levine DS, Longton G, et al. Predictors of progression to cancer in Barrett's esophagus: baseline histology and flow cytometry identify low- and high-risk patient subsets. Am J Gastroenterol 2000;95:1669–1676.
Montgomery E, Bronner MP, Goldblum JR, et al. Reproducibility of the diagnosis of dysplasia in Barrett esophagus: a reaffirmation. Hum Pathol 2001;32:368–378.
Montgomery E . Is there a way for pathologists to decrease interobserver variability in the diagnosis of dysplasia? Arch Pathol Lab Med 2005;129:174–176.
Reid BJ, Haggitt RC, Rubin CE, et al. Observer variation in the diagnosis of dysplasia in Barrett's esophagus. Hum Pathol 1988;19:166–178.
Lembo T, Ippoliti AF, Ramers C, et al. Inflammation of the gastroesophageal junction (carditis) in patients with symptomatic gastroesophageal reflux disease: a prospective study. Gut 1999;45:484–488.
Alikhan M, Rex D, Khan A, et al. Variable pathologic interpretation of columnar lined esophagus by general pathologists in community practice. Gastrointest Endoscopy 1999;50:23–26.
Fennerty MB, Sampliner RE, Garewal HS . Review article: Barrett's oesophagus—cancer risk, biology and therapeutic management. Aliment Pharmacol Ther 1993;7:339–345.
Rabinovitch PS, Longton G, Blount PL, et al. Predictors of progression in Barrett's esophagus III: baseline flow cytometric variables. Am J Gastroenterol 2001;96:3071–3083.
Koppert LB, Wijnhoven BP, van Dekken H, et al. The molecular biology of esophageal adenocarcinoma. J Surg Oncol 2005;92:169–190.
Montgomery EA, Hartmann DP, Carr NJ, et al. Barrett esophagus with dysplasia. Flow cytometric DNA analysis of routine, paraffin-embedded mucosal biopsies. Am J Clin Pathol 1996;106:298–304.
Maley CC, Galipeau PC, Li X, et al. The combination of genetic instability and clonal expansion predicts progression to esophageal adenocarcinoma. Cancer Res 2004;64:7629–7633.
Mueller J, Werner M, Stolte M . Barrett's esophagus: histopathologic definitions and diagnostic criteria. World J Surg 2004;28:148–154.
Baak JP, Ten Kate FJ, Offerhaus GJ, et al. Routine morphometrical analysis can improve reproducibility of dysplasia grade in Barrett's oesophagus surveillance biopsies. J Clin Pathol 2002;55:910–916.
Polkowski W, Baak JP, van Lanschot JJ, et al. Clinical decision making in Barrett's oesophagus can be supported by computerized immunoquantitation and morphometry of features associated with proliferation and differentiation. J Pathol 1998;184:161–168.
van Sandick JW, Baak JP, van Lanschot JJ, et al. Computerized quantitative pathology for the grading of dysplasia in surveillance biopsies of Barrett's oesophagus. J Pathol 2000;190:177–183.
Young IT, Verbeek PW, Mayall BH . Characterization of chromatin distribution in cell nuclei. Cytometry 1986;7:467–474.
Sabo E, Gibrat M, Sova Y, et al. Validation of the novel indices of nuclear pleomorphism, polarity and spatial distribution in the grading of urothelial carcinoma. Anal Quant Cytol Histol 2003;25:53–62.
Transnational College of Lex. Who is Fourier? A Mathematical Adventure. Language Research Foundation: Boston, 1995.
Oleksiak MF, Churchill GA, Crawford DL . Variation in gene expression within and among natural populations. Nat Genet 2002;32:261–266.
Adler PN, Liu J, Charlton J . Cell size and the morphogenesis of wing hairs in drosophila. Genesis 2000;28:82–91.
Russel S, Norvig P . Artificial Intelligence. Modern Approach. Prentice Hall International Editions: Los Angeles, 1995.
Schnell TG, Sontag SJ, Chejfec G, et al. Long-term non-surgical management of Barrett's oesophagus with high-grade dysplasia. Gastroenterology 2001;120:1607–1619.
Buttar NS, Wang KK, Sebo TJ, et al. Extent of high-grade dysplasia in Barrett's oesophagus correlates with risk of adenocarcinoma. Gastroenterology 2001;120:1630–1639.
Stein HJ . Esophageal cancer: screening and surveillance. Dis Esophagus 1996;9:3.
van den Boogert J, van Hillegersberg R, Siersema PD, et al. Endoscopic ablation therapy for Barrett's esophagus with high-grade dysplasia: a review. Am J Gastroenterol 1999;94:1153–1160.
Burke AP, Sobin LH, Shekitka KM, et al. Dysplasia of the stomach and Barrett esophagus: a follow-up study. Mod Pathol 1991;4:336–341.
Stein HJ, Feith M, Müller J, et al. Limited resection for early adenocarcinoma in Barrett's esophagus. Ann Surg 2000;232:733–742.
Einstein AJ, Wu HS, Sanchez M, et al. Fractal characterization of chromatin appearance for diagnosis in breast cytology. J Pathol 1998;185:366–381.
Guillaud M . Subvisual chromatin changes in cervical epithelium measured by texture image analysis and correlated with HPV. Gynecol Oncol 2005;99:S16–S23.
Friedrich K, Dimmer V, Haroske G, et al. Correlation between p53 status, DNA ploidy, proliferation rate and nuclear morphology in breast cancer. An image cytometric study. Anal Cell Pathol 1997;15:85–97.
Khan MA, Walsh PC, Miller MC, et al. Quantitative alterations in nuclear structure predict prostate carcinoma distant metastasis and death in men with biochemical recurrence after radical prostatectomy. Cancer 2003;98:2583–2591.
Bartels PH, da Silva VD, Montironi R, et al. Chromatin texture signatures in nuclei from prostate lesions. Anal Quant Cytol Histol 1998;5:407–416.
Deligdish L, Gil J, Kerner H, et al. Ovarian dysplasia in prophylactic oophorectomy specimens: cytogenetic and morphometric correlations. Cancer 1999;86:1544–1550.
Weyn B, Jacob W, da Silva VD, et al. Data representation and reduction for chromatin texture in nuclei from premalignant prostatic, esophageal, and colonic lesions. Cytometry 2000;41:133–138.
Zink D, Fisher AH, Nickerson JA . Nuclear structure in cancer cells. Nat Rev 2004;4:677–688.
Kelloff GJ, Boone CW, Crowell JA, et al. Strategies for phase II cancer chemoprevention trials: cervix, endometrium, and ovary. J Cell Biochem Supp 1995;23:1–9.
Veltri RW, Partin AW, Miller MC . Quantitative Nuclear Grade (QNG): a new image analysis-based biomarker of clinically relevant nuclear structure alterations. J Cell Biochem 2000;S35:151–157.
Bartels PH, Montironi R, Hamilton PW, et al. Nuclear chromatin texture in prostatic lesions. I: PIN and adenocarcinoma. Anal Quant Cytol Histol 1998;20:389–396.
Bartels PH, Montironi R, Hamilton PW, et al. Nuclear chromatin texture in prostatic lesions. II: PIN and malignancy associated changes. Anal Quant Cytol Histol 1998;20:397–406.
Ricco R, Serio G, Caniglia DM, et al. Size and shape evaluation of astrocytoma nuclei with the shape analytical morphometry software system. Anal Quant Cytol Histol 1994;16:345–350.
Bufo R, Ricco R, Potente F, et al. Using analytical morphometry to distinguish severe dysplasia and large bowel carcinoma. Boll Soc Ital Biol Sper 1990;66:143–150.
de Vries HJ, Enomoto DN, van Marle J, et al. Dermal organization in scleroderma: the fast Fourier transform and the laser scatter method objectify fibrosis in nonlesional as well as lesional skin. Lab Invest 2000;80:1281–1289.
Har-Shai Y, Sabo E, Rohde E, et al. Intralesional cryosurgery enhances the involution of recalcitrant auricular keloids: a new clinical approach supported by experimental studies. Wound Repair Regen 2006;14:8–27.
Polkowski W, van Lanschot JJ, van Sandick JW, et al. The role of quantitative pathology in clinical decision making for Barrett's oesophagus. Anal Cell Pathol 2003;25:123–127.
Gil J, Wu H, Wang BY . Image analysis and morphometry in the diagnosis of breast cancer. Microsc Res Tech 2002;59:109–118.
Sharma P, Bansal A, Mathur S, et al. The utility of a novel narrow band imaging endoscopy system in patients with Barrett's esophagus. Gastrointest Endosc 2006;64:167–175.
Wax A, Pyhtila JW, Graf RN, et al. Prospective grading of neoplastic change in rat esophagus epithelium using angle-resolved low-coherence interferometry. J Biomed Opt 2005;10:051604.
Georgakoudi I, Feld MS . The combined use of fluorescence, reflectance, and light-scattering spectroscopy for evaluating dysplasia in Barrett's esophagus. Gastrointest Endosc Clin N Am 2004;14:519–537.
Badizadegan K, Backman V, Boone CW, et al. Spectroscopic diagnosis and imaging of invisible pre-cancer. Faraday Discuss 2004;126:265–279.
Acknowledgements
This project was supported by the Molecular Pathology Core of the COBRE Center for Cancer Research Development, NIH #P20 RR17695, awarded by the National Center for Research Resources, Institutional Development Award (IDeA) Program.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sabo, E., Beck, A., Montgomery, E. et al. Computerized morphometry as an aid in determining the grade of dysplasia and progression to adenocarcinoma in Barrett's esophagus. Lab Invest 86, 1261–1271 (2006). https://doi.org/10.1038/labinvest.3700481
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700481
Keywords
This article is cited by
-
Histopathological distinction of non-invasive and invasive bladder cancers using machine learning approaches
BMC Medical Informatics and Decision Making (2020)
-
Does GRASP affect DCE-MRI quantitative parameters and texture analysis in patients with esophageal cancer receiving preoperative neoadjuvant chemotherapy?
Chinese Journal of Academic Radiology (2019)
-
Large-scale DNA organization is a prognostic marker of breast cancer survival
Medical Oncology (2018)
-
Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features
Nature Communications (2016)
-
Dysplasia discrimination in intestinal-type neoplasia of the esophagus and colon via digital image analysis
Virchows Archiv (2016)