Abstract
A signaling pathway that induces programmed necrotic cell death (necroptosis) was reported to be activated in cells by several cytokines and various pathogen components. The major proteins participating in that pathway are the protein kinases RIPK1 and RIPK3 and the pseudokinase mixed lineage kinase domain-like protein (MLKL). Recent studies have suggested that MLKL, once activated, mediates necroptosis by binding to cellular membranes, thereby triggering ion fluxes. However, our knowledge of both the sequence of molecular events leading to MLKL activation and the subcellular sites of these events is fragmentary. Here we report that the association of MLKL with the cell membrane in necroptotic death is preceded by the translocation of phosphorylated MLKL, along with RIPK1 and RIPK3, to the nucleus.
Similar content being viewed by others
Main
Apart from the apoptotic cell death pathway that ligands of the tumor necrosis factor (TNF) family can activate, these ligands and various other inducers, including the interferons and various pathogen components, have in recent years been found also to trigger a signaling cascade that induces programmed necrotic death (necroptosis). This cascade encompasses sequential activation of the protein kinases RIPK1 and RIPK3 and the pseudokinase mixed lineage kinase domain-like protein (MLKL).1, 2, 3, 4, 5 RIPK3-mediated phosphorylation of MLKL triggers its oligomerization, which is necessary and sufficient for the induction of cell death,6, 7, 8 and can also trigger some non-deadly functions.9 MLKL was recently suggested to trigger cell death by binding to cellular membranes and initiating ion fluxes through them.6, 7, 8, 10 However, its exact molecular target in death induction is contentious.6, 8, 10, 11, 12 Current knowledge of the subcellular sites of MLKL action is based mainly on determination of the location of this protein close to the time of cell death. Here we present a detailed assessment of the cellular location of MLKL at different times following its activation. Our findings indicate that before cell death, MLKL translocates to the nucleus along with RIPK1 and RIPK3.
Results
Induction of necroptosis triggers nuclear translocation of MLKL





On applying specific antibodies to discern endogenous MLKL molecules in HT29 cells, we found that, whereas in the unstimulated cells the protein is located extranuclearly, after the induction of necroptosis by combined treatment with TNF+ BV6+z-VAD.fmk (TBZ) (TNF, 1000 U/ml); the bivalent inhibitor of apoptosis (IAP) antagonist BV6, 1 μM; and benzyloxycarbonyl-Val-Ala-Asp (OMe) fluoromethylketone (z-VAD.fmk, 20 μM), some of the MLKL molecules translocate to the nucleus at an extent that varies greatly among the treated cells (Figure 1a). Three-dimensional (3D) analysis showed that these translocated molecules did not associate with the nuclear membrane (Figure 1b). Nuclear translocation was also discernible when we traced MLKL molecules tagged with green fluorescent protein (GFP; Figure 1c) or with the FLAG epitope (not shown). At early times after TBZ application, the percentage of cells in which this translocation occurred greatly exceeded the percentage of cells that had died (Figure 1d). The frequency of nuclear occurrence of MLKL in cells that had died was much lower than that in cells that were still viable (inset in Figure 1d). Nuclear translocation of MLKL before cell death could also be induced by TBZ in HeLa cells transfected with RIPK3 (data not shown), as well as in both mouse embryonic fibroblasts (MEFs) and mouse L929 cells (see below).
Induction of necroptosis triggers, independent of cell death, translocation of MLKL to the nucleus. (a) Immunocytochemical analysis of MLKL localization in HT29 cells before and after the induction of necroptosis by application of TBZ for 4 h. Unless otherwise indicated, immunocytochemical analyses of MLKL in this paper are presented as merged confocal images of immunostained MLKL (green) and lamin (red, a marker of the nuclear membrane). Scale bars, 10 μm. (b) 3D presentation of immunocytochemical analysis of MLKL localization in HT29 cells, carried out as in a. Blue, cell surface; red, nuclear membrane; green, MLKL. (c).TBZ-induced nuclear translocation of MLKL that was fused N-terminally to GFP (GFP-MLKL) and expressed constitutively in the HT29 cells. Shown are merged confocal images of GFP fluorescence (green) and immunostaining for lamin (red). Scale bars, 10 μm. (d) Kinetics of MLKL nuclear translocation and of death in HT29 cells. () Cells with PI-stained nuclei. () Cells in which only the cytosol stained for MLKL. () Cells in which both the nucleus and the cytosol stained for MLKL. Inset, PI-positive cells in which MLKL staining (as a percentage of total cells in the culture) was observed only in the cytosol () or in both the cytosol and the nucleus (). Shown are the results obtained from 400 counted cells. (e) Western analysis of the induced nuclear accumulation of MLKL. CE, cytosol extract; NE, isolated nuclei. OCT-1 (a nuclear protein), VDAC (an outer mitochondrial membrane protein), and LDH (a cytosolic protein) served as markers for cross-contamination of the subcellular fractions. (f) Blocking of cell death with necrosulfonamide (NSA)4 does not hamper the induced nuclear accumulation of MLKL. Cell viability was assessed by the XTT test. (g) Treatment of caspase-8-deficient dendritic cells with LPS (1 μg/ml), which does not induce death of these cells but rather activates the NALP3 inflammasome via MLKL activation, also triggers nuclear accumulation of MLKL. Each of the experiments described in this article was carried out twice, with qualitatively identical results. All viability tests were done in tetraplicate and in all of them the means±S.D. of the tetraplicates are shown
Further confirming this finding, MLKL could be discerned by both western blotting analysis (Figure 1e) and immunocytochemical analysis (Supplementary Figure S1) of nuclei isolated from HT29 cells that were treated with TBZ, but not after treatment with any of its three constituent compounds alone.
Translocation of MLKL to the nucleus is not a consequence of necroptosis
A hallmark of necrotic death and one of its early events is the disruption of the nuclear membrane. It could, therefore, be suspected that the observed presence of MLKL within the nuclei of cells in which necrotic cell death was induced is merely a manifestation of the failure of the nuclear membrane in necrotic cells to withhold entry of cytosolic proteins. Several observations, however, ruled that out. First, the translocation of MLKL to nuclei occurred before the latter could be stained with propidium iodide (PI), a marker of nuclear membrane permeabilization (Figure 1d). Second, no effect on the nuclear translocation of MLKL was obtained when necroptosis was blocked in human HT29 cells by necrosulfonamide4 (Figure 1f) or in mouse L929 cells by the reducing agent N-acetylcysteine (NAC)13, 14 or the free radical-scavenging agent butylated hydroxyanisole (BHA)15 (Supplementary Figure S2). Third, N-terminal tagging of MLKL molecules, which makes them incapable of triggering necrotic death,7, 16 did not prevent their translocation to the nuclei in response to TBZ (Figure 1c). Finally, MLKL was also found to translocate to the nuclei of caspase-8-deficient mouse dendritic cells treated with lipopolysaccharide (LPS; Figure 1g), which in these cells triggers assembly of the NLRP3 inflammasome in a way that depends on RIPK1, RIPK3, and MLKL and yet is not associated with cell death.9
The nuclearly located MLKL is associated with modified forms of RIPK1 and RIPK3
Western analysis of RIPK1 and RIPK3 in subcellular fractions of HT29 cells (Figure 2a) and of MEFs (Figure 2b) revealed that the induction of necroptosis was followed also by the translocation of some amounts of these proteins to the nuclei. The apparent molecular sizes of most of the nuclear RIPK1 and RIPK3 proteins were larger than those of the RIPK1 and RIPK3 in the noninduced cells, suggesting that the translocation was associated with covalent modifications of these proteins. Co-immunoprecipitation tests revealed that both in the nuclear fraction and in the cytosol of the stimulated cells, the MLKL, RIPK1, and RIPK3 molecules were physically associated (Figure 2c).
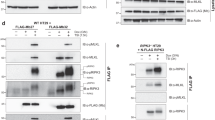
Similar to MLKL, RIPK1 and RIPK3 translocate to the nucleus, and their function is required for MLKL nuclear translocation. (a and b) Western analyses of the nuclear accumulation of MLKL, RIPK1, and RIPK3 in (a) HT29 cells and (b) MEFs after TBZ treatment. (c) Co-immunoprecipitation of RIPK1 and RIPK3 with MLKL from the cytosolic and nuclear extracts of HT29 cells. (d and e) Western analyses of the impact of the knockdown (d) of RIPK1 expression in HT29 cells and (e) of RIPK3 expression in MEFs on the co-translocation of the three proteins to the nucleus (CT, control; K/D, knockdown). (f) Western analysis of the effect of Nec-1 (5 μM) on TBZ-induced nuclear translocation of MLKL
As none of the currently available antibodies against RIPK1 and RIPK3 proved suitable for reliable immunocytochemical localization of the proteins, we expressed in cells RIPK1 and RIPK3 molecules fused to the streptavidin-binding peptide (SBP). As shown in Supplementary Figures S3 and S4, staining of these cells with fluorescent streptavidin further confirmed that both tagged RIPK1 and tagged RIPK3 molecules translocated to the nuclei following TBZ treatment.
Phosphorylation of MLKL targets it to the nucleus
Knockdown of either RIPK1 (Figure 2d) or RIPK3 (Figure 2e) expression, or blocking of the protein kinase activity of RIPK1 using the chemical inhibitor necrostatin-1 (Nec-1; Figure 2f), abolished the TBZ-induced translocation of MLKL to the nucleus, suggesting that this translocation depends on the function of these two protein kinases.
Once activated by RIPK1, RIPK3 phosphorylates MLKL at threonine 357 and serine 358, thereby triggering its mediation of necrotic cell death.4 Immunocytochemical and western analyses using an antibody against the phosphorylated T357/S358 disclosed that in addition to the translocation of some of the phosphorylated MLKL molecules to the cell membrane, a large portion of these molecules translocated to the nucleus (Figures 3a and b). The pattern of immunocytochemical staining with this antibody indicated that the phosphorylated MLKL molecules form large aggregates (Figure 3a). Consistently, the apparent molecular weight of part of the phosphorylated protein detected by western analysis of the nuclear extracts was very large, suggesting the formation of large aggregates that could not be fully dissociated by SDS (Figure 3b).
Nuclear translocation of MLKL is triggered by its phosphorylation and the resulting conformational change in MLKL. (a and b) Immunocytochemical and western analyses of the nuclear translocation of MLKL using an antibody against phosphorylated T357/S358 (ph-MLKL). (a) Immunocytochemical analysis. Arrows indicate association with the plasma membrane. Scale bars, 10 μm. (b) Western analysis. Analysis of the same protein samples with antibody against nonphosphorylated MLKL is shown for comparison. Note that anti-T357/S358 recognizes a high-molecular-weight band that probably corresponds to a large aggregate of the protein in which the epitope recognized by the antibody against nonphosphorylated MLKL is hindered. (c and d) Impact of mutation of the phosphorylation sites in MLKL on the nuclear translocation of MLKL (c and d) and of RIPK1 and RIPK3 (d) in HT29 cells in response to TBZ treatment. Scale bars, 10 μm. (e) Immunocytochemical analysis of the subcellular localizations of WT and ‘phosphomimetic’ MLKL mutants whose expression is induced in HT29 cells. Location of MLKL by the cell membranes (arrows) was determined on the basis of the transmission microscopy images shown at the right. Scale bars, 10 μm
Mutational replacement of threonine 357 or serine 358 in MLKL with alanine blocks the necroptotic activity of this protein (Sun et al.4 and see Figure 4f), whereas their replacement with aspartic or glutamic acid mimics its activation.17 To further define the role of MLKL phosphorylation in the nuclear translocation of this protein, we generated these two kinds of MLKL mutants and expressed them in cells in which the endogenous MLKL had been knocked down. As shown in Figures 3c and d, unlike the wild-type (WT) protein, the ‘nonphosphorylatable’ T357A/S358A MLKL mutant did not translocate to nuclei upon TBZ treatment. In contrast, mere expression of the T357E or S358D MLKL ‘phosphomimetic’ mutant, or (to an even greater extent) of the ‘combined’ T357E/S358D mutant, triggered the formation of large aggregates of the protein that translocated spontaneously to the nuclei (Figure 3e). Some amounts of these mutated proteins also translocated to the plasma membranes.
Nuclear translocation of MLKL is not required for necroptosis but might facilitate it. (a) Immunocytochemical analysis of MLKL expression in HT29 cells induced to express the MLKL K230M/Q356A and the MLKL(1−180) mutants. Scale bars, 10 μm. (b) Impacts of the various mutations in MLKL on the induction of death merely by their expression. TBZ was applied on cells induced to express the WT protein 2 h after 4OHT application. Viability was assessed by the LDH-release assay. (c) Western analysis of the subcellular localization of the indicated MLKL mutants whose expression is induced in HT29 cells for 6 h and 12 h. CE (Aq), cytosolic extract; M (Det), membranous fraction; NE (insoluble), nuclear extract, isolated by phase separation using Triton X-114 as described.8 (d–f) Effect of NLS mutation in MLKL (NLSmt) on (d and e) subcellular localization of the protein and on (f) the extent of cell death in TBZ-treated HT29 cells. The effect of mutations of the phosphorylation sites in MLKL is also shown in (f). Scale bars in d, 10 μm. In f, viability was assessed by the LDH-release assay. (g) Hypothetical model of the impact of the conformational change in phosphorylated MLKL on its subcellular sites, and of the possible interrelationships between the MLKL molecules located at these different sites
These findings indicated that the phosphorylation of MLKL is required for its translocation to the nucleus and, moreover, is sufficient to impose such translocation.
Interestingly, expression of the MLKL T357A/S358A mutant in HT29 cells also blocked the TBZ-induced nuclear translocation of RIPK1 and RIPK3 (Figure 3d), suggesting that the translocation of the two protein kinases depends on that of MLKL.
Translocation of MLKL to the nucleus is dictated by the conformational state induced by its phosphorylation
The RIPK3-mediated phosphorylation of threonine 357 and serine 358 in MLKL – residues that occur in the activation loop of this protein – is believed to trigger ‘opening up’ of a folded form of the protein in which the N-terminal, death-mediating, four-helix bundle (amino acids 1–180) is tethered to the C-terminal pseudokinase domain.17 As a result of this change, these MLKL molecules aggregate through association of their exposed N-terminal helix regions, yielding death-inducing oligomers. Activation of MLKL can also be dictated by mutation of lysine 230 and glutamine 356 in its ATP-binding pocket.17 As shown in Figure 4a, the K230M/Q356A mutant form of MLKL aggregated spontaneously and translocated both to the nuclei and to the plasma membranes.
Nuclear translocation of MLKL is not required for necroptosis but might facilitate it
Deletion of the C-terminal pseudokinase domain of MLKL was reported to render the MLKL molecule spontaneously cytotoxic.7, 10, 16 However, as shown in Figure 4b, the extent of cell death observed in cells expressing the MLKL(1−180) deletion mutant was significantly lower than that induced by the various spontaneously toxic mutants of the full-length molecule. Titration of the effects of these various mutants at different levels of their expression indicated that the observed differences in their death induction are not due to differences in effectiveness of the expression of the proteins or in the extent of their oligomerization, but rather genuinely reflects an inferior ability of the MLKL(1−180) mutant to mediate cell death (Supplementary Figure S5).
Immunocytochemical and western analyses revealed that MLKL(1−180), unlike the other spontaneously cytotoxic MLKL mutants examined in this study, was excluded from the nucleus (Figures 4a and c), suggesting that the C-terminal part of MLKL is required for the nuclear translocation of this protein. Sequence analysis revealed that the C-terminal region contains a bipartite nuclear localization signal (NLS, amino acids 224−256). RIPK3 also contains an NLS, which has been shown to affect the localization of RIPK3 in cells overexpressing this protein kinase.18 However, in view of the failure of RIPK3 to translocate to the nucleus in cells expressing the nonphosphorylatable MLKL mutant (Figure 3d), we reasoned that the NLS found in the latter protein might be more crucially involved in its nuclear translocation. Mutation of this motif indeed significantly decreased both the TBZ-induced nuclear translocation of MLKL, RIPK1, and RIPK3 (Figures 4d and e) and cell death (Figure 4f).
Discussion
Accumulating evidence indicates that MLKL and the protein kinases RIPK1 and RIPK3 contribute to inflammatory processes, both through the induction of necroptotic cell death and through other cellular changes.19, 20 Our finding that the activation of these molecules triggers their translocation to the nucleus is therefore likely to shed new light on the mechanisms underlying the pathogenesis of inflammatory diseases and on the contribution of cell death to inflammation.
The N-terminal four-helix bundle portion of MLKL (the MLKL(1−180) mutant) was recently shown to be capable of triggering necrotic cell death.7, 10, 16 This mutant, as shown here, does not occur in the nucleus, implying that the deadly effector function of MLKL may be independent of any nuclear function of this protein. This is consistent with the evidence that MLKL kills cells upon binding of its N-terminal four-helix bundle to cellular membranes. However, the low effectiveness of death induction by the MLKL(1−180) mutant and the decreased cytotoxicity of MLKL molecules in which the NLS is mutated raise the possibility that nuclear translocation nevertheless does make some contribution to the cytotoxic function of MLKL, either by participating in its activation or by modulating its effector function. We cannot yet exclude the possibility, however, that these mutations interfere with death induction through some other effect on MLKL, unrelated to their impact on its subcellular localization. Accordingly, we cannot rule it out that the nuclear translocation of the signaling proteins for necroptosis actually serves a role unrelated to cell death induction or even has the opposite effect, namely that it reduces signaling for cell death by decreasing the translocation of MLKL to the cell membrane.
Our findings suggest that the conformational change imposed on the MLKL molecule by its phosphorylation results not only in the triggering of cell death mediated by the N-terminal four-helix bundle portion of the protein, but also in the translocation of this molecule to the nucleus through an effect of the NLS motif found in the C-terminal pseudokinase region. Like the N-terminal four-helix motif, the C-terminal NLS motif is apparently also exposed as a result of the conformational change imposed by the phosphorylation of MLKL. The time gap between the initial nuclear accumulation of activated MLKL molecules and cell death suggests that the nuclear translocation of this protein precedes translocation to the cell membrane. It remains to be clarified whether the early translocation of activated MLKL to the nucleus is an initiating step that leads to the eventual translocation of the same molecules to the plasma membrane, or constitutes a different process affecting distinct MLKL molecules (Figure 4g). However, the mere fact that MLKL translocates to the nucleus suggests that besides mediating the effects of cytoplasmic signaling proteins like RIPK1 and RIPK3, this protein is also subject to functional modulation by the activities of some nuclear constituents. Elucidation of the molecular interactions of MLKL, RIPK1, and RIPK3 that specifically occur in the nucleus will make it possible to nail down the functional implications of this translocation and its relationship to the functions already known to be mediated by these signaling proteins.
Materials and methods
Cell culture
Cells of the human HT29 colorectal adenocarcinoma line were grown in McCoy’s 5A medium. Cells of the HeLa cervical adenocarcinoma line and HeLa cells infected with the GEV16/pF5 × UAS lentiviral system21 inducibly expressing RIPK3, as well as MEFs immortalized by expression of the SV40 large T antigen and cells of the mouse L929 line, were cultured in Dulbecco’s modified Eagle’s medium. Both media were supplemented with 10% fetal calf serum, 100 U/ml penicillin, and 100 μg/ml streptomycin.
Mouse bone marrow-derived dendritic cells from which caspase-8 was specifically deleted were produced as described.9
Reagents
TNF was obtained from Ybdy (Seoul, South Korea). The bivalent IAP antagonist BV622 and the caspase inhibitor z-VAD.fmk were purchased from WUXI APPTEC (Shanghai, China). BHA, NAC, bacterial LPS, Nec-1, PI, 4-hydroxytamoxifen (4OHT), and Triton X-114 were from Sigma-Aldrich (St. Louis, MO, USA).
Antibodies
Antibodies against MLKL were purchased from the following sources: GTX107538 for western analysis and immunoprecipitation of human MLKL was from GeneTex (Hsinchu, Taiwan); M6697 for immunocytochemical analysis of human MLKL from Sigma-Aldrich; AP14272b for detection of mouse MLKL from Abgent (San Diego, CA, USA); and Ab187091 for western analysis and immunocytochemistry of phosphorylated T357/S358 in human MLKL8 from Abcam (Cambridge, UK). An antibody (610459) against human and mouse RIPK1 was purchased from BD Biosciences (San Jose, CA, USA). The antibodies 12107 against human RIPK3 and IMG-5523-1 against mouse RIPK3 were purchased from Cell Signalling (Danvers, MA, USA) and Novus Biologicals (Littleton, CO, USA), respectively. The antibodies Ab8980 against human lamin, GTX59803 against OCT-19, 4866 against VDAC, and sc-27230 against lactic dehydrogenase (LDH) were purchased from Abcam, GeneTex, Cell Signalling, and Santacruz (Dallas, TX, USA), respectively. Cy2-conjugated goat anti-rabbit and Cy3-conjugated goat anti-mouse antibodies, as well as Cy2-conjugated streptavidin from Jackson ImmunoResearch Laboratories (West Grow, PA, USA), were used for immunofluorescence staining. Horseradish peroxidase-conjugated mouse anti-rabbit IgG (conformation specific) antibody from Cell Signalling and horseradish peroxidase-conjugated goat anti-mouse light chain specific antibody from Jackson ImmunoResearch Laboratories were used for western blotting analysis after immunoprecipitation. Horseradish peroxidase-conjugated antibodies from Jackson ImmunoResearch were used for other western analyses.
Cytological and immunocytological analyses
For immunofluorescence staining, the cells were fixed with 4% paraformaldehyde in PBS, permeabilized with ice-cold methanol for 5 min at −20 °C, and blocked with a buffer containing 20% normal goat serum and 2% bovine serum albumin in 0.1% Tween-20, 150 mM NaCl, 0.3 M glycine, and 10 mM sodium phosphate (pH 7.2). The cells were then incubated overnight with the indicated antibodies and then with Cy2- or Cy3-conjugated secondary antibodies. Cells expressing proteins fused to the SBP were immunostained with Cy2-conjugated streptavidin.
Images of the immunostained cells and of cells expressing GFP fused to MLKL were captured with the Olympus IX 81 confocal microscope (Olympus imaging inc., Centre Valley, PA, USA), using the UPLSAPO × 60/1.35 NA oil objective and were processed using the Fluoview FV1000 software (Olympus imaging inc.). The 3D images were reconstructed from stacks of serial cross-sections of the cells obtained by confocal microscopy. The image stacks were reconstructed with the surface function in the Imaris software from Bitplane AG (Zurich, Switzerland).
Necroptosis induction
In all experiments, necroptosis was induced by combined treatment with TBZ (TNF, 1000 units/ml; BV6, 1 μM; z-VAD.fmk, 20 μM) for the indicated durations.
Cell death quantification
To quantify cell death, we applied the following tests: (a) nuclear staining by incubation of the cells for 20 min at 37 oC with 5 μg/ml PI in PBS, followed by fixation in 4% paraformaldehyde in PBS and determination of the frequency of the stained cells by microscopic inspection; (b) quantification of the LDH released as a result of cell membrane rupture, by use of the Cytotoxicity Detection Kit (Roche Life Science, Penzberg, Germany); and (c) determination of the amounts of live cells, by use of the XTT Cell Proliferation Kit (Biological Industries, Beit Haemek, Israel).
Nuclear analysis
Nuclei were isolated as described,23 using an extraction buffer containing 10 mM HEPES pH 7.9, 10 mM KCl, 0.1 mM EDTA, 0.5 mM PMSF, 40 mM beta-glycerophosphate, 50 mM NaF, and 1 mM sodium vanadate. To immunoprecipitate the intranuclear proteins, the nuclei were first ruptured by incubation for 20 min at 4 oC in RIPA buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 1% NP-40, 1% sodium deoxycholate, 0.1% SDS). In all western analyses of cytosolic and nuclear extracts, cross-contamination by nuclear, mitochondrial, and cytosolic proteins was assessed by western analyses for the presence of OCT-1 (a nuclear protein), VDAC (a mitochondrial protein), and LDH (a cytosolic protein) in the extracts.
Enrichment of membrane-associated proteins by phase separation
Membrane proteins in Triton X-114 solution were phase separated as described.8
Immunoprecipitation and western analyses
Immunoprecipitation followed by western blotting analysis was done as described.24 To estimate the extent of oligomerization of MLKL, cells were extracted with 1% NP-40 lysis buffer including protease and phosphatase inhibitors, and this was followed by solubilization in Laemmli SDS−PAGE buffer devoid of any reducing agent, as described by Cai et al.6 In all western analyses where some nonspecific bands could be seen, arrows indicate the specific bands.
mRNA knockdown
To knock down the expression of human RIPK1, we used RIPK1-specific On-TARGETplus SMARTpool siRNA (Dharmacon, Lafayette, CO, USA). Expression of mouse RIPK3 was knocked down using lentiviral-expressed shRNA and human MLKL expression was knocked down using 3′-UTR targeting lentiviral shRNA (both from Sigma-Aldrich).
Inducible and constitutive expression of MLKL mutants and fusion proteins
N-terminal fusions of MLKL with GFP and with the FLAG epitope were expressed in the pEF-IRES-Puro and the pcDNA3.1 vectors, respectively. Human RIPK1 that was fused to the SBP tag in the pIRESpuro-GLUE vector was transiently expressed in HT29 cells. Mouse RIPK3 that was fused to the SBP was expressed in MEFs by retroviral infection, and this was followed by selection for cells expressing the infected proteins by growth of the cells in 2 μg/ml of puromycin.
The NLS motif in MLKL, comprising amino acids 224−RAPVAIKVFKKLQAGSIAIVRQTFNKEIKTMKK−256, was identified using the cNLS Mapper software (http://nls-mapper.iab.keio.ac.jp/cgi-bin/NLS_Mapper_form.cgi). To inactivate this motif, it was mutated to the sequence 224−RAPVAIKVFAALQAGSIAIVRQTFNAEIKTMAA−256.
The various mutants of MLKL were expressed constitutively in HT29 cells by their infection with the FUGW lentiviral vector25 and were expressed inducibly by lentiviral infection with the GEV16/pF5 × UAS system.21 Unless otherwise indicated, induction was for 12 h with 20 nM 4OHT.
Abbreviations
- GFP:
-
green fluorescent protein
- IAP:
-
inhibitor of apoptosis
- MLKL:
-
mixed lineage kinase domain-like protein
- NLS:
-
nuclear localization signal
- NSA:
-
necrosulfamide
- PBS:
-
phosphate buffered saline
- PI:
-
propidium iodide
- TBZ:
-
TNF+ BV6+z-VAD.fmk
- z-VAD.fmk:
-
benzyloxycarbonyl-Val-Ala-Asp (OMe) fluoromethylketone
References
Zhang DW, Shao J, Lin J, Zhang N, Lu BJ, Lin SC et al. RIP3, an energy metabolism regulator that switches TNF-induced cell death from apoptosis to necrosis. Science 2009; 325: 332–336.
He S, Wang L, Miao L, Wang T, Du F, Zhao L et al. Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell 2009; 137: 1100–1111.
Cho YS, Challa S, Moquin D, Genga R, Ray TD, Guildford M et al. Phosphorylation-driven assembly of the RIP1-RIP3 complex regulates programmed necrosis and virus-induced inflammation. Cell 2009; 137: 1112–1123.
Sun L, Wang H, Wang Z, He S, Chen S, Liao D et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 2012; 148: 213–227.
Zhao J, Jitkaew S, Cai Z, Choksi S, Li Q, Luo J et al. Mixed lineage kinase domain-like is a key receptor interacting protein 3 downstream component of TNF-induced necrosis. Proc Natl Acad Sci USA 2012; 109: 5322–5327.
Cai Z, Jitkaew S, Zhao J, Chiang HC, Choksi S, Liu J et al. Plasma membrane translocation of trimerized MLKL protein is required for TNF-induced necroptosis. Nat Cell Biol 2014; 16: 55–65.
Chen X, Li W, Ren J, Huang D, He WT, Song Y et al. Translocation of mixed lineage kinase domain-like protein to plasma membrane leads to necrotic cell death. Cell Res 2014; 24: 105–121.
Wang H, Sun L, Su L, Rizo J, Liu L, Wang LF et al. Mixed lineage kinase domain-like protein mlkl causes necrotic membrane disruption upon phosphorylation by RIP3. Mol Cell 2014; 54: 133–146.
Kang TB, Yang SH, Toth B, Kovalenko A, Wallach D . Caspase-8 blocks kinase RIPK3-mediated activation of the NLRP3 inflammasome. Immunity 2013; 38: 27–40.
Dondelinger Y, Declercq W, Montessuit S, Roelandt R, Goncalves A, Bruggeman I et al. MLKL compromises plasma membrane integrity by binding to phosphatidylinositol phosphates. Cell Rep 2014; 7: 971–981.
Wang Z, Jiang H, Chen S, Du F, Wang X . The mitochondrial phosphatase PGAM5 functions at the convergence point of multiple necrotic death pathways. Cell 2012; 148: 228–243.
Chen W, Zhou Z, Li L, Zhong CQ, Zheng X, Wu X et al. Diverse sequence determinants control human and mouse receptor interacting protein 3 (RIP3) and mixed lineage kinase domain-like (MLKL) interaction in necroptotic signaling. J Biol Chem 2013; 288: 16247–16261.
Schulze-Osthoff K, Bakker AC, Vanhaesebroeck B, Beyaert R, Jacob WA, Fiers W . Cytotoxic activity of tumor necrosis factor is mediated by early damage of mitochondrial functions. Evidence for the involvement of mitochondrial radical generation. J Biol Chem 1992; 267: 5317–5323.
Mayer M, Noble M . N-acetyl-L-cysteine is a pluripotent protector against cell death and enhancer of trophic factor-mediated cell survival in vitro. Proc Natl Acad Sci USA 1994; 91: 7496–7500.
Vercammen D, Beyaert R, Denecker G, Goossens V, Van Loo G, Declercq W et al. Inhibition of caspases increases the sensitivity of L929 cells to necrosis mediated by tumor necrosis factor. J Exp Med 1998; 187: 1477–1485.
Hildebrand JM, Tanzer MC, Lucet IS, Young SN, Spall SK, Sharma P et al. Activation of the pseudokinase MLKL unleashes the four-helix bundle domain to induce membrane localization and necroptotic cell death. Proc Natl Acad Sci USA 2014; 111: 15072–15077.
Murphy JM, Czabotar PE, Hildebrand JM, Lucet IS, Zhang JG, Alvarez-Diaz S et al. The pseudokinase MLKL mediates necroptosis via a molecular switch mechanism. Immunity 2013; 39: 443–453.
Yang Y, Ma J, Chen Y, Wu M . Nucleocytoplasmic shuttling of receptor-interacting protein 3 (RIP3): identification of novel nuclear export and import signals in RIP3. J Biol Chem 2004; 279: 38820–38829.
Ofengeim D, Yuan J . Regulation of RIP1 kinase signalling at the crossroads of inflammation and cell death. Nat Rev Mol Cell Biol 2013; 14: 727–736.
Chan FK, Luz NF, Moriwaki K . Programmed necrosis in the cross talk of cell death and inflammation. Annu Rev Immunol 2015; 33: 79–106.
Dunning CJ, McKenzie M, Sugiana C, Lazarou M, Silke J, Connelly A et al. Human CIA30 is involved in the early assembly of mitochondrial complex I and mutations in its gene cause disease. EMBO J 2007; 26: 3227–3237.
Varfolomeev E, Blankenship JW, Wayson SM, Fedorova AV, Kayagaki N, Garg P et al. IAP antagonists induce autoubiquitination of c-IAPs, NF-kappaB activation, and TNFalpha-dependent apoptosis. Cell 2007; 131: 669–681.
Schreiber E, Matthias P, Muller MM, Schaffner W . Rapid detection of octamer binding proteins with ‘mini-extracts’, prepared from a small number of cells. Nucleic Acids Res 1989; 17: 6419.
Rajput A, Kovalenko A, Bogdanov K, Yang SH, Kang TB, Kim JC et al. RIG-I RNA helicase activation of IRF3 transcription factor is negatively regulated by caspase-8-mediated cleavage of the RIP1 protein. Immunity 2011; 34: 340–351.
Lois C, Hong EJ, Pease S, Brown EJ, Baltimore D . Germline transmission and tissue-specific expression of transgenes delivered by lentiviral vectors. Science 2002; 295: 868–872.
Acknowledgements
We thank Dr. Tae-Bong Kang for useful advice and Vladimir Kiss and Dr. Reinat Nevo for their advice and help with the immunocytochemical analyses. This study was supported in part by grants from the Kekst Family Center for Medical Genetics and the Shapell Family Center for Genetic Disorders Research at The Weizmann Institute of Science. DW is the incumbent of the Joseph and Bessie Feinberg Professorial Chair at The Weizmann Institute of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by S Nagata
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Yoon, S., Bogdanov, K., Kovalenko, A. et al. Necroptosis is preceded by nuclear translocation of the signaling proteins that induce it. Cell Death Differ 23, 253–260 (2016). https://doi.org/10.1038/cdd.2015.92
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2015.92
This article is cited by
-
Site-specific ubiquitination of MLKL targets it to endosomes and targets Listeria and Yersinia to the lysosomes
Cell Death & Differentiation (2022)
-
Quantitative analysis of phosphoproteome in necroptosis reveals a role of TRIM28 phosphorylation in promoting necroptosis-induced cytokine production
Cell Death & Disease (2021)
-
RIPK3 signaling and its role in the pathogenesis of cancers
Cellular and Molecular Life Sciences (2021)
-
Neuron-specific activation of necroptosis signaling in multiple sclerosis cortical grey matter
Acta Neuropathologica (2021)
-
A necroptotic-independent function of MLKL in regulating endothelial cell adhesion molecule expression
Cell Death & Disease (2020)